Daniel Golden
CoCa-CXR: Contrastive Captioners Learn Strong Temporal Structures for Chest X-Ray Vision-Language Understanding
Feb 27, 2025Abstract:Vision-language models have proven to be of great benefit for medical image analysis since they learn rich semantics from both images and reports. Prior efforts have focused on better alignment of image and text representations to enhance image understanding. However, though explicit reference to a prior image is common in Chest X-Ray (CXR) reports, aligning progression descriptions with the semantics differences in image pairs remains under-explored. In this work, we propose two components to address this issue. (1) A CXR report processing pipeline to extract temporal structure. It processes reports with a large language model (LLM) to separate the description and comparison contexts, and extracts fine-grained annotations from reports. (2) A contrastive captioner model for CXR, namely CoCa-CXR, to learn how to both describe images and their temporal progressions. CoCa-CXR incorporates a novel regional cross-attention module to identify local differences between paired CXR images. Extensive experiments show the superiority of CoCa-CXR on both progression analysis and report generation compared to previous methods. Notably, on MS-CXR-T progression classification, CoCa-CXR obtains 65.0% average testing accuracy on five pulmonary conditions, outperforming the previous state-of-the-art (SOTA) model BioViL-T by 4.8%. It also achieves a RadGraph F1 of 24.2% on MIMIC-CXR, which is comparable to the Med-Gemini foundation model.
Health AI Developer Foundations
Nov 26, 2024



Abstract:Robust medical Machine Learning (ML) models have the potential to revolutionize healthcare by accelerating clinical research, improving workflows and outcomes, and producing novel insights or capabilities. Developing such ML models from scratch is cost prohibitive and requires substantial compute, data, and time (e.g., expert labeling). To address these challenges, we introduce Health AI Developer Foundations (HAI-DEF), a suite of pre-trained, domain-specific foundation models, tools, and recipes to accelerate building ML for health applications. The models cover various modalities and domains, including radiology (X-rays and computed tomography), histopathology, dermatological imaging, and audio. These models provide domain specific embeddings that facilitate AI development with less labeled data, shorter training times, and reduced computational costs compared to traditional approaches. In addition, we utilize a common interface and style across these models, and prioritize usability to enable developers to integrate HAI-DEF efficiently. We present model evaluations across various tasks and conclude with a discussion of their application and evaluation, covering the importance of ensuring efficacy, fairness, and equity. Finally, while HAI-DEF and specifically the foundation models lower the barrier to entry for ML in healthcare, we emphasize the importance of validation with problem- and population-specific data for each desired usage setting. This technical report will be updated over time as more modalities and features are added.
PathAlign: A vision-language model for whole slide images in histopathology
Jun 27, 2024Abstract:Microscopic interpretation of histopathology images underlies many important diagnostic and treatment decisions. While advances in vision-language modeling raise new opportunities for analysis of such images, the gigapixel-scale size of whole slide images (WSIs) introduces unique challenges. Additionally, pathology reports simultaneously highlight key findings from small regions while also aggregating interpretation across multiple slides, often making it difficult to create robust image-text pairs. As such, pathology reports remain a largely untapped source of supervision in computational pathology, with most efforts relying on region-of-interest annotations or self-supervision at the patch-level. In this work, we develop a vision-language model based on the BLIP-2 framework using WSIs paired with curated text from pathology reports. This enables applications utilizing a shared image-text embedding space, such as text or image retrieval for finding cases of interest, as well as integration of the WSI encoder with a frozen large language model (LLM) for WSI-based generative text capabilities such as report generation or AI-in-the-loop interactions. We utilize a de-identified dataset of over 350,000 WSIs and diagnostic text pairs, spanning a wide range of diagnoses, procedure types, and tissue types. We present pathologist evaluation of text generation and text retrieval using WSI embeddings, as well as results for WSI classification and workflow prioritization (slide-level triaging). Model-generated text for WSIs was rated by pathologists as accurate, without clinically significant error or omission, for 78% of WSIs on average. This work demonstrates exciting potential capabilities for language-aligned WSI embeddings.
Advancing Multimodal Medical Capabilities of Gemini
May 06, 2024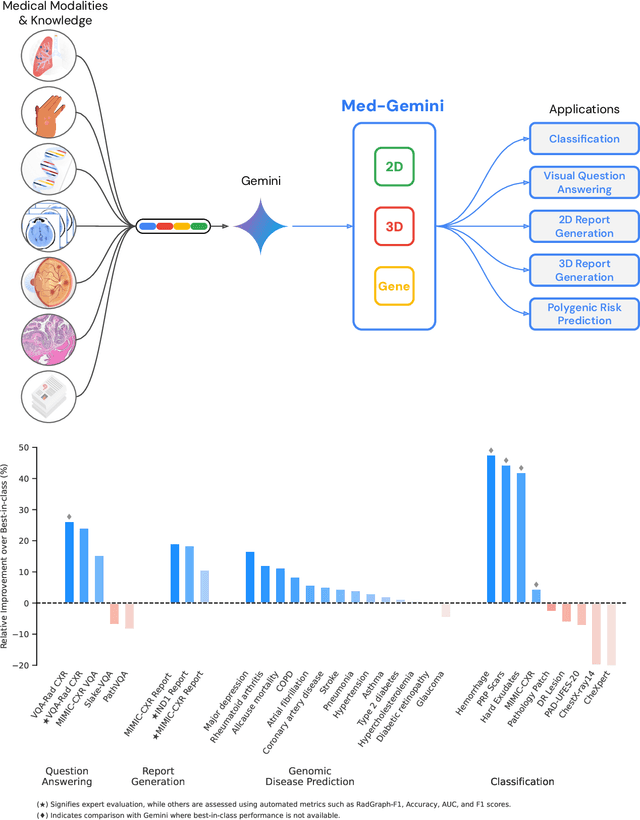
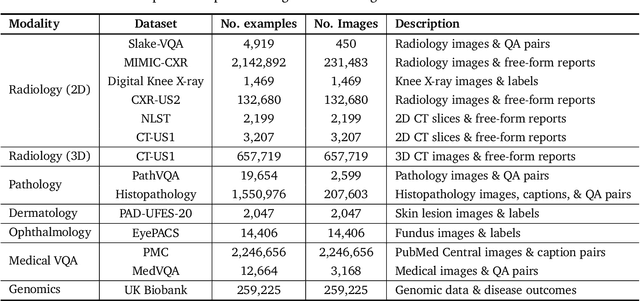

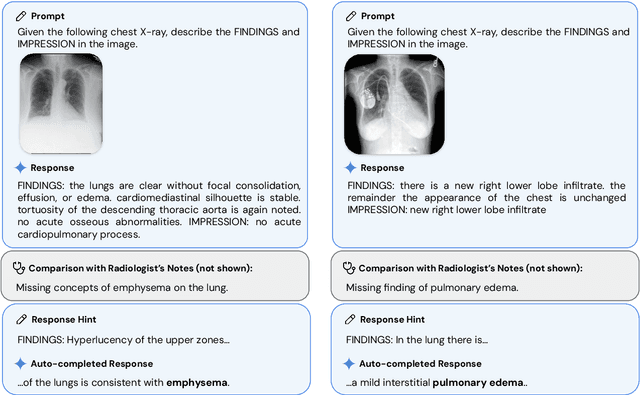
Abstract:Many clinical tasks require an understanding of specialized data, such as medical images and genomics, which is not typically found in general-purpose large multimodal models. Building upon Gemini's multimodal models, we develop several models within the new Med-Gemini family that inherit core capabilities of Gemini and are optimized for medical use via fine-tuning with 2D and 3D radiology, histopathology, ophthalmology, dermatology and genomic data. Med-Gemini-2D sets a new standard for AI-based chest X-ray (CXR) report generation based on expert evaluation, exceeding previous best results across two separate datasets by an absolute margin of 1% and 12%, where 57% and 96% of AI reports on normal cases, and 43% and 65% on abnormal cases, are evaluated as "equivalent or better" than the original radiologists' reports. We demonstrate the first ever large multimodal model-based report generation for 3D computed tomography (CT) volumes using Med-Gemini-3D, with 53% of AI reports considered clinically acceptable, although additional research is needed to meet expert radiologist reporting quality. Beyond report generation, Med-Gemini-2D surpasses the previous best performance in CXR visual question answering (VQA) and performs well in CXR classification and radiology VQA, exceeding SoTA or baselines on 17 of 20 tasks. In histopathology, ophthalmology, and dermatology image classification, Med-Gemini-2D surpasses baselines across 18 out of 20 tasks and approaches task-specific model performance. Beyond imaging, Med-Gemini-Polygenic outperforms the standard linear polygenic risk score-based approach for disease risk prediction and generalizes to genetically correlated diseases for which it has never been trained. Although further development and evaluation are necessary in the safety-critical medical domain, our results highlight the potential of Med-Gemini across a wide range of medical tasks.
ELIXR: Towards a general purpose X-ray artificial intelligence system through alignment of large language models and radiology vision encoders
Aug 02, 2023Abstract:Our approach, which we call Embeddings for Language/Image-aligned X-Rays, or ELIXR, leverages a language-aligned image encoder combined or grafted onto a fixed LLM, PaLM 2, to perform a broad range of tasks. We train this lightweight adapter architecture using images paired with corresponding free-text radiology reports from the MIMIC-CXR dataset. ELIXR achieved state-of-the-art performance on zero-shot chest X-ray (CXR) classification (mean AUC of 0.850 across 13 findings), data-efficient CXR classification (mean AUCs of 0.893 and 0.898 across five findings (atelectasis, cardiomegaly, consolidation, pleural effusion, and pulmonary edema) for 1% (~2,200 images) and 10% (~22,000 images) training data), and semantic search (0.76 normalized discounted cumulative gain (NDCG) across nineteen queries, including perfect retrieval on twelve of them). Compared to existing data-efficient methods including supervised contrastive learning (SupCon), ELIXR required two orders of magnitude less data to reach similar performance. ELIXR also showed promise on CXR vision-language tasks, demonstrating overall accuracies of 58.7% and 62.5% on visual question answering and report quality assurance tasks, respectively. These results suggest that ELIXR is a robust and versatile approach to CXR AI.
ScarGAN: Chained Generative Adversarial Networks to Simulate Pathological Tissue on Cardiovascular MR Scans
Aug 14, 2018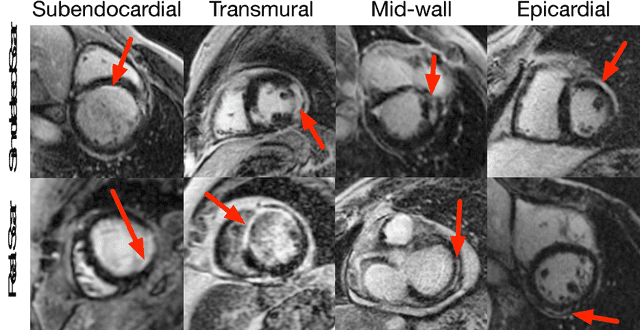
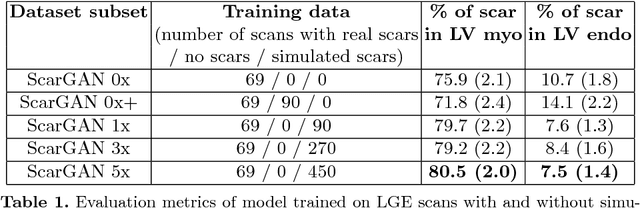
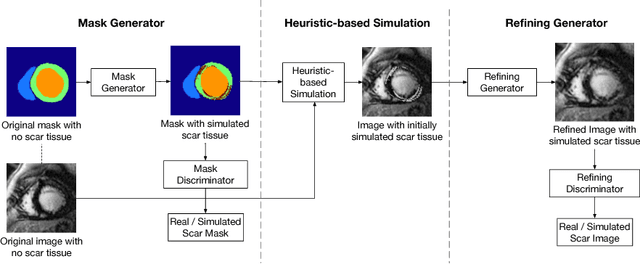
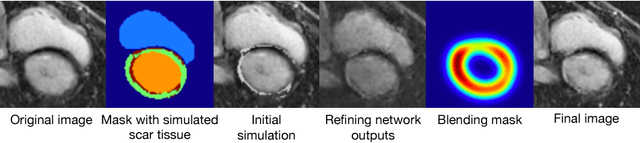
Abstract:Medical images with specific pathologies are scarce, but a large amount of data is usually required for a deep convolutional neural network (DCNN) to achieve good accuracy. We consider the problem of segmenting the left ventricular (LV) myocardium on late gadolinium enhancement (LGE) cardiovascular magnetic resonance (CMR) scans of which only some of the scans have scar tissue. We propose ScarGAN to simulate scar tissue on healthy myocardium using chained generative adversarial networks (GAN). Our novel approach factorizes the simulation process into 3 steps: 1) a mask generator to simulate the shape of the scar tissue; 2) a domain-specific heuristic to produce the initial simulated scar tissue from the simulated shape; 3) a refining generator to add details to the simulated scar tissue. Unlike other approaches that generate samples from scratch, we simulate scar tissue on normal scans resulting in highly realistic samples. We show that experienced radiologists are unable to distinguish between real and simulated scar tissue. Training a U-Net with additional scans with scar tissue simulated by ScarGAN increases the percentage of scar pixels correctly included in LV myocardium prediction from 75.9% to 80.5%.
Computationally efficient cardiac views projection using 3D Convolutional Neural Networks
Nov 03, 2017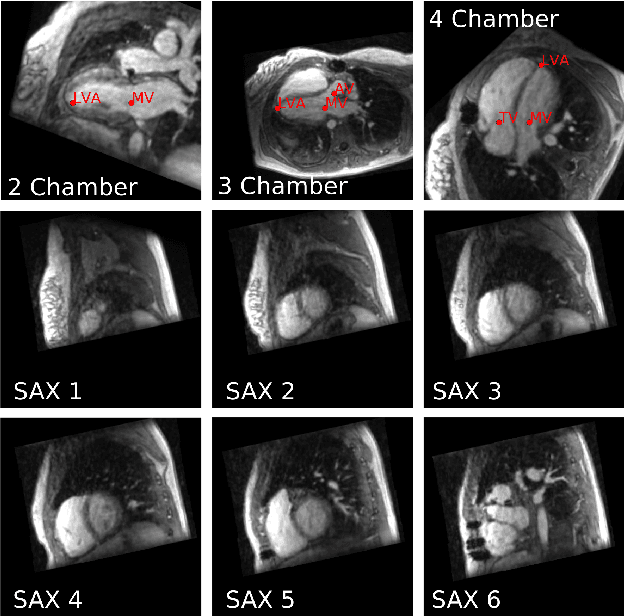
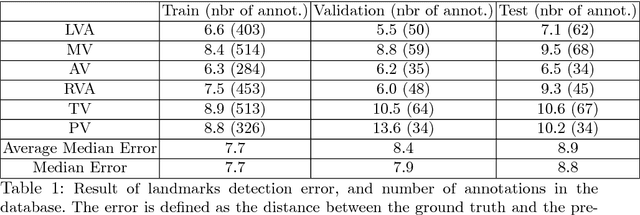
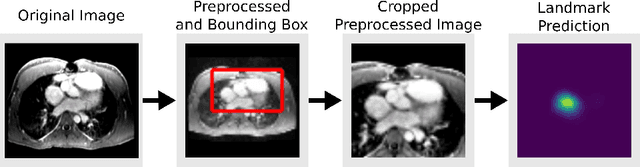
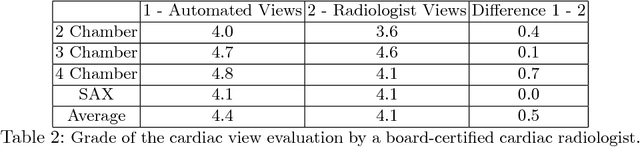
Abstract:4D Flow is an MRI sequence which allows acquisition of 3D images of the heart. The data is typically acquired volumetrically, so it must be reformatted to generate cardiac long axis and short axis views for diagnostic interpretation. These views may be generated by placing 6 landmarks: the left and right ventricle apex, and the aortic, mitral, pulmonary, and tricuspid valves. In this paper, we propose an automatic method to localize landmarks in order to compute the cardiac views. Our approach consists of first calculating a bounding box that tightly crops the heart, followed by a landmark localization step within this bounded region. Both steps are based on a 3D extension of the recently introduced ENet. We demonstrate that the long and short axis projections computed with our automated method are of equivalent quality to projections created with landmarks placed by an experienced cardiac radiologist, based on a blinded test administered to a different cardiac radiologist.
FastVentricle: Cardiac Segmentation with ENet
Apr 13, 2017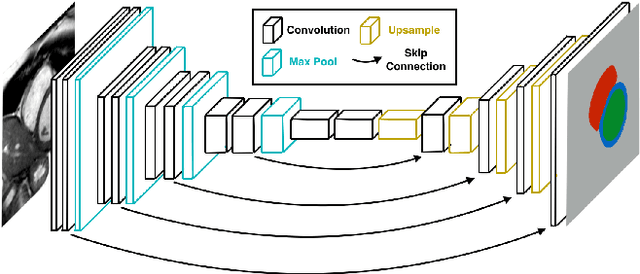
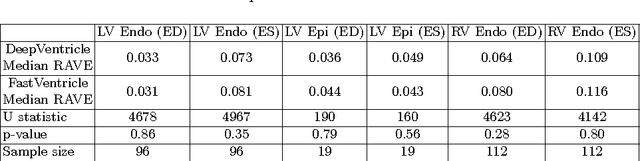
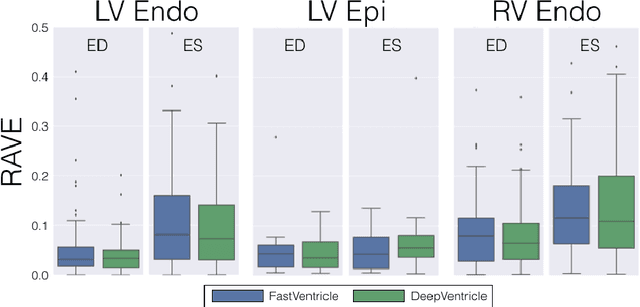
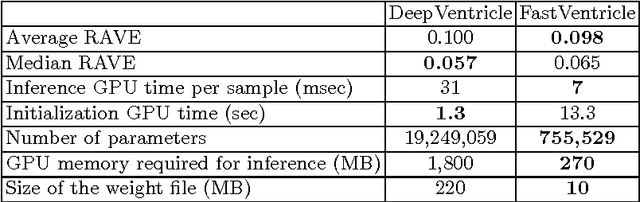
Abstract:Cardiac Magnetic Resonance (CMR) imaging is commonly used to assess cardiac structure and function. One disadvantage of CMR is that post-processing of exams is tedious. Without automation, precise assessment of cardiac function via CMR typically requires an annotator to spend tens of minutes per case manually contouring ventricular structures. Automatic contouring can lower the required time per patient by generating contour suggestions that can be lightly modified by the annotator. Fully convolutional networks (FCNs), a variant of convolutional neural networks, have been used to rapidly advance the state-of-the-art in automated segmentation, which makes FCNs a natural choice for ventricular segmentation. However, FCNs are limited by their computational cost, which increases the monetary cost and degrades the user experience of production systems. To combat this shortcoming, we have developed the FastVentricle architecture, an FCN architecture for ventricular segmentation based on the recently developed ENet architecture. FastVentricle is 4x faster and runs with 6x less memory than the previous state-of-the-art ventricular segmentation architecture while still maintaining excellent clinical accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge