Dominic Masters
Skillful joint probabilistic weather forecasting from marginals
Jun 12, 2025
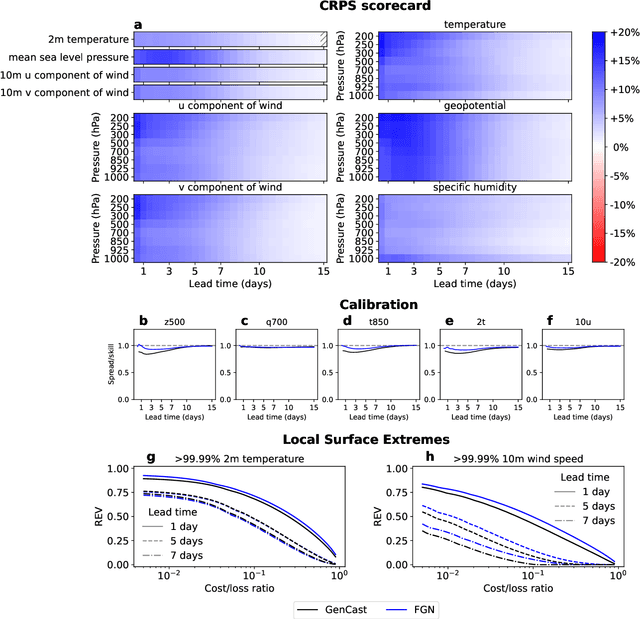
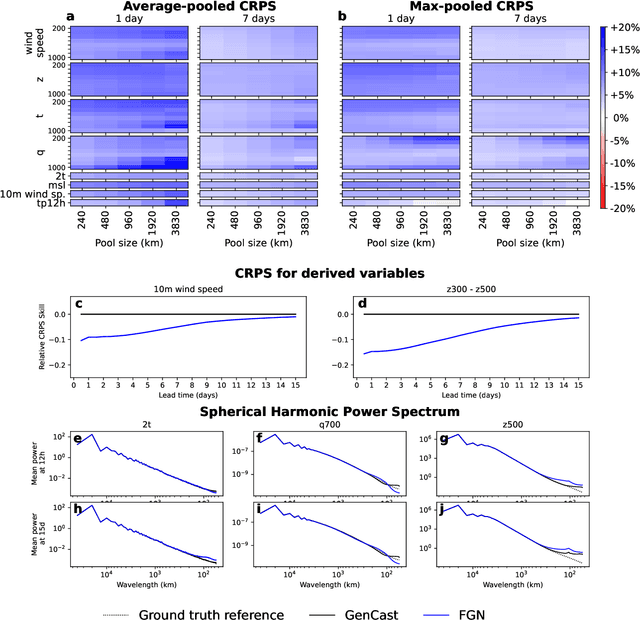
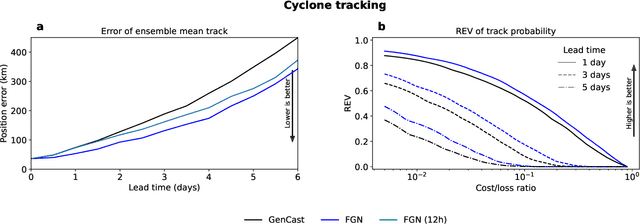
Abstract:Machine learning (ML)-based weather models have rapidly risen to prominence due to their greater accuracy and speed than traditional forecasts based on numerical weather prediction (NWP), recently outperforming traditional ensembles in global probabilistic weather forecasting. This paper presents FGN, a simple, scalable and flexible modeling approach which significantly outperforms the current state-of-the-art models. FGN generates ensembles via learned model-perturbations with an ensemble of appropriately constrained models. It is trained directly to minimize the continuous rank probability score (CRPS) of per-location forecasts. It produces state-of-the-art ensemble forecasts as measured by a range of deterministic and probabilistic metrics, makes skillful ensemble tropical cyclone track predictions, and captures joint spatial structure despite being trained only on marginals.
Generating QM1B with PySCF$_{\text{IPU}}$
Nov 02, 2023Abstract:The emergence of foundation models in Computer Vision and Natural Language Processing have resulted in immense progress on downstream tasks. This progress was enabled by datasets with billions of training examples. Similar benefits are yet to be unlocked for quantum chemistry, where the potential of deep learning is constrained by comparatively small datasets with 100k to 20M training examples. These datasets are limited in size because the labels are computed using the accurate (but computationally demanding) predictions of Density Functional Theory (DFT). Notably, prior DFT datasets were created using CPU supercomputers without leveraging hardware acceleration. In this paper, we take a first step towards utilising hardware accelerators by introducing the data generator PySCF$_{\text{IPU}}$ using Intelligence Processing Units (IPUs). This allowed us to create the dataset QM1B with one billion training examples containing 9-11 heavy atoms. We demonstrate that a simple baseline neural network (SchNet 9M) improves its performance by simply increasing the amount of training data without additional inductive biases. To encourage future researchers to use QM1B responsibly, we highlight several limitations of QM1B and emphasise the low-resolution of our DFT options, which also serves as motivation for even larger, more accurate datasets. Code and dataset are available on Github: http://github.com/graphcore-research/pyscf-ipu
Towards Foundational Models for Molecular Learning on Large-Scale Multi-Task Datasets
Oct 18, 2023



Abstract:Recently, pre-trained foundation models have enabled significant advancements in multiple fields. In molecular machine learning, however, where datasets are often hand-curated, and hence typically small, the lack of datasets with labeled features, and codebases to manage those datasets, has hindered the development of foundation models. In this work, we present seven novel datasets categorized by size into three distinct categories: ToyMix, LargeMix and UltraLarge. These datasets push the boundaries in both the scale and the diversity of supervised labels for molecular learning. They cover nearly 100 million molecules and over 3000 sparsely defined tasks, totaling more than 13 billion individual labels of both quantum and biological nature. In comparison, our datasets contain 300 times more data points than the widely used OGB-LSC PCQM4Mv2 dataset, and 13 times more than the quantum-only QM1B dataset. In addition, to support the development of foundational models based on our proposed datasets, we present the Graphium graph machine learning library which simplifies the process of building and training molecular machine learning models for multi-task and multi-level molecular datasets. Finally, we present a range of baseline results as a starting point of multi-task and multi-level training on these datasets. Empirically, we observe that performance on low-resource biological datasets show improvement by also training on large amounts of quantum data. This indicates that there may be potential in multi-task and multi-level training of a foundation model and fine-tuning it to resource-constrained downstream tasks.
PopSparse: Accelerated block sparse matrix multiplication on IPU
Apr 05, 2023



Abstract:Reducing the computational cost of running large scale neural networks using sparsity has attracted great attention in the deep learning community. While much success has been achieved in reducing FLOP and parameter counts while maintaining acceptable task performance, achieving actual speed improvements has typically been much more difficult, particularly on general purpose accelerators (GPAs) such as NVIDIA GPUs using low precision number formats. In this work we introduce PopSparse, a library that enables fast sparse operations on Graphcore IPUs by leveraging both the unique hardware characteristics of IPUs as well as any block structure defined in the data. We target two different types of sparsity: static, where the sparsity pattern is fixed at compile-time; and dynamic, where it can change each time the model is run. We present benchmark results for matrix multiplication for both of these modes on IPU with a range of block sizes, matrix sizes and densities. Results indicate that the PopSparse implementations are faster than dense matrix multiplications on IPU at a range of sparsity levels with large matrix size and block size. Furthermore, static sparsity in general outperforms dynamic sparsity. While previous work on GPAs has shown speedups only for very high sparsity (typically 99\% and above), the present work demonstrates that our static sparse implementation outperforms equivalent dense calculations in FP16 at lower sparsity (around 90%). IPU code is available to view and run at ipu.dev/sparsity-benchmarks, GPU code will be made available shortly.
GPS++: Reviving the Art of Message Passing for Molecular Property Prediction
Feb 06, 2023



Abstract:We present GPS++, a hybrid Message Passing Neural Network / Graph Transformer model for molecular property prediction. Our model integrates a well-tuned local message passing component and biased global attention with other key ideas from prior literature to achieve state-of-the-art results on large-scale molecular dataset PCQM4Mv2. Through a thorough ablation study we highlight the impact of individual components and, contrary to expectations set by recent trends, find that nearly all of the model's performance can be maintained without any use of global self-attention. We also show that our approach is significantly more accurate than prior art when 3D positional information is not available.
GPS++: An Optimised Hybrid MPNN/Transformer for Molecular Property Prediction
Dec 06, 2022



Abstract:This technical report presents GPS++, the first-place solution to the Open Graph Benchmark Large-Scale Challenge (OGB-LSC 2022) for the PCQM4Mv2 molecular property prediction task. Our approach implements several key principles from the prior literature. At its core our GPS++ method is a hybrid MPNN/Transformer model that incorporates 3D atom positions and an auxiliary denoising task. The effectiveness of GPS++ is demonstrated by achieving 0.0719 mean absolute error on the independent test-challenge PCQM4Mv2 split. Thanks to Graphcore IPU acceleration, GPS++ scales to deep architectures (16 layers), training at 3 minutes per epoch, and large ensemble (112 models), completing the final predictions in 1 hour 32 minutes, well under the 4 hour inference budget allocated. Our implementation is publicly available at: https://github.com/graphcore/ogb-lsc-pcqm4mv2.
8-bit Numerical Formats for Deep Neural Networks
Jun 06, 2022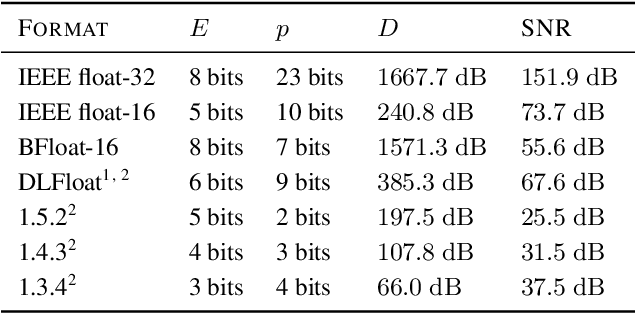
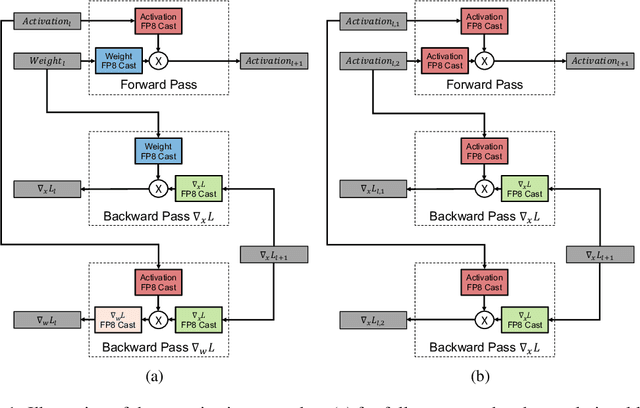
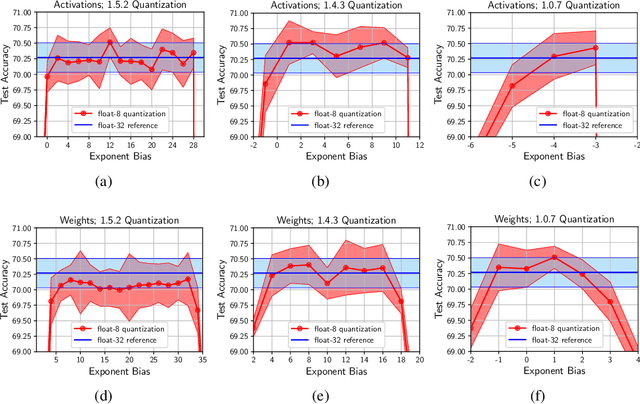
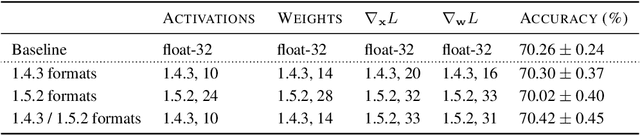
Abstract:Given the current trend of increasing size and complexity of machine learning architectures, it has become of critical importance to identify new approaches to improve the computational efficiency of model training. In this context, we address the advantages of floating-point over fixed-point representation, and present an in-depth study on the use of 8-bit floating-point number formats for activations, weights, and gradients for both training and inference. We explore the effect of different bit-widths for exponents and significands and different exponent biases. The experimental results demonstrate that a suitable choice of these low-precision formats enables faster training and reduced power consumption without any degradation in accuracy for a range of deep learning models for image classification and language processing.
Making EfficientNet More Efficient: Exploring Batch-Independent Normalization, Group Convolutions and Reduced Resolution Training
Jun 15, 2021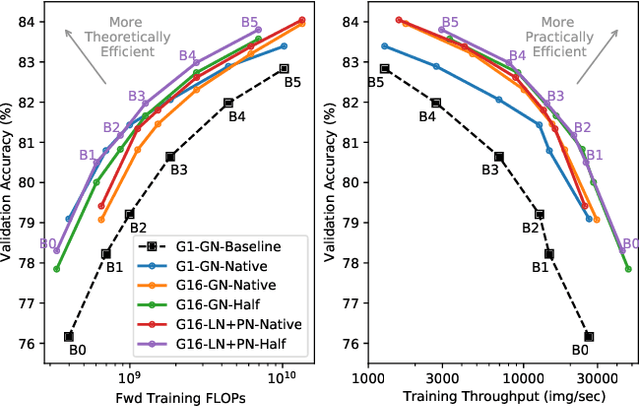

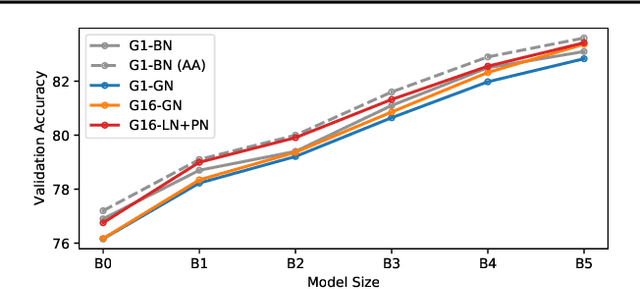
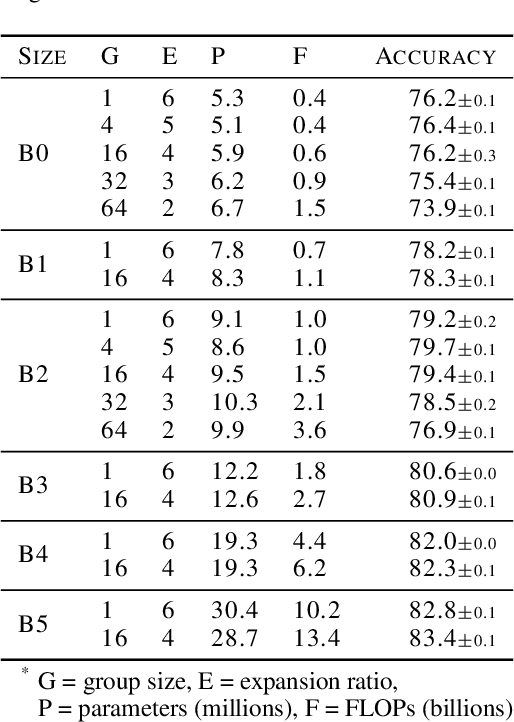
Abstract:Much recent research has been dedicated to improving the efficiency of training and inference for image classification. This effort has commonly focused on explicitly improving theoretical efficiency, often measured as ImageNet validation accuracy per FLOP. These theoretical savings have, however, proven challenging to achieve in practice, particularly on high-performance training accelerators. In this work, we focus on improving the practical efficiency of the state-of-the-art EfficientNet models on a new class of accelerator, the Graphcore IPU. We do this by extending this family of models in the following ways: (i) generalising depthwise convolutions to group convolutions; (ii) adding proxy-normalized activations to match batch normalization performance with batch-independent statistics; (iii) reducing compute by lowering the training resolution and inexpensively fine-tuning at higher resolution. We find that these three methods improve the practical efficiency for both training and inference. Our code will be made available online.
Proxy-Normalizing Activations to Match Batch Normalization while Removing Batch Dependence
Jun 11, 2021
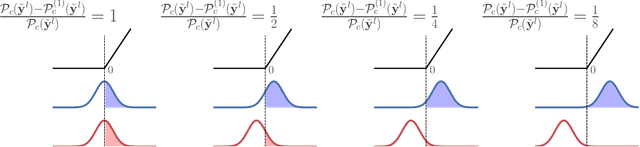

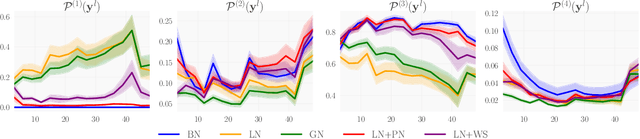
Abstract:We investigate the reasons for the performance degradation incurred with batch-independent normalization. We find that the prototypical techniques of layer normalization and instance normalization both induce the appearance of failure modes in the neural network's pre-activations: (i) layer normalization induces a collapse towards channel-wise constant functions; (ii) instance normalization induces a lack of variability in instance statistics, symptomatic of an alteration of the expressivity. To alleviate failure mode (i) without aggravating failure mode (ii), we introduce the technique "Proxy Normalization" that normalizes post-activations using a proxy distribution. When combined with layer normalization or group normalization, this batch-independent normalization emulates batch normalization's behavior and consistently matches or exceeds its performance.
Revisiting Small Batch Training for Deep Neural Networks
Apr 20, 2018



Abstract:Modern deep neural network training is typically based on mini-batch stochastic gradient optimization. While the use of large mini-batches increases the available computational parallelism, small batch training has been shown to provide improved generalization performance and allows a significantly smaller memory footprint, which might also be exploited to improve machine throughput. In this paper, we review common assumptions on learning rate scaling and training duration, as a basis for an experimental comparison of test performance for different mini-batch sizes. We adopt a learning rate that corresponds to a constant average weight update per gradient calculation (i.e., per unit cost of computation), and point out that this results in a variance of the weight updates that increases linearly with the mini-batch size $m$. The collected experimental results for the CIFAR-10, CIFAR-100 and ImageNet datasets show that increasing the mini-batch size progressively reduces the range of learning rates that provide stable convergence and acceptable test performance. On the other hand, small mini-batch sizes provide more up-to-date gradient calculations, which yields more stable and reliable training. The best performance has been consistently obtained for mini-batch sizes between $m = 2$ and $m = 32$, which contrasts with recent work advocating the use of mini-batch sizes in the thousands.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge