Eloy Geenjaar
Mapping minds not averages: a scalable subject-specific manifold learning framework for neuroimaging data
Apr 30, 2025Abstract:Mental and cognitive representations are believed to reside on low-dimensional, non-linear manifolds embedded within high-dimensional brain activity. Uncovering these manifolds is key to understanding individual differences in brain function, yet most existing machine learning methods either rely on population-level spatial alignment or assume data that is temporally structured, either because data is aligned among subjects or because event timings are known. We introduce a manifold learning framework that can capture subject-specific spatial variations across both structured and temporally unstructured neuroimaging data. On simulated data and two naturalistic fMRI datasets (Sherlock and Forrest Gump), our framework outperforms group-based baselines by recovering more accurate and individualized representations. We further show that the framework scales efficiently to large datasets and generalizes well to new subjects. To test this, we apply the framework to temporally unstructured resting-state fMRI data from individuals with schizophrenia and healthy controls. We further apply our method to a large resting-state fMRI dataset comprising individuals with schizophrenia and controls. In this setting, we demonstrate that the framework scales efficiently to large populations and generalizes robustly to unseen subjects. The learned subject-specific spatial maps our model finds reveal clinically relevant patterns, including increased activation in the basal ganglia, visual, auditory, and somatosensory regions, and decreased activation in the insula, inferior frontal gyrus, and angular gyrus. These findings suggest that our framework can uncover clinically relevant subject-specific brain activity patterns. Our approach thus provides a scalable and individualized framework for modeling brain activity, with applications in computational neuroscience and clinical research.
CiTrus: Squeezing Extra Performance out of Low-data Bio-signal Transfer Learning
Dec 16, 2024



Abstract:Transfer learning for bio-signals has recently become an important technique to improve prediction performance on downstream tasks with small bio-signal datasets. Recent works have shown that pre-training a neural network model on a large dataset (e.g. EEG) with a self-supervised task, replacing the self-supervised head with a linear classification head, and fine-tuning the model on different downstream bio-signal datasets (e.g., EMG or ECG) can dramatically improve the performance on those datasets. In this paper, we propose a new convolution-transformer hybrid model architecture with masked auto-encoding for low-data bio-signal transfer learning, introduce a frequency-based masked auto-encoding task, employ a more comprehensive evaluation framework, and evaluate how much and when (multimodal) pre-training improves fine-tuning performance. We also introduce a dramatically more performant method of aligning a downstream dataset with a different temporal length and sampling rate to the original pre-training dataset. Our findings indicate that the convolution-only part of our hybrid model can achieve state-of-the-art performance on some low-data downstream tasks. The performance is often improved even further with our full model. In the case of transformer-based models we find that pre-training especially improves performance on downstream datasets, multimodal pre-training often increases those gains further, and our frequency-based pre-training performs the best on average for the lowest and highest data regimes.
Learning low-dimensional dynamics from whole-brain data improves task capture
May 18, 2023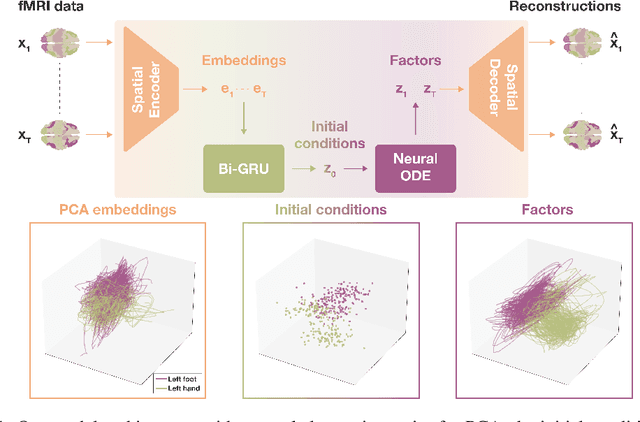
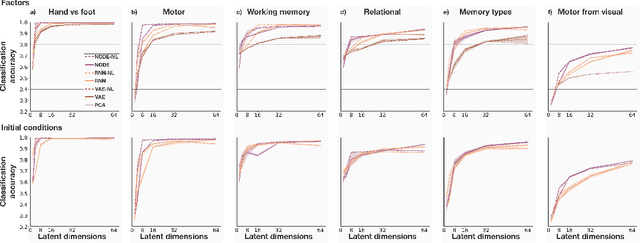
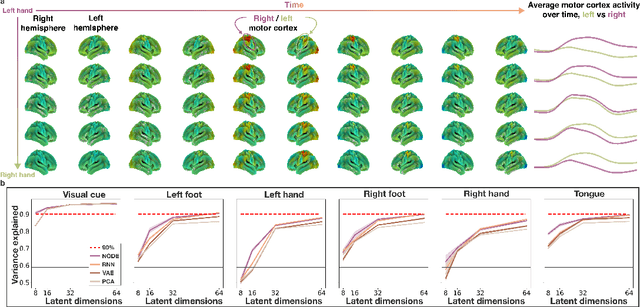
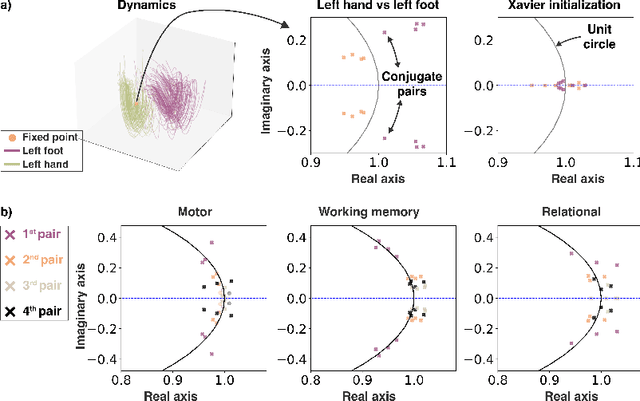
Abstract:The neural dynamics underlying brain activity are critical to understanding cognitive processes and mental disorders. However, current voxel-based whole-brain dimensionality reduction techniques fall short of capturing these dynamics, producing latent timeseries that inadequately relate to behavioral tasks. To address this issue, we introduce a novel approach to learning low-dimensional approximations of neural dynamics by using a sequential variational autoencoder (SVAE) that represents the latent dynamical system via a neural ordinary differential equation (NODE). Importantly, our method finds smooth dynamics that can predict cognitive processes with accuracy higher than classical methods. Our method also shows improved spatial localization to task-relevant brain regions and identifies well-known structures such as the motor homunculus from fMRI motor task recordings. We also find that non-linear projections to the latent space enhance performance for specific tasks, offering a promising direction for future research. We evaluate our approach on various task-fMRI datasets, including motor, working memory, and relational processing tasks, and demonstrate that it outperforms widely used dimensionality reduction techniques in how well the latent timeseries relates to behavioral sub-tasks, such as left-hand or right-hand tapping. Additionally, we replace the NODE with a recurrent neural network (RNN) and compare the two approaches to understand the importance of explicitly learning a dynamical system. Lastly, we analyze the robustness of the learned dynamical systems themselves and find that their fixed points are robust across seeds, highlighting our method's potential for the analysis of cognitive processes as dynamical systems.
CommsVAE: Learning the brain's macroscale communication dynamics using coupled sequential VAEs
Oct 07, 2022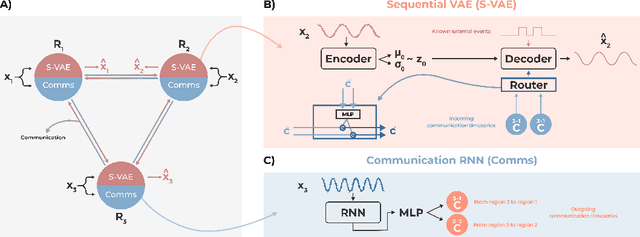
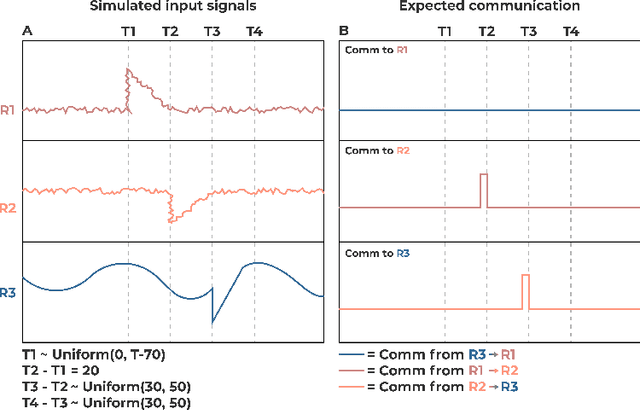
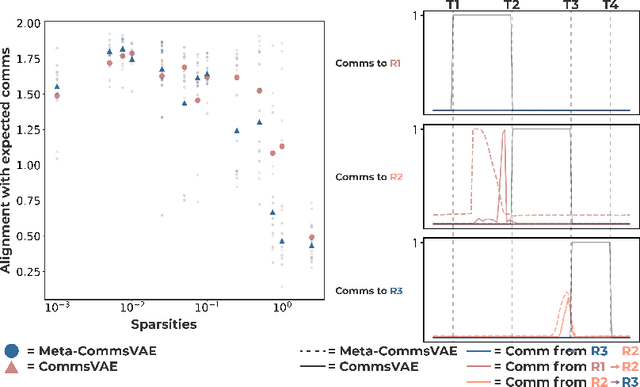
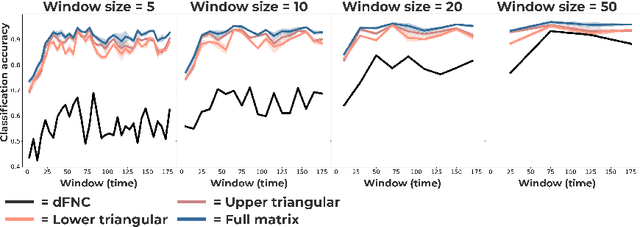
Abstract:Communication within or between complex systems is commonplace in the natural sciences and fields such as graph neural networks. The brain is a perfect example of such a complex system, where communication between brain regions is constantly being orchestrated. To analyze communication, the brain is often split up into anatomical regions that each perform certain computations. These regions must interact and communicate with each other to perform tasks and support higher-level cognition. On a macroscale, these regions communicate through signal propagation along the cortex and along white matter tracts over longer distances. When and what types of signals are communicated over time is an unsolved problem and is often studied using either functional or structural data. In this paper, we propose a non-linear generative approach to communication from functional data. We address three issues with common connectivity approaches by explicitly modeling the directionality of communication, finding communication at each timestep, and encouraging sparsity. To evaluate our model, we simulate temporal data that has sparse communication between nodes embedded in it and show that our model can uncover the expected communication dynamics. Subsequently, we apply our model to temporal neural data from multiple tasks and show that our approach models communication that is more specific to each task. The specificity of our method means it can have an impact on the understanding of psychiatric disorders, which are believed to be related to highly specific communication between brain regions compared to controls. In sum, we propose a general model for dynamic communication learning on graphs, and show its applicability to a subfield of the natural sciences, with potential widespread scientific impact.
Self-supervised multimodal neuroimaging yields predictive representations for a spectrum of Alzheimer's phenotypes
Sep 07, 2022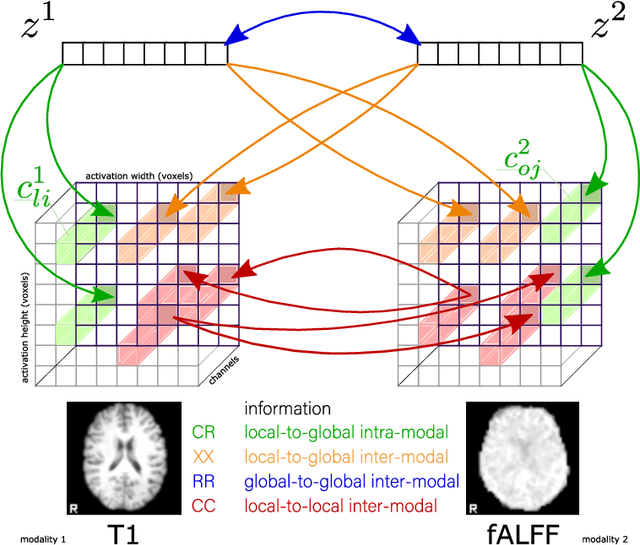
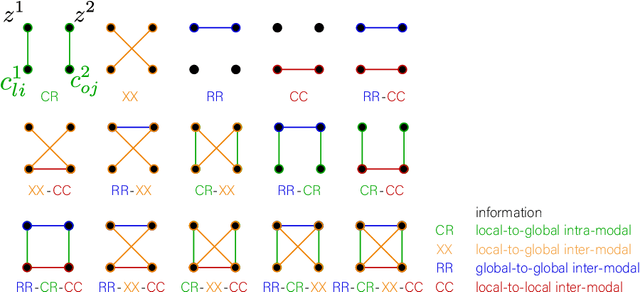
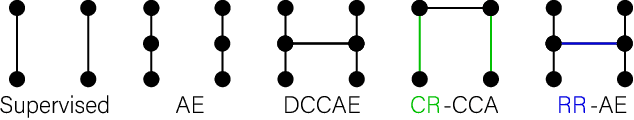
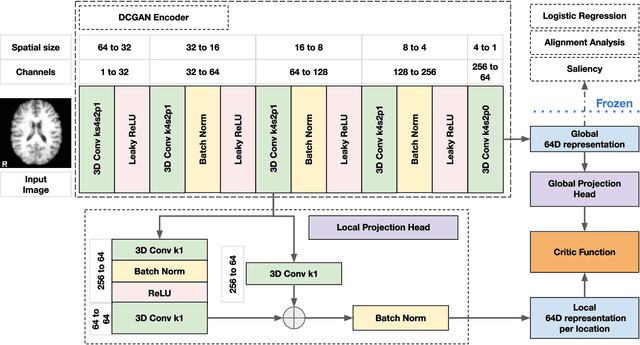
Abstract:Recent neuroimaging studies that focus on predicting brain disorders via modern machine learning approaches commonly include a single modality and rely on supervised over-parameterized models.However, a single modality provides only a limited view of the highly complex brain. Critically, supervised models in clinical settings lack accurate diagnostic labels for training. Coarse labels do not capture the long-tailed spectrum of brain disorder phenotypes, which leads to a loss of generalizability of the model that makes them less useful in diagnostic settings. This work presents a novel multi-scale coordinated framework for learning multiple representations from multimodal neuroimaging data. We propose a general taxonomy of informative inductive biases to capture unique and joint information in multimodal self-supervised fusion. The taxonomy forms a family of decoder-free models with reduced computational complexity and a propensity to capture multi-scale relationships between local and global representations of the multimodal inputs. We conduct a comprehensive evaluation of the taxonomy using functional and structural magnetic resonance imaging (MRI) data across a spectrum of Alzheimer's disease phenotypes and show that self-supervised models reveal disorder-relevant brain regions and multimodal links without access to the labels during pre-training. The proposed multimodal self-supervised learning yields representations with improved classification performance for both modalities. The concomitant rich and flexible unsupervised deep learning framework captures complex multimodal relationships and provides predictive performance that meets or exceeds that of a more narrow supervised classification analysis. We present elaborate quantitative evidence of how this framework can significantly advance our search for missing links in complex brain disorders.
Spatio-temporally separable non-linear latent factor learning: an application to somatomotor cortex fMRI data
May 26, 2022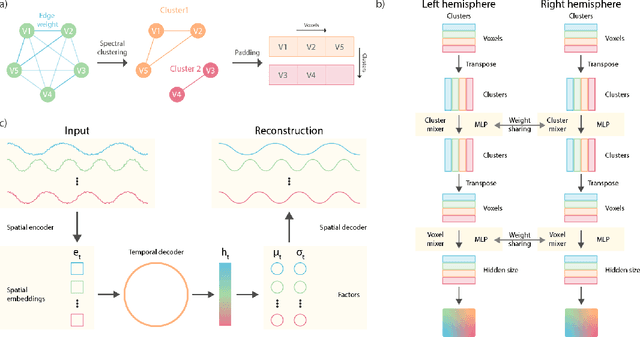
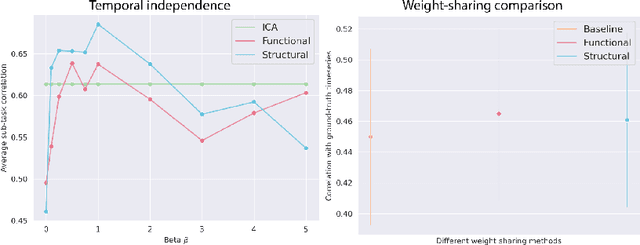
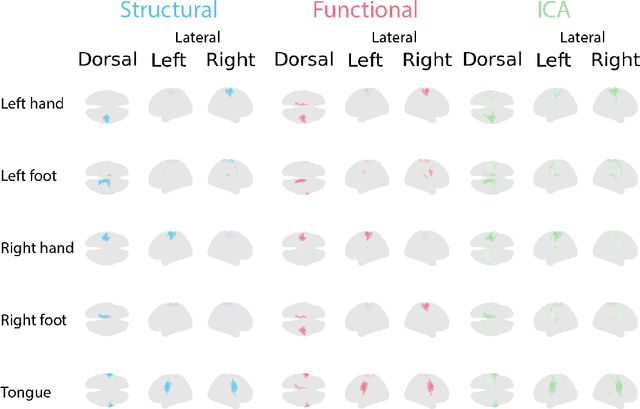
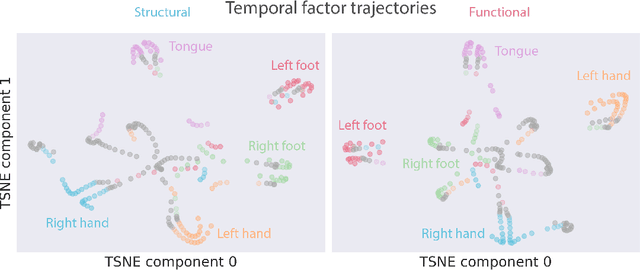
Abstract:Functional magnetic resonance imaging (fMRI) data contain complex spatiotemporal dynamics, thus researchers have developed approaches that reduce the dimensionality of the signal while extracting relevant and interpretable dynamics. Models of fMRI data that can perform whole-brain discovery of dynamical latent factors are understudied. The benefits of approaches such as linear independent component analysis models have been widely appreciated, however, nonlinear extensions of these models present challenges in terms of identification. Deep learning methods provide a way forward, but new methods for efficient spatial weight-sharing are critical to deal with the high dimensionality of the data and the presence of noise. Our approach generalizes weight sharing to non-Euclidean neuroimaging data by first performing spectral clustering based on the structural and functional similarity between voxels. The spectral clusters and their assignments can then be used as patches in an adapted multi-layer perceptron (MLP)-mixer model to share parameters among input points. To encourage temporally independent latent factors, we use an additional total correlation term in the loss. Our approach is evaluated on data with multiple motor sub-tasks to assess whether the model captures disentangled latent factors that correspond to each sub-task. Then, to assess the latent factors we find further, we compare the spatial location of each latent factor to the motor homunculus. Finally, we show that our approach captures task effects better than the current gold standard of source signal separation, independent component analysis (ICA).
Variational voxelwise rs-fMRI representation learning: Evaluation of sex, age, and neuropsychiatric signatures
Aug 29, 2021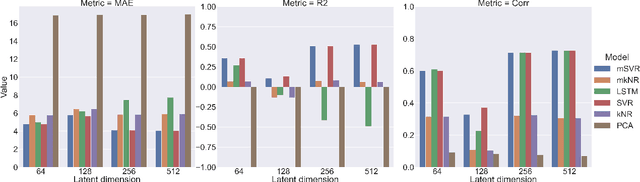
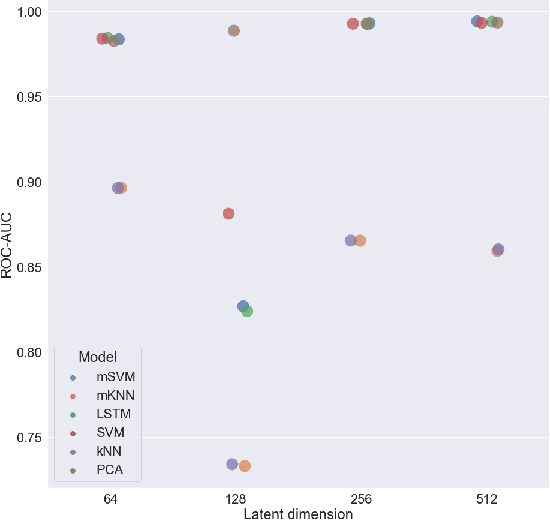
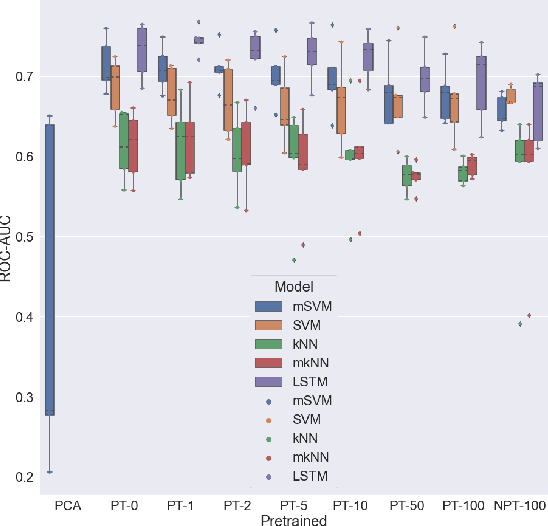
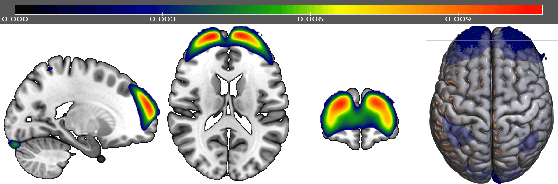
Abstract:We propose to apply non-linear representation learning to voxelwise rs-fMRI data. Learning the non-linear representations is done using a variational autoencoder (VAE). The VAE is trained on voxelwise rs-fMRI data and performs non-linear dimensionality reduction that retains meaningful information. The retention of information in the model's representations is evaluated using downstream age regression and sex classification tasks. The results on these tasks are highly encouraging and a linear regressor trained with the representations of our unsupervised model performs almost as well as a supervised neural network, trained specifically for age regression on the same dataset. The model is also evaluated with a schizophrenia diagnosis prediction task, to assess its feasibility as a dimensionality reduction method for neuropsychiatric datasets. These results highlight the potential for pre-training on a larger set of individuals who do not have mental illness, to improve the downstream neuropsychiatric task results. The pre-trained model is fine-tuned for a variable number of epochs on a schizophrenia dataset and we find that fine-tuning for 1 epoch yields the best results. This work therefore not only opens up non-linear dimensionality reduction for voxelwise rs-fMRI data but also shows that pre-training a deep learning model on voxelwise rs-fMRI datasets greatly increases performance even on smaller datasets. It also opens up the ability to look at the distribution of rs-fMRI time series in the latent space of the VAE for heterogeneous neuropsychiatric disorders like schizophrenia in future work. This can be complemented with the generative aspect of the model that allows us to reconstruct points from the model's latent space back into brain space and obtain an improved understanding of the relation that the VAE learns between subjects, timepoints, and a subject's characteristics.
Fusing multimodal neuroimaging data with a variational autoencoder
May 03, 2021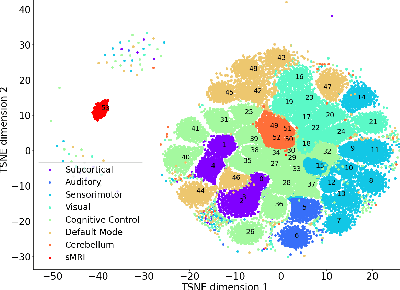
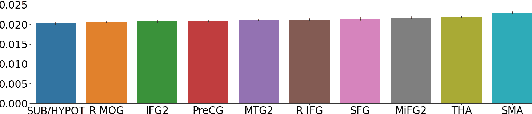
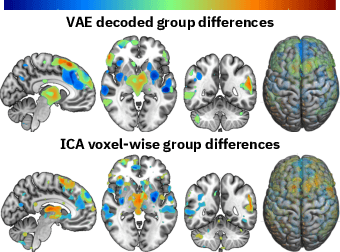
Abstract:Neuroimaging studies often involve the collection of multiple data modalities. These modalities contain both shared and mutually exclusive information about the brain. This work aims at finding a scalable and interpretable method to fuse the information of multiple neuroimaging modalities using a variational autoencoder (VAE). To provide an initial assessment, this work evaluates the representations that are learned using a schizophrenia classification task. A support vector machine trained on the representations achieves an area under the curve for the classifier's receiver operating characteristic (ROC-AUC) of 0.8610.
Tasting the cake: evaluating self-supervised generalization on out-of-distribution multimodal MRI data
Apr 20, 2021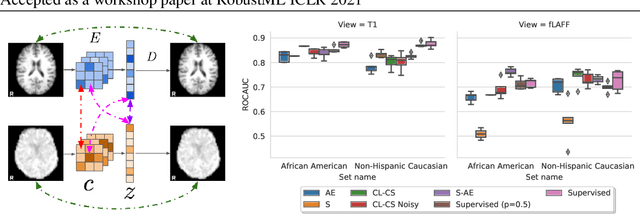
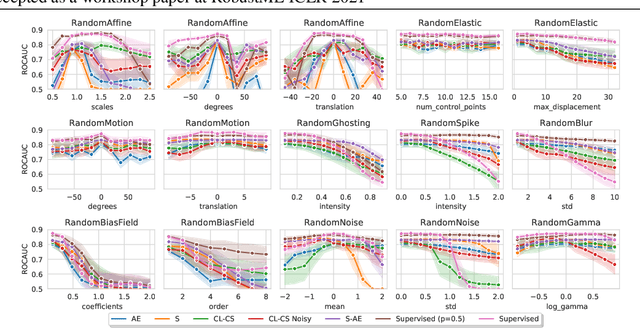
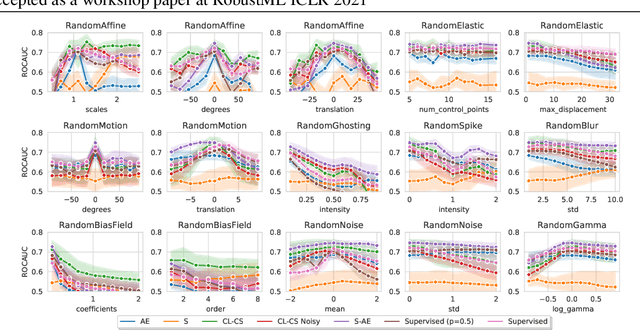

Abstract:Self-supervised learning has enabled significant improvements on natural image benchmarks. However, there is less work in the medical imaging domain in this area. The optimal models have not yet been determined among the various options. Moreover, little work has evaluated the current applicability limits of novel self-supervised methods. In this paper, we evaluate a range of current contrastive self-supervised methods on out-of-distribution generalization in order to evaluate their applicability to medical imaging. We show that self-supervised models are not as robust as expected based on their results in natural imaging benchmarks and can be outperformed by supervised learning with dropout. We also show that this behavior can be countered with extensive augmentation. Our results highlight the need for out-of-distribution generalization standards and benchmarks to adopt the self-supervised methods in the medical imaging community.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge