Haiyun Yuan
"One-Shot" Reduction of Additive Artifacts in Medical Images
Oct 23, 2021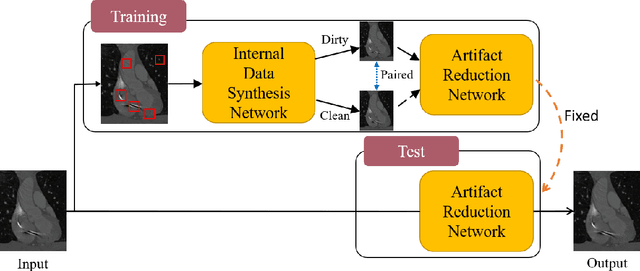
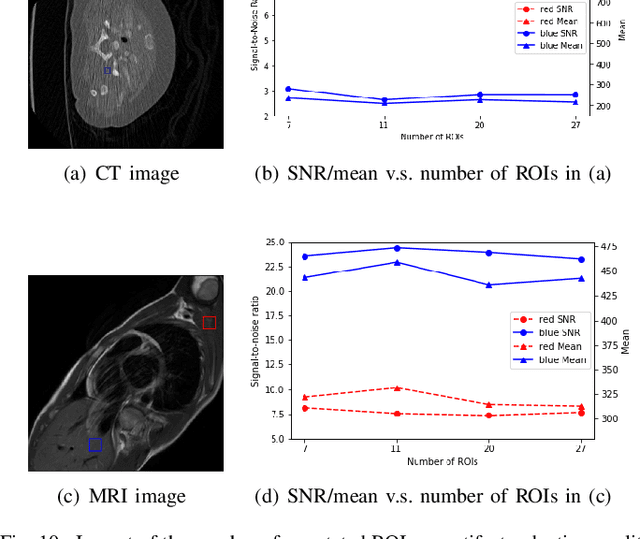
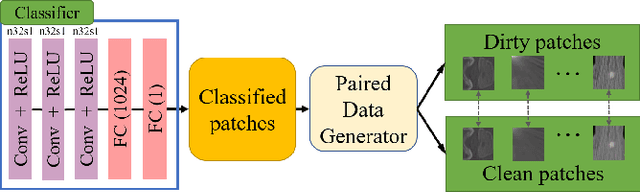

Abstract:Medical images may contain various types of artifacts with different patterns and mixtures, which depend on many factors such as scan setting, machine condition, patients' characteristics, surrounding environment, etc. However, existing deep-learning-based artifact reduction methods are restricted by their training set with specific predetermined artifact types and patterns. As such, they have limited clinical adoption. In this paper, we introduce One-Shot medical image Artifact Reduction (OSAR), which exploits the power of deep learning but without using pre-trained general networks. Specifically, we train a light-weight image-specific artifact reduction network using data synthesized from the input image at test-time. Without requiring any prior large training data set, OSAR can work with almost any medical images that contain varying additive artifacts which are not in any existing data sets. In addition, Computed Tomography (CT) and Magnetic Resonance Imaging (MRI) are used as vehicles and show that the proposed method can reduce artifacts better than state-of-the-art both qualitatively and quantitatively using shorter test time.
Hardware-aware Real-time Myocardial Segmentation Quality Control in Contrast Echocardiography
Sep 14, 2021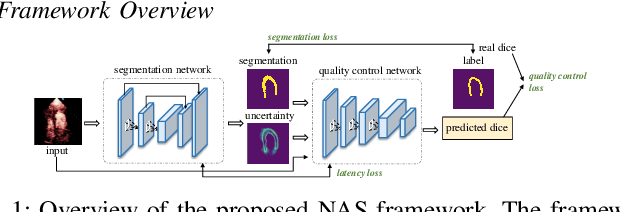
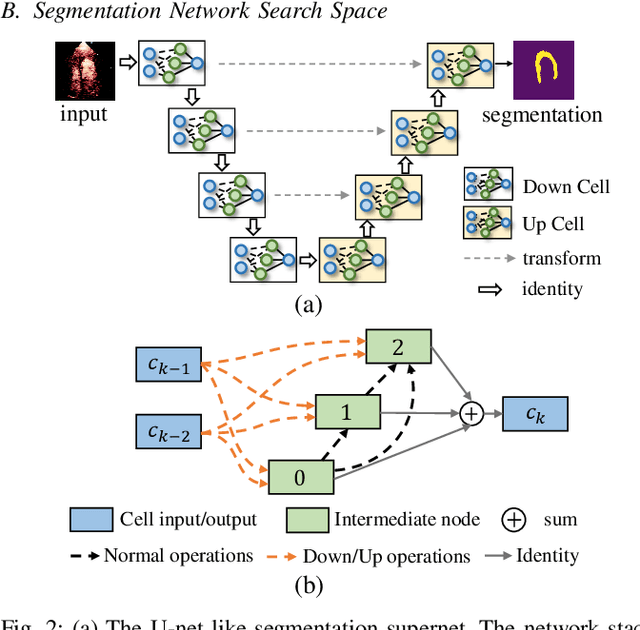
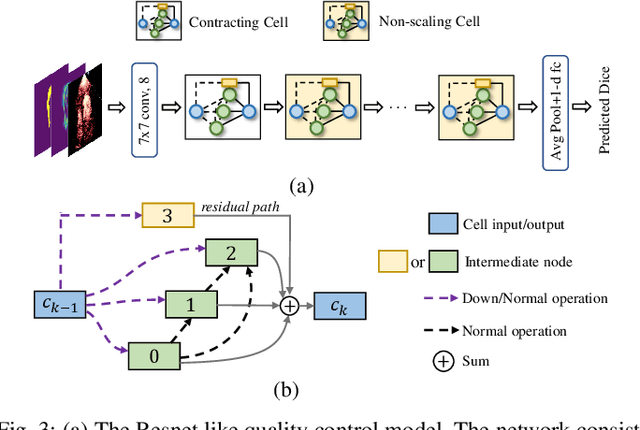
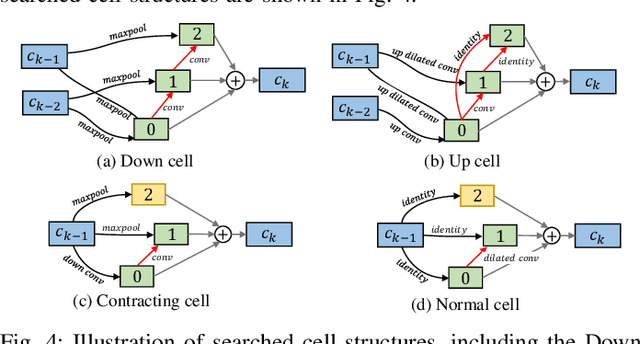
Abstract:Automatic myocardial segmentation of contrast echocardiography has shown great potential in the quantification of myocardial perfusion parameters. Segmentation quality control is an important step to ensure the accuracy of segmentation results for quality research as well as its clinical application. Usually, the segmentation quality control happens after the data acquisition. At the data acquisition time, the operator could not know the quality of the segmentation results. On-the-fly segmentation quality control could help the operator to adjust the ultrasound probe or retake data if the quality is unsatisfied, which can greatly reduce the effort of time-consuming manual correction. However, it is infeasible to deploy state-of-the-art DNN-based models because the segmentation module and quality control module must fit in the limited hardware resource on the ultrasound machine while satisfying strict latency constraints. In this paper, we propose a hardware-aware neural architecture search framework for automatic myocardial segmentation and quality control of contrast echocardiography. We explicitly incorporate the hardware latency as a regularization term into the loss function during training. The proposed method searches the best neural network architecture for the segmentation module and quality prediction module with strict latency.
Positional Contrastive Learning for Volumetric Medical Image Segmentation
Jun 18, 2021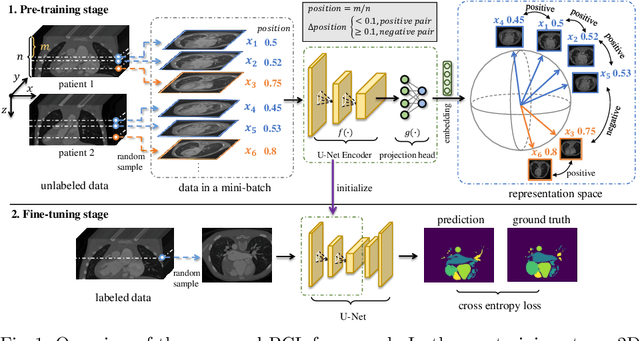
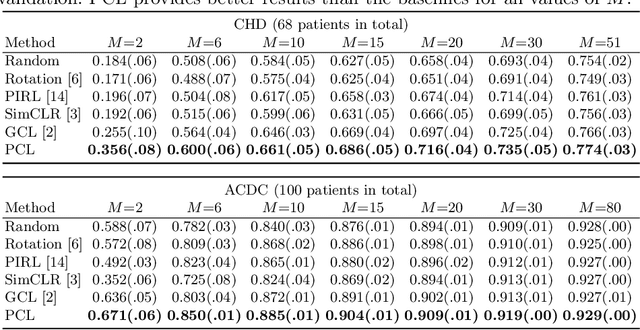
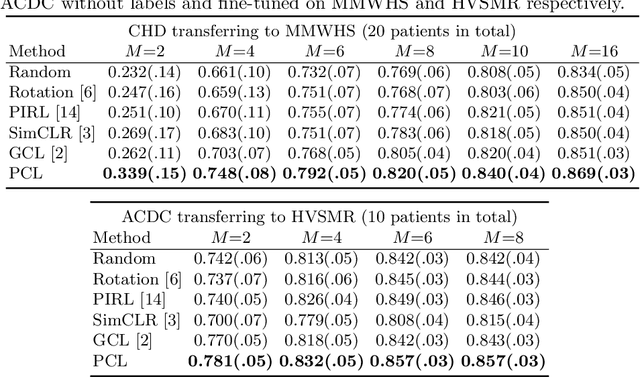
Abstract:The success of deep learning heavily depends on the availability of large labeled training sets. However, it is hard to get large labeled datasets in medical image domain because of the strict privacy concern and costly labeling efforts. Contrastive learning, an unsupervised learning technique, has been proved powerful in learning image-level representations from unlabeled data. The learned encoder can then be transferred or fine-tuned to improve the performance of downstream tasks with limited labels. A critical step in contrastive learning is the generation of contrastive data pairs, which is relatively simple for natural image classification but quite challenging for medical image segmentation due to the existence of the same tissue or organ across the dataset. As a result, when applied to medical image segmentation, most state-of-the-art contrastive learning frameworks inevitably introduce a lot of false-negative pairs and result in degraded segmentation quality. To address this issue, we propose a novel positional contrastive learning (PCL) framework to generate contrastive data pairs by leveraging the position information in volumetric medical images. Experimental results on CT and MRI datasets demonstrate that the proposed PCL method can substantially improve the segmentation performance compared to existing methods in both semi-supervised setting and transfer learning setting.
Multi-Cycle-Consistent Adversarial Networks for Edge Denoising of Computed Tomography Images
Apr 25, 2021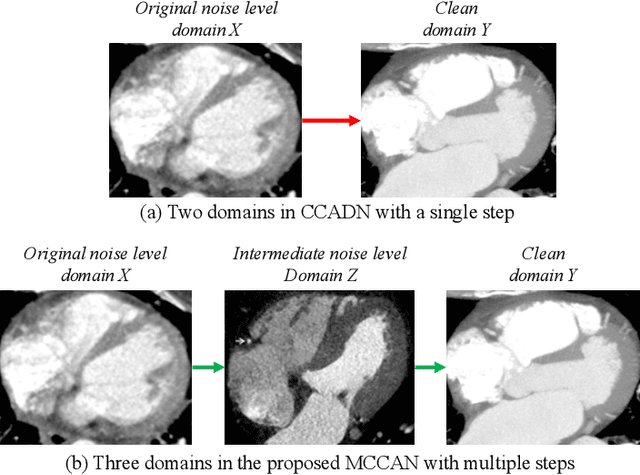
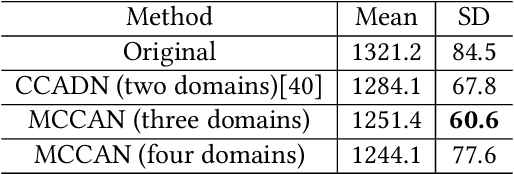

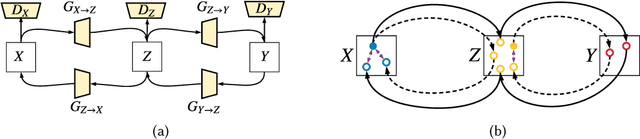
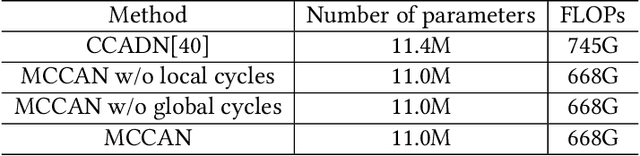
Abstract:As one of the most commonly ordered imaging tests, computed tomography (CT) scan comes with inevitable radiation exposure that increases the cancer risk to patients. However, CT image quality is directly related to radiation dose, thus it is desirable to obtain high-quality CT images with as little dose as possible. CT image denoising tries to obtain high dose like high-quality CT images (domain X) from low dose low-quality CTimages (domain Y), which can be treated as an image-to-image translation task where the goal is to learn the transform between a source domain X (noisy images) and a target domain Y (clean images). In this paper, we propose a multi-cycle-consistent adversarial network (MCCAN) that builds intermediate domains and enforces both local and global cycle-consistency for edge denoising of CT images. The global cycle-consistency couples all generators together to model the whole denoising process, while the local cycle-consistency imposes effective supervision on the process between adjacent domains. Experiments show that both local and global cycle-consistency are important for the success of MCCAN, which outperformsCCADN in terms of denoising quality with slightly less computation resource consumption.
ImageCHD: A 3D Computed Tomography Image Dataset for Classification of Congenital Heart Disease
Jan 26, 2021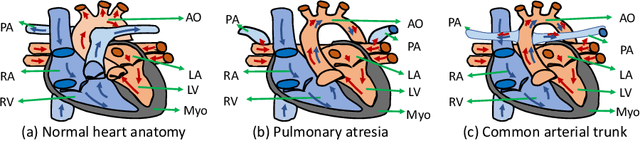
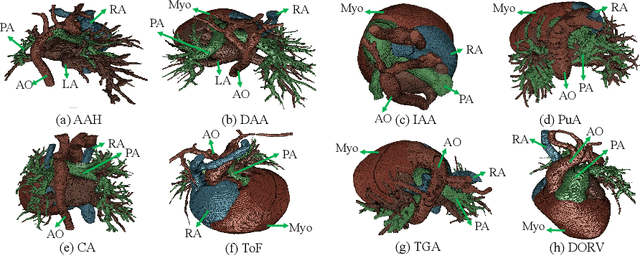
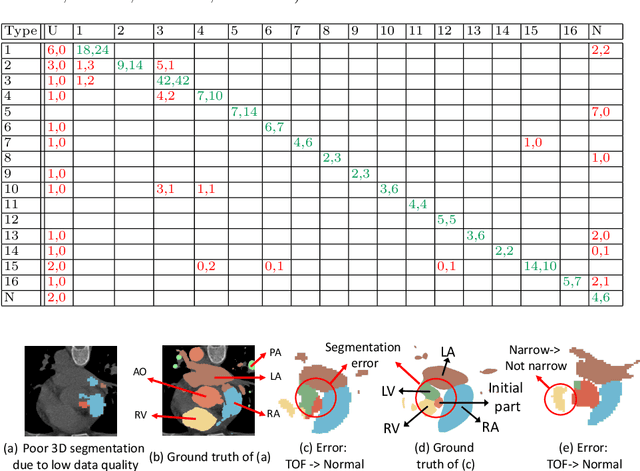
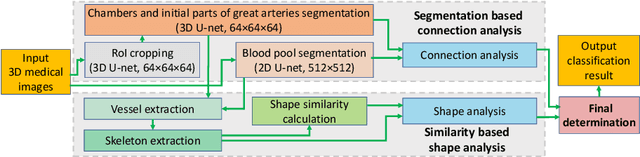
Abstract:Congenital heart disease (CHD) is the most common type of birth defect, which occurs 1 in every 110 births in the United States. CHD usually comes with severe variations in heart structure and great artery connections that can be classified into many types. Thus highly specialized domain knowledge and the time-consuming human process is needed to analyze the associated medical images. On the other hand, due to the complexity of CHD and the lack of dataset, little has been explored on the automatic diagnosis (classification) of CHDs. In this paper, we present ImageCHD, the first medical image dataset for CHD classification. ImageCHD contains 110 3D Computed Tomography (CT) images covering most types of CHD, which is of decent size Classification of CHDs requires the identification of large structural changes without any local tissue changes, with limited data. It is an example of a larger class of problems that are quite difficult for current machine-learning-based vision methods to solve. To demonstrate this, we further present a baseline framework for the automatic classification of CHD, based on a state-of-the-art CHD segmentation method. Experimental results show that the baseline framework can only achieve a classification accuracy of 82.0\% under a selective prediction scheme with 88.4\% coverage, leaving big room for further improvement. We hope that ImageCHD can stimulate further research and lead to innovative and generic solutions that would have an impact in multiple domains. Our dataset is released to the public compared with existing medical imaging datasets.
Myocardial Segmentation of Cardiac MRI Sequences with Temporal Consistency for Coronary Artery Disease Diagnosis
Dec 29, 2020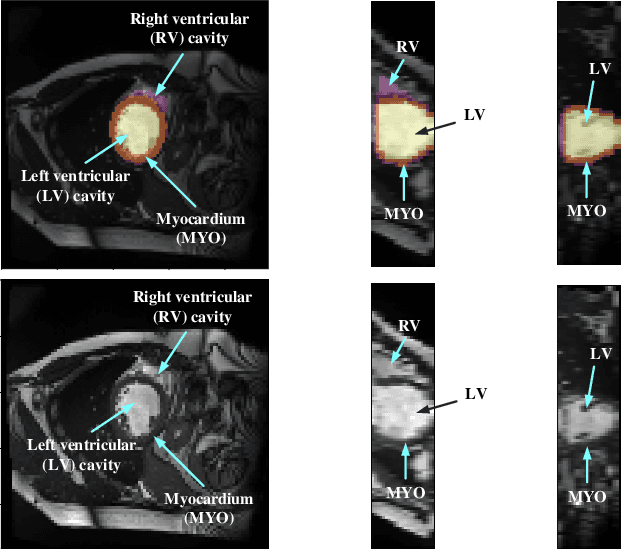
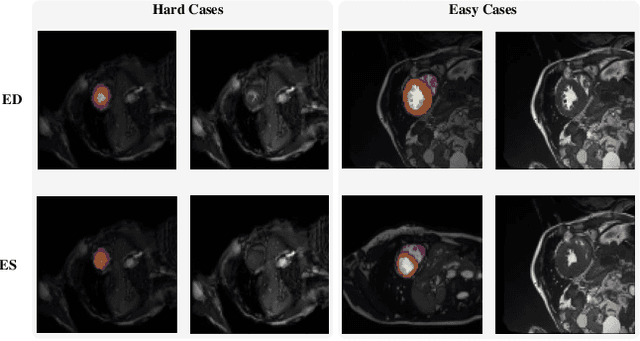
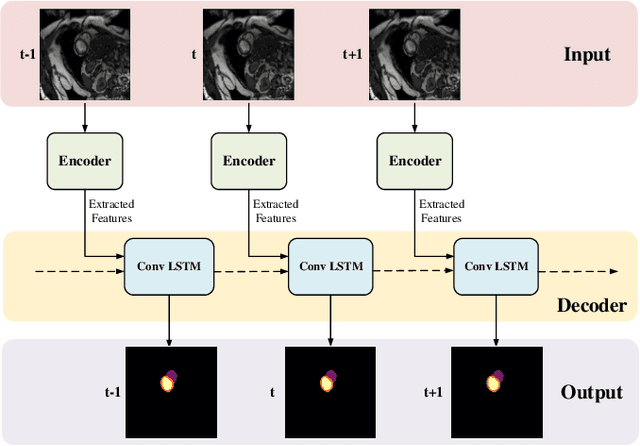
Abstract:Coronary artery disease (CAD) is the most common cause of death globally, and its diagnosis is usually based on manual myocardial segmentation of Magnetic Resonance Imaging (MRI) sequences. As the manual segmentation is tedious, time-consuming and with low applicability, automatic myocardial segmentation using machine learning techniques has been widely explored recently. However, almost all the existing methods treat the input MRI sequences independently, which fails to capture the temporal information between sequences, e.g., the shape and location information of the myocardium in sequences along time. In this paper, we propose a myocardial segmentation framework for sequence of cardiac MRI (CMR) scanning images of left ventricular cavity, right ventricular cavity, and myocardium. Specifically, we propose to combine conventional networks and recurrent networks to incorporate temporal information between sequences to ensure temporal consistent. We evaluated our framework on the Automated Cardiac Diagnosis Challenge (ACDC) dataset. Experiment results demonstrate that our framework can improve the segmentation accuracy by up to 2% in Dice coefficient.
Towards Cardiac Intervention Assistance: Hardware-aware Neural Architecture Exploration for Real-Time 3D Cardiac Cine MRI Segmentation
Aug 17, 2020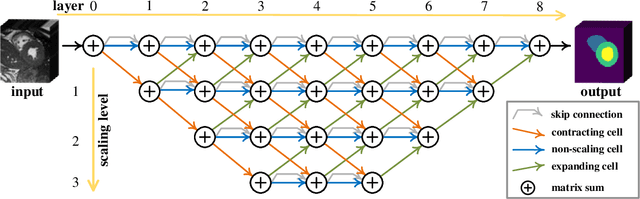
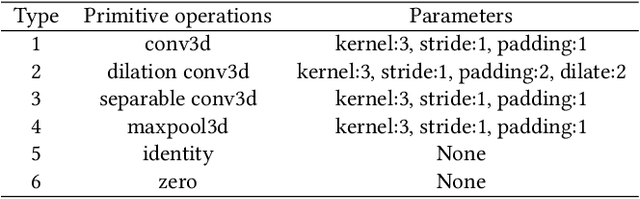
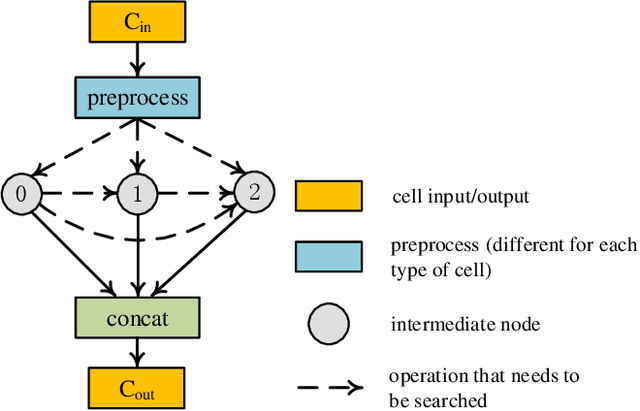
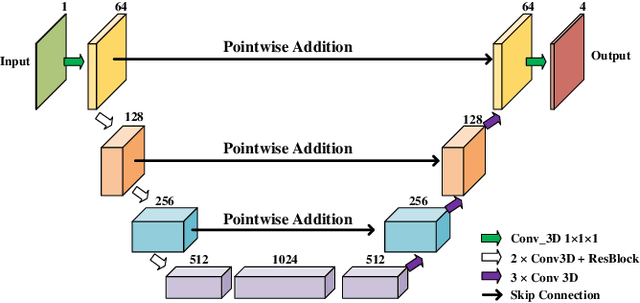
Abstract:Real-time cardiac magnetic resonance imaging (MRI) plays an increasingly important role in guiding various cardiac interventions. In order to provide better visual assistance, the cine MRI frames need to be segmented on-the-fly to avoid noticeable visual lag. In addition, considering reliability and patient data privacy, the computation is preferably done on local hardware. State-of-the-art MRI segmentation methods mostly focus on accuracy only, and can hardly be adopted for real-time application or on local hardware. In this work, we present the first hardware-aware multi-scale neural architecture search (NAS) framework for real-time 3D cardiac cine MRI segmentation. The proposed framework incorporates a latency regularization term into the loss function to handle real-time constraints, with the consideration of underlying hardware. In addition, the formulation is fully differentiable with respect to the architecture parameters, so that stochastic gradient descent (SGD) can be used for optimization to reduce the computation cost while maintaining optimization quality. Experimental results on ACDC MICCAI 2017 dataset demonstrate that our hardware-aware multi-scale NAS framework can reduce the latency by up to 3.5 times and satisfy the real-time constraints, while still achieving competitive segmentation accuracy, compared with the state-of-the-art NAS segmentation framework.
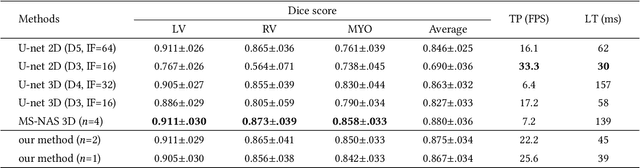
ICA-UNet: ICA Inspired Statistical UNet for Real-time 3D Cardiac Cine MRI Segmentation
Jul 18, 2020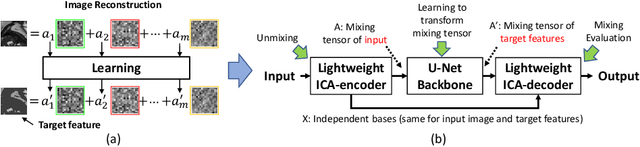
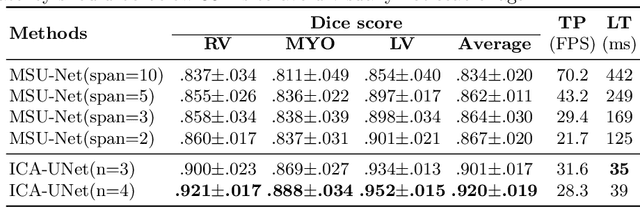
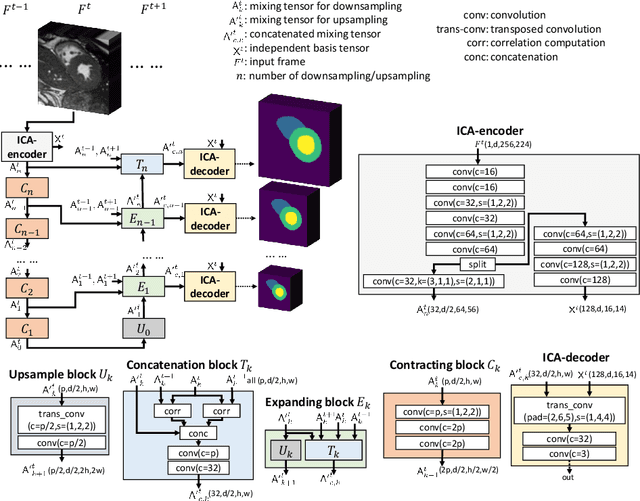
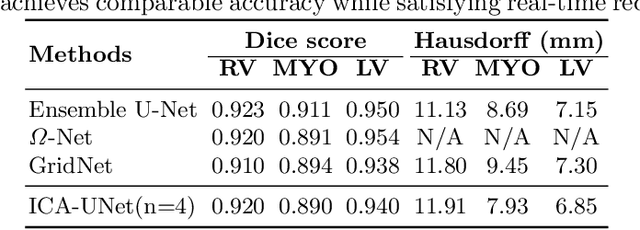
Abstract:Real-time cine magnetic resonance imaging (MRI) plays an increasingly important role in various cardiac interventions. In order to enable fast and accurate visual assistance, the temporal frames need to be segmented on-the-fly. However, state-of-the-art MRI segmentation methods are used either offline because of their high computation complexity, or in real-time but with significant accuracy loss and latency increase (causing visually noticeable lag). As such, they can hardly be adopted to assist visual guidance. In this work, inspired by a new interpretation of Independent Component Analysis (ICA) for learning, we propose a novel ICA-UNet for real-time 3D cardiac cine MRI segmentation. Experiments using the MICCAI ACDC 2017 dataset show that, compared with the state-of-the-arts, ICA-UNet not only achieves higher Dice scores, but also meets the real-time requirements for both throughput and latency (up to 12.6X reduction), enabling real-time guidance for cardiac interventions without visual lag.
MSU-Net: Multiscale Statistical U-Net for Real-time 3D Cardiac MRI Video Segmentation
Sep 15, 2019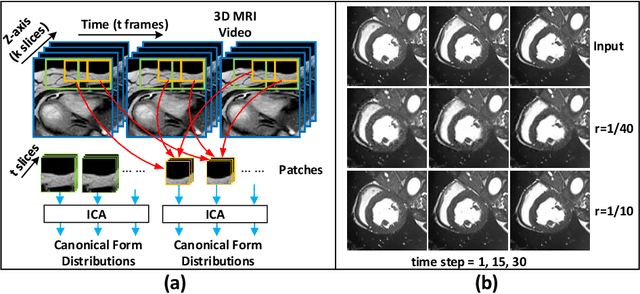
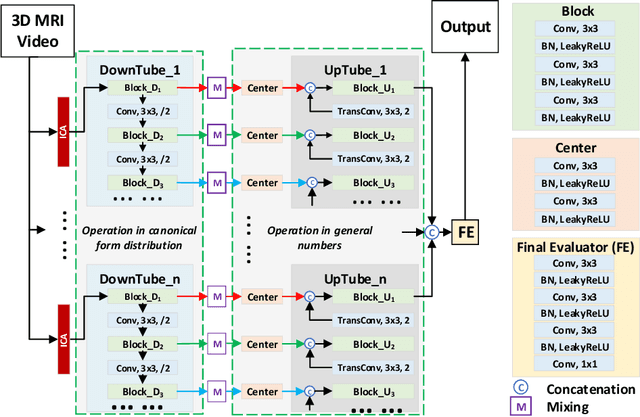
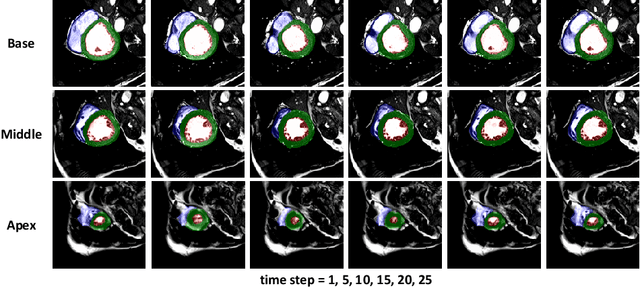
Abstract:Cardiac magnetic resonance imaging (MRI) is an essential tool for MRI-guided surgery and real-time intervention. The MRI videos are expected to be segmented on-the-fly in real practice. However, existing segmentation methods would suffer from drastic accuracy loss when modified for speedup. In this work, we propose Multiscale Statistical U-Net (MSU-Net) for real-time 3D MRI video segmentation in cardiac surgical guidance. Our idea is to model the input samples as multiscale canonical form distributions for speedup, while the spatio-temporal correlation is still fully utilized. A parallel statistical U-Net is then designed to efficiently process these distributions. The fast data sampling and efficient parallel structure of MSU-Net endorse the fast and accurate inference. Compared with vanilla U-Net and a modified state-of-the-art method GridNet, our method achieves up to 268% and 237% speedup with 1.6% and 3.6% increased Dice scores.
Accurate Congenital Heart Disease Model Generation for 3D Printing
Jul 12, 2019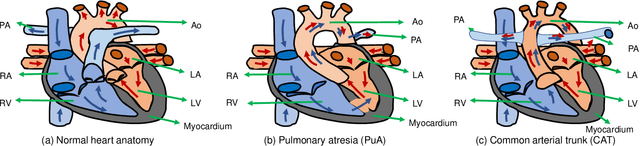
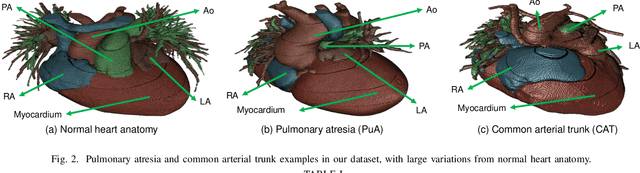
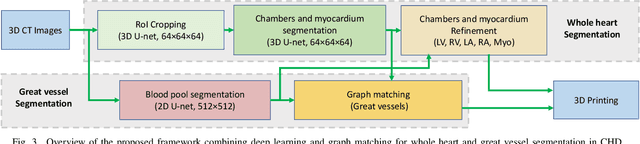
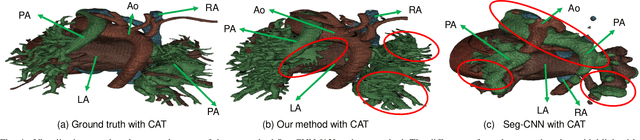
Abstract:3D printing has been widely adopted for clinical decision making and interventional planning of Congenital heart disease (CHD), while whole heart and great vessel segmentation is the most significant but time-consuming step in the model generation for 3D printing. While various automatic whole heart and great vessel segmentation frameworks have been developed in the literature, they are ineffective when applied to medical images in CHD, which have significant variations in heart structure and great vessel connections. To address the challenge, we leverage the power of deep learning in processing regular structures and that of graph algorithms in dealing with large variations and propose a framework that combines both for whole heart and great vessel segmentation in CHD. Particularly, we first use deep learning to segment the four chambers and myocardium followed by the blood pool, where variations are usually small. We then extract the connection information and apply graph matching to determine the categories of all the vessels. Experimental results using 683D CT images covering 14 types of CHD show that our method can increase Dice score by 11.9% on average compared with the state-of-the-art whole heart and great vessel segmentation method in normal anatomy. The segmentation results are also printed out using 3D printers for validation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge