Hongjie Yan
LEREL: Lipschitz Continuity-Constrained Emotion Recognition Ensemble Learning For Electroencephalography
Apr 12, 2025


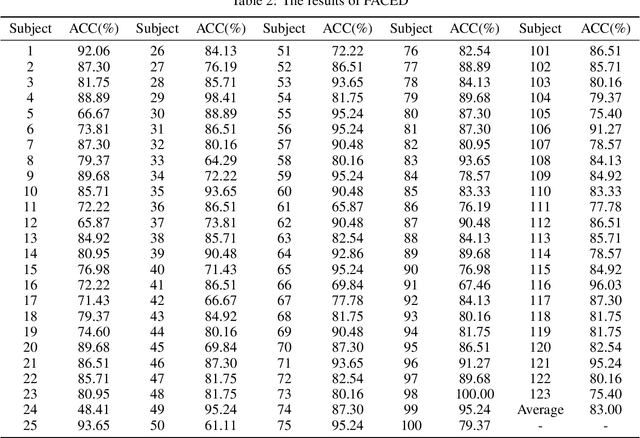
Abstract:Accurate and efficient perception of emotional states in oneself and others is crucial, as emotion-related disorders are associated with severe psychosocial impairments. While electroencephalography (EEG) offers a powerful tool for emotion detection, current EEG-based emotion recognition (EER) methods face key limitations: insufficient model stability, limited accuracy in processing high-dimensional nonlinear EEG signals, and poor robustness against intra-subject variability and signal noise. To address these challenges, we propose LEREL (Lipschitz continuity-constrained Emotion Recognition Ensemble Learning), a novel framework that significantly enhances both the accuracy and robustness of emotion recognition performance. The LEREL framework employs Lipschitz continuity constraints to enhance model stability and generalization in EEG emotion recognition, reducing signal variability and noise susceptibility while maintaining strong performance on small-sample datasets. The ensemble learning strategy reduces single-model bias and variance through multi-classifier decision fusion, further optimizing overall performance. Experimental results on three public benchmark datasets (EAV, FACED and SEED) demonstrate LEREL's effectiveness, achieving average recognition accuracies of 76.43%, 83.00% and 89.22%, respectively.
Information Bottleneck-Guided Heterogeneous Graph Learning for Interpretable Neurodevelopmental Disorder Diagnosis
Feb 28, 2025
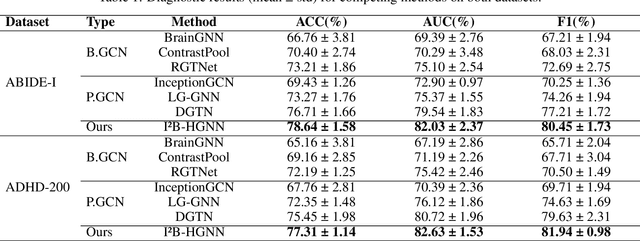
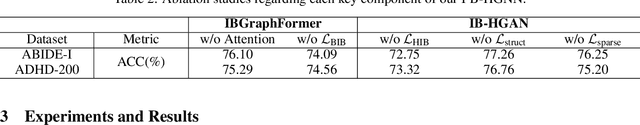
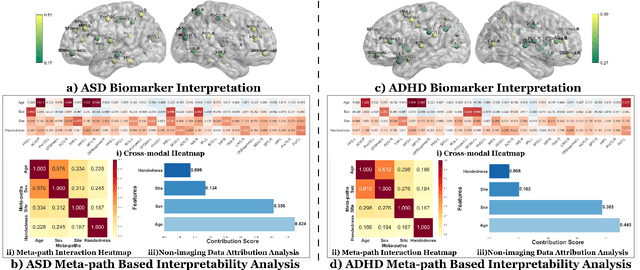
Abstract:Developing interpretable models for diagnosing neurodevelopmental disorders (NDDs) is highly valuable yet challenging, primarily due to the complexity of encoding, decoding and integrating imaging and non-imaging data. Many existing machine learning models struggle to provide comprehensive interpretability, often failing to extract meaningful biomarkers from imaging data, such as functional magnetic resonance imaging (fMRI), or lacking mechanisms to explain the significance of non-imaging data. In this paper, we propose the Interpretable Information Bottleneck Heterogeneous Graph Neural Network (I2B-HGNN), a novel framework designed to learn from fine-grained local patterns to comprehensive global multi-modal interactions. This framework comprises two key modules. The first module, the Information Bottleneck Graph Transformer (IBGraphFormer) for local patterns, integrates global modeling with brain connectomic-constrained graph neural networks to identify biomarkers through information bottleneck-guided pooling. The second module, the Information Bottleneck Heterogeneous Graph Attention Network (IB-HGAN) for global multi-modal interactions, facilitates interpretable multi-modal fusion of imaging and non-imaging data using heterogeneous graph neural networks. The results of the experiments demonstrate that I2B-HGNN excels in diagnosing NDDs with high accuracy, providing interpretable biomarker identification and effective analysis of non-imaging data.
STARFormer: A Novel Spatio-Temporal Aggregation Reorganization Transformer of FMRI for Brain Disorder Diagnosis
Dec 31, 2024



Abstract:Many existing methods that use functional magnetic resonance imaging (fMRI) classify brain disorders, such as autism spectrum disorder (ASD) and attention deficit hyperactivity disorder (ADHD), often overlook the integration of spatial and temporal dependencies of the blood oxygen level-dependent (BOLD) signals, which may lead to inaccurate or imprecise classification results. To solve this problem, we propose a Spatio-Temporal Aggregation eorganization ransformer (STARFormer) that effectively captures both spatial and temporal features of BOLD signals by incorporating three key modules. The region of interest (ROI) spatial structure analysis module uses eigenvector centrality (EC) to reorganize brain regions based on effective connectivity, highlighting critical spatial relationships relevant to the brain disorder. The temporal feature reorganization module systematically segments the time series into equal-dimensional window tokens and captures multiscale features through variable window and cross-window attention. The spatio-temporal feature fusion module employs a parallel transformer architecture with dedicated temporal and spatial branches to extract integrated features. The proposed STARFormer has been rigorously evaluated on two publicly available datasets for the classification of ASD and ADHD. The experimental results confirm that the STARFormer achieves state-of-the-art performance across multiple evaluation metrics, providing a more accurate and reliable tool for the diagnosis of brain disorders and biomedical research. The codes will be available at: https://github.com/NZWANG/STARFormer.
Neural-MCRL: Neural Multimodal Contrastive Representation Learning for EEG-based Visual Decoding
Dec 23, 2024



Abstract:Decoding neural visual representations from electroencephalogram (EEG)-based brain activity is crucial for advancing brain-machine interfaces (BMI) and has transformative potential for neural sensory rehabilitation. While multimodal contrastive representation learning (MCRL) has shown promise in neural decoding, existing methods often overlook semantic consistency and completeness within modalities and lack effective semantic alignment across modalities. This limits their ability to capture the complex representations of visual neural responses. We propose Neural-MCRL, a novel framework that achieves multimodal alignment through semantic bridging and cross-attention mechanisms, while ensuring completeness within modalities and consistency across modalities. Our framework also features the Neural Encoder with Spectral-Temporal Adaptation (NESTA), a EEG encoder that adaptively captures spectral patterns and learns subject-specific transformations. Experimental results demonstrate significant improvements in visual decoding accuracy and model generalization compared to state-of-the-art methods, advancing the field of EEG-based neural visual representation decoding in BMI. Codes will be available at: https://github.com/NZWANG/Neural-MCRL.
STANet: A Novel Spatio-Temporal Aggregation Network for Depression Classification with Small and Unbalanced FMRI Data
Jul 31, 2024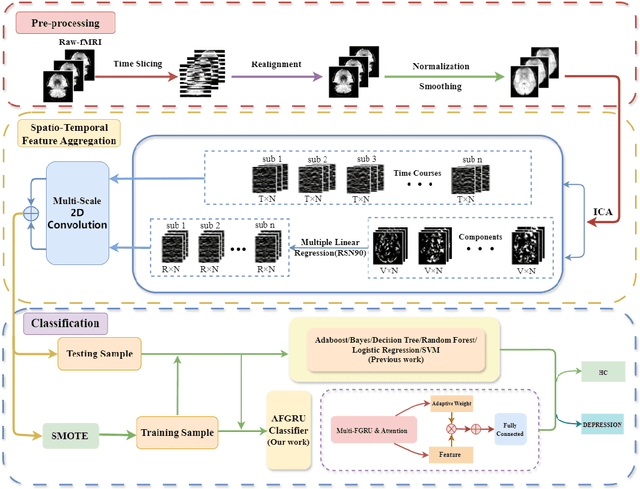
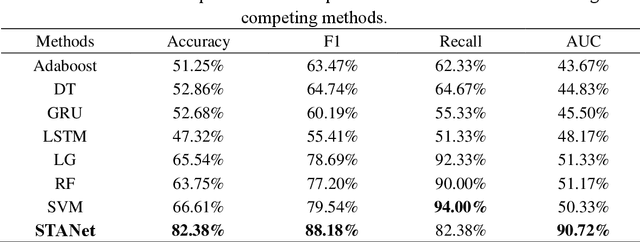
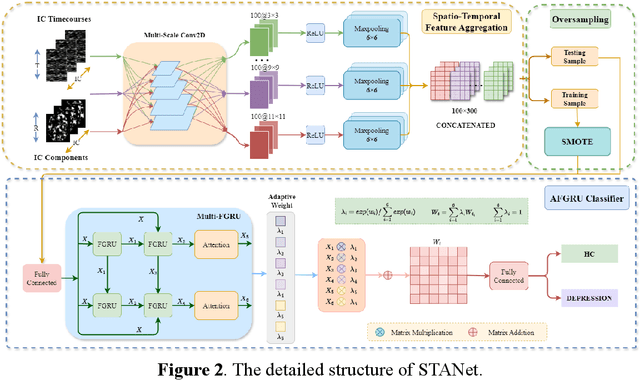
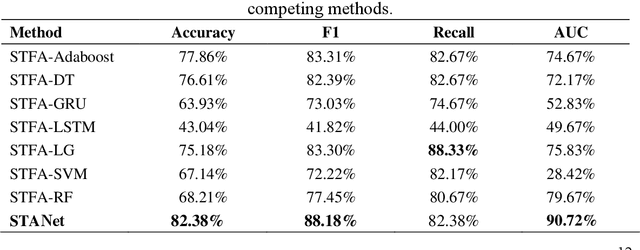
Abstract:Accurate diagnosis of depression is crucial for timely implementation of optimal treatments, preventing complications and reducing the risk of suicide. Traditional methods rely on self-report questionnaires and clinical assessment, lacking objective biomarkers. Combining fMRI with artificial intelligence can enhance depression diagnosis by integrating neuroimaging indicators. However, the specificity of fMRI acquisition for depression often results in unbalanced and small datasets, challenging the sensitivity and accuracy of classification models. In this study, we propose the Spatio-Temporal Aggregation Network (STANet) for diagnosing depression by integrating CNN and RNN to capture both temporal and spatial features of brain activity. STANet comprises the following steps:(1) Aggregate spatio-temporal information via ICA. (2) Utilize multi-scale deep convolution to capture detailed features. (3) Balance data using the SMOTE to generate new samples for minority classes. (4) Employ the AFGRU classifier, which combines Fourier transformation with GRU, to capture long-term dependencies, with an adaptive weight assignment mechanism to enhance model generalization. The experimental results demonstrate that STANet achieves superior depression diagnostic performance with 82.38% accuracy and a 90.72% AUC. The STFA module enhances classification by capturing deeper features at multiple scales. The AFGRU classifier, with adaptive weights and stacked GRU, attains higher accuracy and AUC. SMOTE outperforms other oversampling methods. Additionally, spatio-temporal aggregated features achieve better performance compared to using only temporal or spatial features. STANet outperforms traditional or deep learning classifiers, and functional connectivity-based classifiers, as demonstrated by ten-fold cross-validation.
Neural Modulation Alteration to Positive and Negative Emotions in Depressed Patients: Insights from fMRI Using Positive/Negative Emotion Atlas
Jul 26, 2024
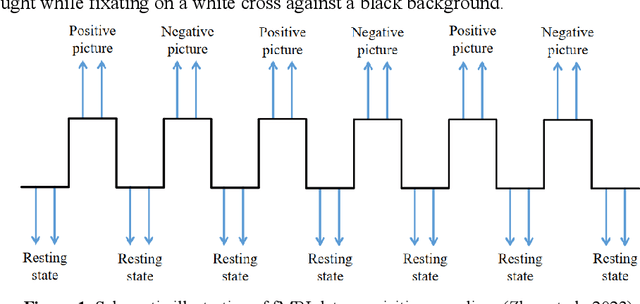


Abstract:Background: Although it has been noticed that depressed patients show differences in processing emotions, the precise neural modulation mechanisms of positive and negative emotions remain elusive. FMRI is a cutting-edge medical imaging technology renowned for its high spatial resolution and dynamic temporal information, making it particularly suitable for the neural dynamics of depression research. Methods: To address this gap, our study firstly leveraged fMRI to delineate activated regions associated with positive and negative emotions in healthy individuals, resulting in the creation of positive emotion atlas (PEA) and negative emotion atlas (NEA). Subsequently, we examined neuroimaging changes in depression patients using these atlases and evaluated their diagnostic performance based on machine learning. Results: Our findings demonstrate that the classification accuracy of depressed patients based on PEA and NEA exceeded 0.70, a notable improvement compared to the whole-brain atlases. Furthermore, ALFF analysis unveiled significant differences between depressed patients and healthy controls in eight functional clusters during the NEA, focusing on the left cuneus, cingulate gyrus, and superior parietal lobule. In contrast, the PEA revealed more pronounced differences across fifteen clusters, involving the right fusiform gyrus, parahippocampal gyrus, and inferior parietal lobule. Limitations: Due to the limited sample size and subtypes of depressed patients, the efficacy may need further validation in future. Conclusions: These findings emphasize the complex interplay between emotion modulation and depression, showcasing significant alterations in both PEA and NEA among depression patients. This research enhances our understanding of emotion modulation in depression, with implications for diagnosis and treatment evaluation.
A Tale of Single-channel Electroencephalogram: Devices, Datasets, Signal Processing, Applications, and Future Directions
Jul 20, 2024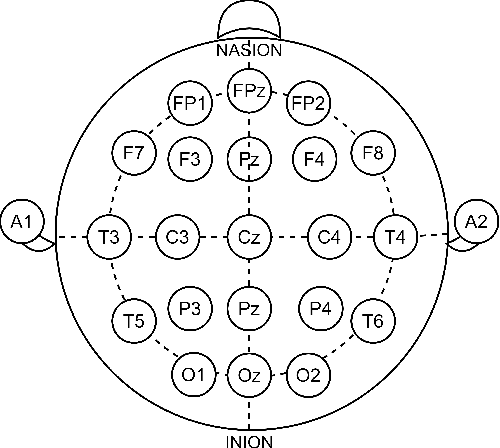
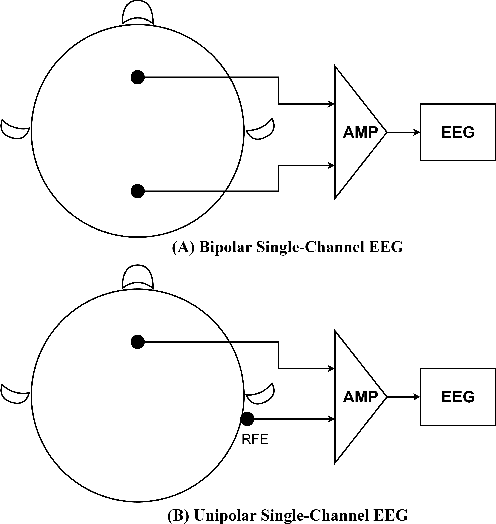
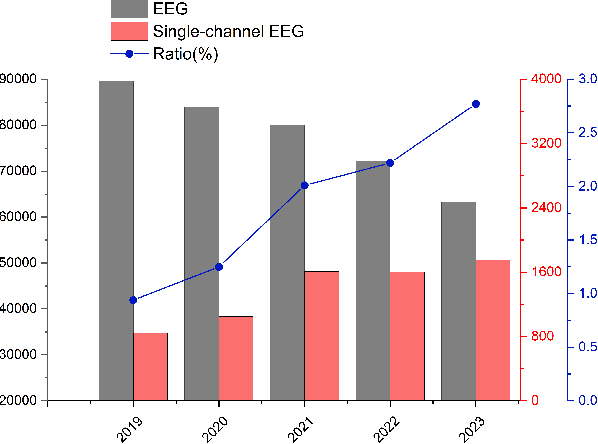

Abstract:Single-channel electroencephalogram (EEG) is a cost-effective, comfortable, and non-invasive method for monitoring brain activity, widely adopted by researchers, consumers, and clinicians. The increasing number and proportion of articles on single-channel EEG underscore its growing potential. This paper provides a comprehensive review of single-channel EEG, focusing on development trends, devices, datasets, signal processing methods, recent applications, and future directions. Definitions of bipolar and unipolar configurations in single-channel EEG are clarified to guide future advancements. Applications mainly span sleep staging, emotion recognition, educational research, and clinical diagnosis. Ongoing advancements of single-channel EEG in AI-based EEG generation techniques suggest potential parity or superiority over multichannel EEG performance.
MHNet: Multi-view High-order Network for Diagnosing Neurodevelopmental Disorders Using Resting-state fMRI
Jul 03, 2024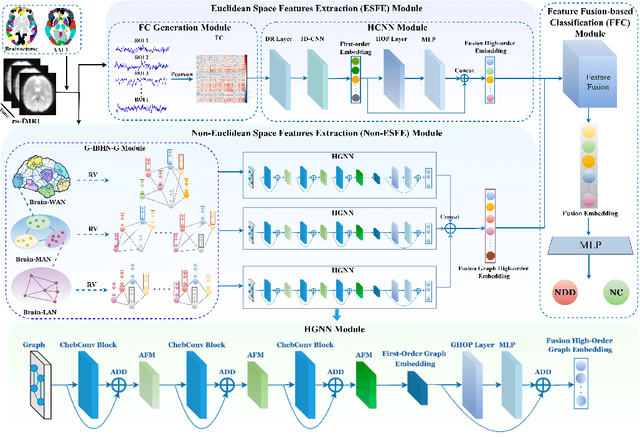
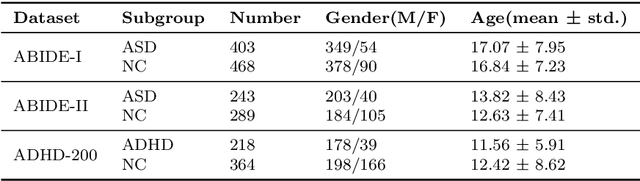
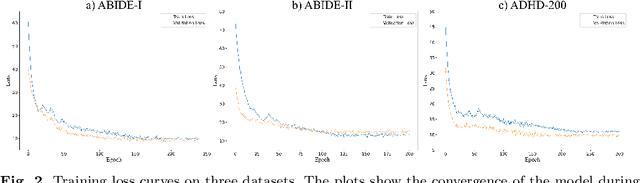
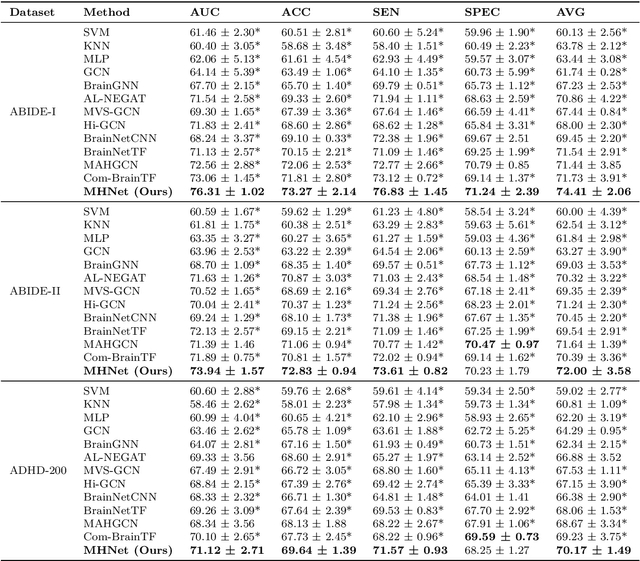
Abstract:Background: Deep learning models have shown promise in diagnosing neurodevelopmental disorders (NDD) like ASD and ADHD. However, many models either use graph neural networks (GNN) to construct single-level brain functional networks (BFNs) or employ spatial convolution filtering for local information extraction from rs-fMRI data, often neglecting high-order features crucial for NDD classification. Methods: We introduce a Multi-view High-order Network (MHNet) to capture hierarchical and high-order features from multi-view BFNs derived from rs-fMRI data for NDD prediction. MHNet has two branches: the Euclidean Space Features Extraction (ESFE) module and the Non-Euclidean Space Features Extraction (Non-ESFE) module, followed by a Feature Fusion-based Classification (FFC) module for NDD identification. ESFE includes a Functional Connectivity Generation (FCG) module and a High-order Convolutional Neural Network (HCNN) module to extract local and high-order features from BFNs in Euclidean space. Non-ESFE comprises a Generic Internet-like Brain Hierarchical Network Generation (G-IBHN-G) module and a High-order Graph Neural Network (HGNN) module to capture topological and high-order features in non-Euclidean space. Results: Experiments on three public datasets show that MHNet outperforms state-of-the-art methods using both AAL1 and Brainnetome Atlas templates. Extensive ablation studies confirm the superiority of MHNet and the effectiveness of using multi-view fMRI information and high-order features. Our study also offers atlas options for constructing more sophisticated hierarchical networks and explains the association between key brain regions and NDD. Conclusion: MHNet leverages multi-view feature learning from both Euclidean and non-Euclidean spaces, incorporating high-order information from BFNs to enhance NDD classification performance.
You Only Acquire Sparse-channel (YOAS): A Unified Framework for Dense-channel EEG Generation
Jun 21, 2024



Abstract:High-precision acquisition of dense-channel electroencephalogram (EEG) signals is often impeded by the costliness and lack of portability of equipment. In contrast, generating dense-channel EEG signals effectively from sparse channels shows promise and economic viability. However, sparse-channel EEG poses challenges such as reduced spatial resolution, information loss, signal mixing, and heightened susceptibility to noise and interference. To address these challenges, we first theoretically formulate the dense-channel EEG generation problem as by optimizing a set of cross-channel EEG signal generation problems. Then, we propose the YOAS framework for generating dense-channel data from sparse-channel EEG signals. The YOAS totally consists of four sequential stages: Data Preparation, Data Preprocessing, Biased-EEG Generation, and Synthetic EEG Generation. Data Preparation and Preprocessing carefully consider the distribution of EEG electrodes and low signal-to-noise ratio problem of EEG signals. Biased-EEG Generation includes sub-modules of BiasEEGGanFormer and BiasEEGDiffFormer, which facilitate long-term feature extraction with attention and generate signals by combining electrode position alignment with diffusion model, respectively. Synthetic EEG Generation synthesizes the final signals, employing a deduction paradigm for multi-channel EEG generation. Extensive experiments confirmed YOAS's feasibility, efficiency, and theoretical validity, even remarkably enhancing data discernibility. This breakthrough in dense-channel EEG signal generation from sparse-channel data opens new avenues for exploration in EEG signal processing and application.
MM-GTUNets: Unified Multi-Modal Graph Deep Learning for Brain Disorders Prediction
Jun 20, 2024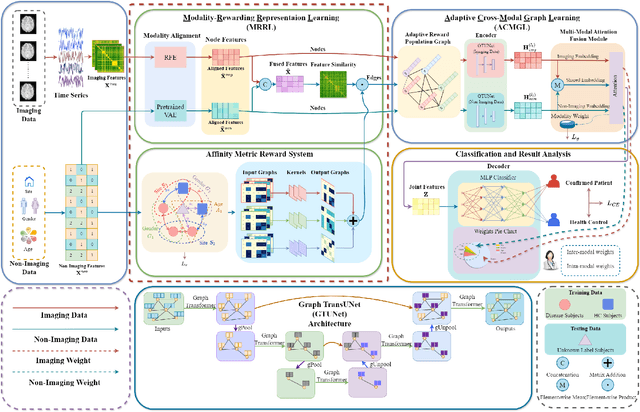

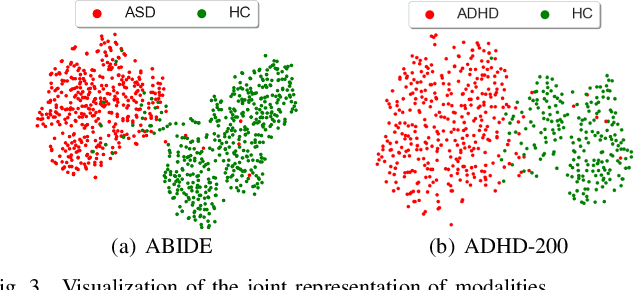
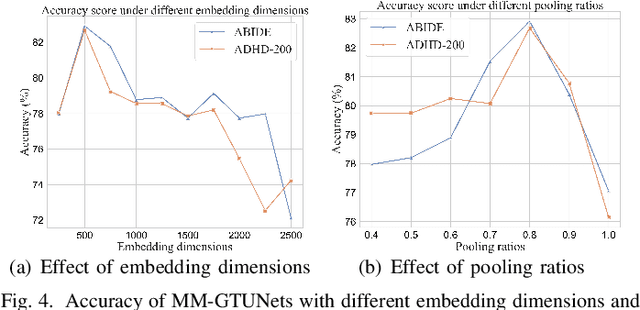
Abstract:Graph deep learning (GDL) has demonstrated impressive performance in predicting population-based brain disorders (BDs) through the integration of both imaging and non-imaging data. However, the effectiveness of GDL based methods heavily depends on the quality of modeling the multi-modal population graphs and tends to degrade as the graph scale increases. Furthermore, these methods often constrain interactions between imaging and non-imaging data to node-edge interactions within the graph, overlooking complex inter-modal correlations, leading to suboptimal outcomes. To overcome these challenges, we propose MM-GTUNets, an end-to-end graph transformer based multi-modal graph deep learning (MMGDL) framework designed for brain disorders prediction at large scale. Specifically, to effectively leverage rich multi-modal information related to diseases, we introduce Modality Reward Representation Learning (MRRL) which adaptively constructs population graphs using a reward system. Additionally, we employ variational autoencoder to reconstruct latent representations of non-imaging features aligned with imaging features. Based on this, we propose Adaptive Cross-Modal Graph Learning (ACMGL), which captures critical modality-specific and modality-shared features through a unified GTUNet encoder taking advantages of Graph UNet and Graph Transformer, and feature fusion module. We validated our method on two public multi-modal datasets ABIDE and ADHD-200, demonstrating its superior performance in diagnosing BDs. Our code is available at https://github.com/NZWANG/MM-GTUNets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge