Jackie Ma
MADE: A Living Benchmark for Multi-Label Text Classification with Uncertainty Quantification of Medical Device Adverse Events
Apr 16, 2026Abstract:Machine learning in high-stakes domains such as healthcare requires not only strong predictive performance but also reliable uncertainty quantification (UQ) to support human oversight. Multi-label text classification (MLTC) is a central task in this domain, yet remains challenging due to label imbalances, dependencies, and combinatorial complexity. Existing MLTC benchmarks are increasingly saturated and may be affected by training data contamination, making it difficult to distinguish genuine reasoning capabilities from memorization. We introduce MADE, a living MLTC benchmark derived from {m}edical device {ad}verse {e}vent reports and continuously updated with newly published reports to prevent contamination. MADE features a long-tailed distribution of hierarchical labels and enables reproducible evaluation with strict temporal splits. We establish baselines across more than 20 encoder- and decoder-only models under fine-tuning and few-shot settings (instruction-tuned/reasoning variants, local/API-accessible). We systematically assess entropy-/consistency-based and self-verbalized UQ methods. Results show clear trade-offs: smaller discriminatively fine-tuned decoders achieve the strongest head-to-tail accuracy while maintaining competitive UQ; generative fine-tuning delivers the most reliable UQ; large reasoning models improve performance on rare labels yet exhibit surprisingly weak UQ; and self-verbalized confidence is not a reliable proxy for uncertainty. Our work is publicly available at https://hhi.fraunhofer.de/aml-demonstrator/made-benchmark.
Structural Compactness as a Complementary Criterion for Explanation Quality
Mar 31, 2026Abstract:In the evaluation of attribution quality, the quantitative assessment of explanation legibility is particularly difficult, as it is influenced by varying shapes and internal organization of attributions not captured by simple statistics. To address this issue, we introduce Minimum Spanning Tree Compactness (MST-C), a graph-based structural metric that captures higher-order geometric properties of attributions, such as spread and cohesion. These components are combined into a single score that evaluates compactness, favoring attributions with salient points spread across a small area and spatially organized into few but cohesive clusters. We show that MST-C reliably distinguishes between explanation methods, exposes fundamental structural differences between models, and provides a robust, self-contained diagnostic for explanation compactness that complements existing notions of attribution complexity.
X-SYS: A Reference Architecture for Interactive Explanation Systems
Feb 13, 2026Abstract:The explainable AI (XAI) research community has proposed numerous technical methods, yet deploying explainability as systems remains challenging: Interactive explanation systems require both suitable algorithms and system capabilities that maintain explanation usability across repeated queries, evolving models and data, and governance constraints. We argue that operationalizing XAI requires treating explainability as an information systems problem where user interaction demands induce specific system requirements. We introduce X-SYS, a reference architecture for interactive explanation systems, that guides (X)AI researchers, developers and practitioners in connecting interactive explanation user interfaces (XUI) with system capabilities. X-SYS organizes around four quality attributes named STAR (scalability, traceability, responsiveness, and adaptability), and specifies a five-component decomposition (XUI Services, Explanation Services, Model Services, Data Services, Orchestration and Governance). It maps interaction patterns to system capabilities to decouple user interface evolution from backend computation. We implement X-SYS through SemanticLens, a system for semantic search and activation steering in vision-language models. SemanticLens demonstrates how contract-based service boundaries enable independent evolution, offline/online separation ensures responsiveness, and persistent state management supports traceability. Together, this work provides a reusable blueprint and concrete instantiation for interactive explanation systems supporting end-to-end design under operational constraints.
Metric Hub: A metric library and practical selection workflow for use-case-driven data quality assessment in medical AI
Jan 30, 2026Abstract:Machine learning (ML) in medicine has transitioned from research to concrete applications aimed at supporting several medical purposes like therapy selection, monitoring and treatment. Acceptance and effective adoption by clinicians and patients, as well as regulatory approval, require evidence of trustworthiness. A major factor for the development of trustworthy AI is the quantification of data quality for AI model training and testing. We have recently proposed the METRIC-framework for systematically evaluating the suitability (fit-for-purpose) of data for medical ML for a given task. Here, we operationalize this theoretical framework by introducing a collection of data quality metrics - the metric library - for practically measuring data quality dimensions. For each metric, we provide a metric card with the most important information, including definition, applicability, examples, pitfalls and recommendations, to support the understanding and implementation of these metrics. Furthermore, we discuss strategies and provide decision trees for choosing an appropriate set of data quality metrics from the metric library given specific use cases. We demonstrate the impact of our approach exemplarily on the PTB-XL ECG-dataset. This is a first step to enable fit-for-purpose evaluation of training and test data in practice as the base for establishing trustworthy AI in medicine.
Multimodal Deep Learning for Prediction of Progression-Free Survival in Patients with Neuroendocrine Tumors Undergoing 177Lu-based Peptide Receptor Radionuclide Therapy
Nov 07, 2025
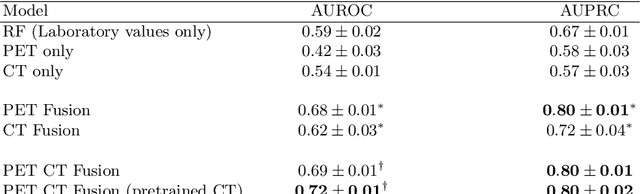
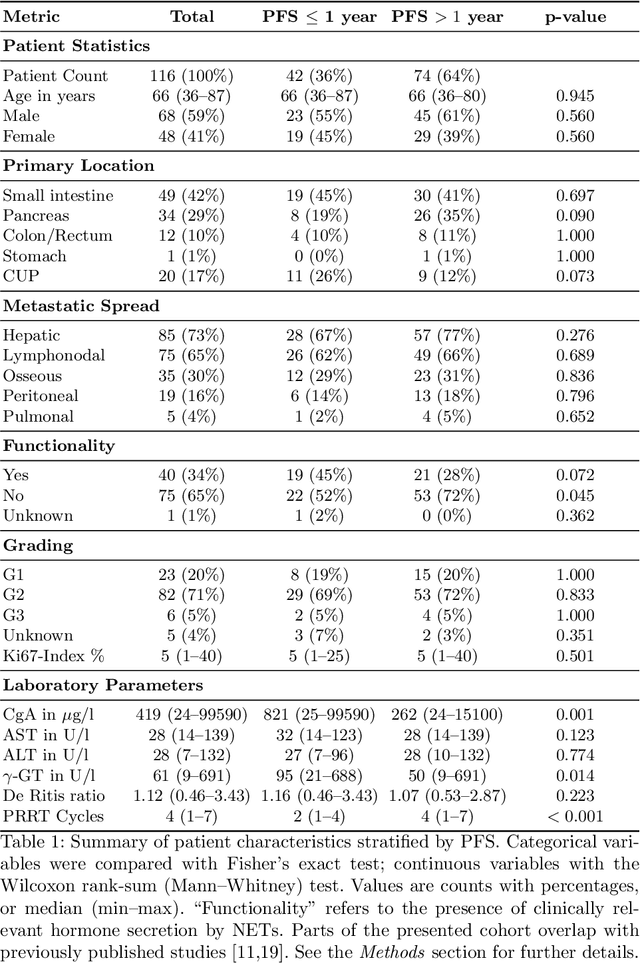


Abstract:Peptide receptor radionuclide therapy (PRRT) is an established treatment for metastatic neuroendocrine tumors (NETs), yet long-term disease control occurs only in a subset of patients. Predicting progression-free survival (PFS) could support individualized treatment planning. This study evaluates laboratory, imaging, and multimodal deep learning models for PFS prediction in PRRT-treated patients. In this retrospective, single-center study 116 patients with metastatic NETs undergoing 177Lu-DOTATOC were included. Clinical characteristics, laboratory values, and pretherapeutic somatostatin receptor positron emission tomography/computed tomographies (SR-PET/CT) were collected. Seven models were trained to classify low- vs. high-PFS groups, including unimodal (laboratory, SR-PET, or CT) and multimodal fusion approaches. Explainability was evaluated by feature importance analysis and gradient maps. Forty-two patients (36%) had short PFS (< 1 year), 74 patients long PFS (>1 year). Groups were similar in most characteristics, except for higher baseline chromogranin A (p = 0.003), elevated gamma-GT (p = 0.002), and fewer PRRT cycles (p < 0.001) in short-PFS patients. The Random Forest model trained only on laboratory biomarkers reached an AUROC of 0.59 +- 0.02. Unimodal three-dimensional convolutional neural networks using SR-PET or CT performed worse (AUROC 0.42 +- 0.03 and 0.54 +- 0.01, respectively). A multimodal fusion model laboratory values, SR-PET, and CT -augmented with a pretrained CT branch - achieved the best results (AUROC 0.72 +- 0.01, AUPRC 0.80 +- 0.01). Multimodal deep learning combining SR-PET, CT, and laboratory biomarkers outperformed unimodal approaches for PFS prediction after PRRT. Upon external validation, such models may support risk-adapted follow-up strategies.
Spatially Resolved Meteorological and Ancillary Data in Central Europe for Rainfall Streamflow Modeling
Jun 04, 2025Abstract:We present a dataset for rainfall streamflow modeling that is fully spatially resolved with the aim of taking neural network-driven hydrological modeling beyond lumped catchments. To this end, we compiled data covering five river basins in central Europe: upper Danube, Elbe, Oder, Rhine, and Weser. The dataset contains meteorological forcings, as well as ancillary information on soil, rock, land cover, and orography. The data is harmonized to a regular 9km times 9km grid and contains daily values that span from October 1981 to September 2011. We also provide code to further combine our dataset with publicly available river discharge data for end-to-end rainfall streamflow modeling.
DiffScale: Continuous Downscaling and Bias Correction of Subseasonal Wind Speed Forecasts using Diffusion Models
Mar 31, 2025Abstract:Renewable resources are strongly dependent on local and large-scale weather situations. Skillful subseasonal to seasonal (S2S) forecasts -- beyond two weeks and up to two months -- can offer significant socioeconomic advantages to the energy sector. This study aims to enhance wind speed predictions using a diffusion model with classifier-free guidance to downscale S2S forecasts of surface wind speed. We propose DiffScale, a diffusion model that super-resolves spatial information for continuous downscaling factors and lead times. Leveraging weather priors as guidance for the generative process of diffusion models, we adopt the perspective of conditional probabilities on sampling super-resolved S2S forecasts. We aim to directly estimate the density associated with the target S2S forecasts at different spatial resolutions and lead times without auto-regression or sequence prediction, resulting in an efficient and flexible model. Synthetic experiments were designed to super-resolve wind speed S2S forecasts from the European Center for Medium-Range Weather Forecast (ECMWF) from a coarse resolution to a finer resolution of ERA5 reanalysis data, which serves as a high-resolution target. The innovative aspect of DiffScale lies in its flexibility to downscale arbitrary scaling factors, enabling it to generalize across various grid resolutions and lead times -without retraining the model- while correcting model errors, making it a versatile tool for improving S2S wind speed forecasts. We achieve a significant improvement in prediction quality, outperforming baselines up to week 3.
Synthetic Datasets for Machine Learning on Spatio-Temporal Graphs using PDEs
Feb 06, 2025Abstract:Many physical processes can be expressed through partial differential equations (PDEs). Real-world measurements of such processes are often collected at irregularly distributed points in space, which can be effectively represented as graphs; however, there are currently only a few existing datasets. Our work aims to make advancements in the field of PDE-modeling accessible to the temporal graph machine learning community, while addressing the data scarcity problem, by creating and utilizing datasets based on PDEs. In this work, we create and use synthetic datasets based on PDEs to support spatio-temporal graph modeling in machine learning for different applications. More precisely, we showcase three equations to model different types of disasters and hazards in the fields of epidemiology, atmospheric particles, and tsunami waves. Further, we show how such created datasets can be used by benchmarking several machine learning models on the epidemiological dataset. Additionally, we show how pre-training on this dataset can improve model performance on real-world epidemiological data. The presented methods enable others to create datasets and benchmarks customized to individual requirements. The source code for our methodology and the three created datasets can be found on https://github.com/github-usr-ano/Temporal_Graph_Data_PDEs.
Spatial Shortcuts in Graph Neural Controlled Differential Equations
Oct 25, 2024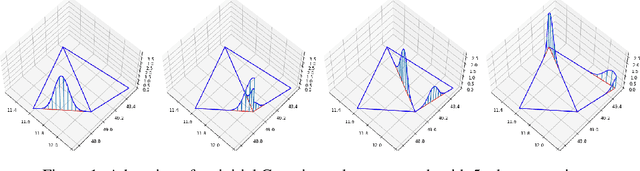

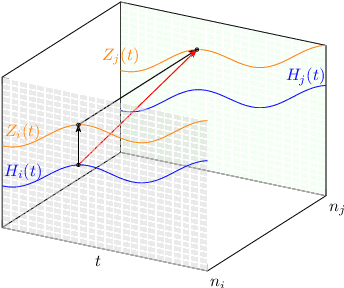

Abstract:We incorporate prior graph topology information into a Neural Controlled Differential Equation (NCDE) to predict the future states of a dynamical system defined on a graph. The informed NCDE infers the future dynamics at the vertices of simulated advection data on graph edges with a known causal graph, observed only at vertices during training. We investigate different positions in the model architecture to inform the NCDE with graph information and identify an outer position between hidden state and control as theoretically and empirically favorable. Our such informed NCDE requires fewer parameters to reach a lower Mean Absolute Error (MAE) compared to previous methods that do not incorporate additional graph topology information.
Synthetic Generation of Dermatoscopic Images with GAN and Closed-Form Factorization
Oct 07, 2024



Abstract:In the realm of dermatological diagnoses, where the analysis of dermatoscopic and microscopic skin lesion images is pivotal for the accurate and early detection of various medical conditions, the costs associated with creating diverse and high-quality annotated datasets have hampered the accuracy and generalizability of machine learning models. We propose an innovative unsupervised augmentation solution that harnesses Generative Adversarial Network (GAN) based models and associated techniques over their latent space to generate controlled semiautomatically-discovered semantic variations in dermatoscopic images. We created synthetic images to incorporate the semantic variations and augmented the training data with these images. With this approach, we were able to increase the performance of machine learning models and set a new benchmark amongst non-ensemble based models in skin lesion classification on the HAM10000 dataset; and used the observed analytics and generated models for detailed studies on model explainability, affirming the effectiveness of our solution.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge