Jesse Lieman-Sifry
ScarGAN: Chained Generative Adversarial Networks to Simulate Pathological Tissue on Cardiovascular MR Scans
Aug 14, 2018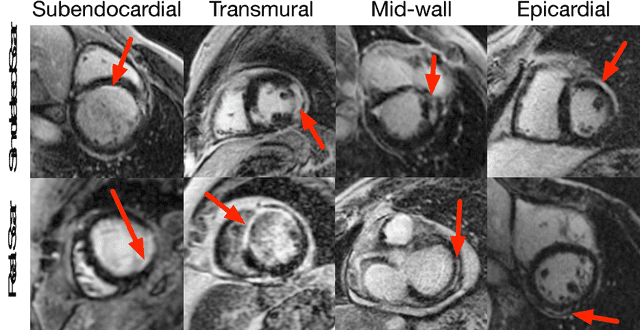
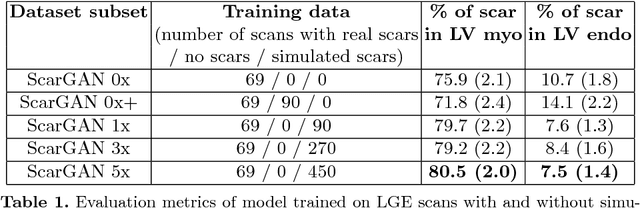
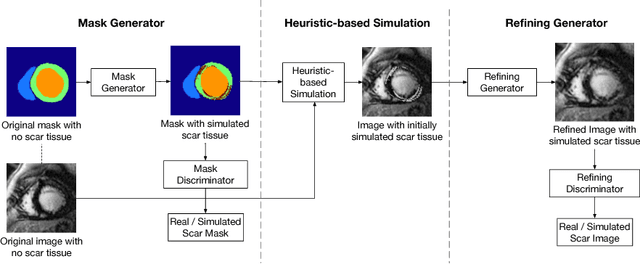
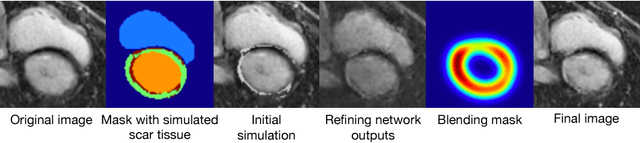
Abstract:Medical images with specific pathologies are scarce, but a large amount of data is usually required for a deep convolutional neural network (DCNN) to achieve good accuracy. We consider the problem of segmenting the left ventricular (LV) myocardium on late gadolinium enhancement (LGE) cardiovascular magnetic resonance (CMR) scans of which only some of the scans have scar tissue. We propose ScarGAN to simulate scar tissue on healthy myocardium using chained generative adversarial networks (GAN). Our novel approach factorizes the simulation process into 3 steps: 1) a mask generator to simulate the shape of the scar tissue; 2) a domain-specific heuristic to produce the initial simulated scar tissue from the simulated shape; 3) a refining generator to add details to the simulated scar tissue. Unlike other approaches that generate samples from scratch, we simulate scar tissue on normal scans resulting in highly realistic samples. We show that experienced radiologists are unable to distinguish between real and simulated scar tissue. Training a U-Net with additional scans with scar tissue simulated by ScarGAN increases the percentage of scar pixels correctly included in LV myocardium prediction from 75.9% to 80.5%.
Computationally efficient cardiac views projection using 3D Convolutional Neural Networks
Nov 03, 2017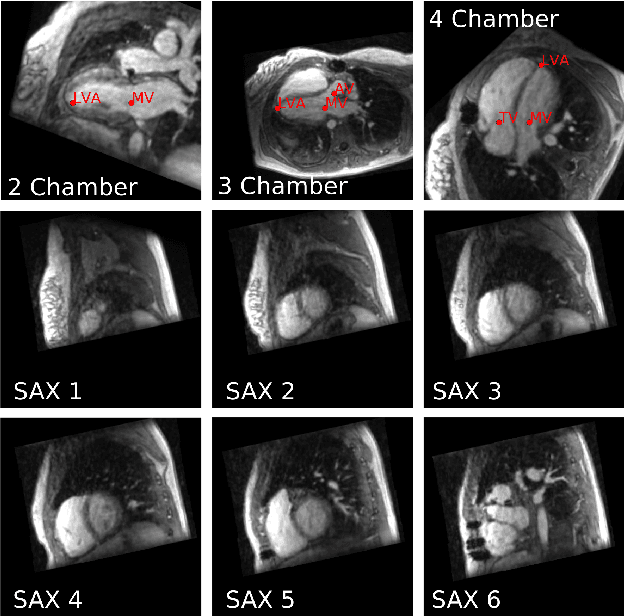
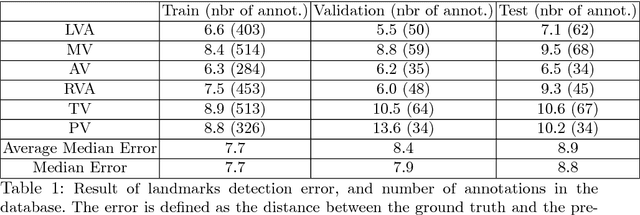
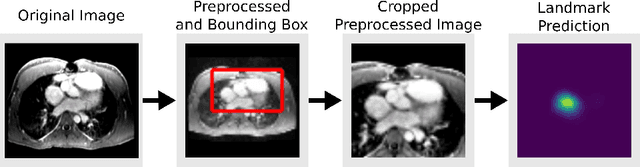
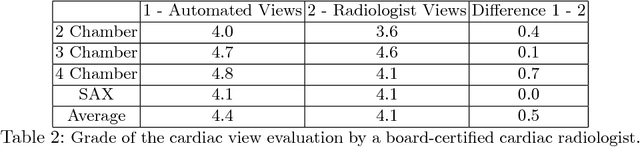
Abstract:4D Flow is an MRI sequence which allows acquisition of 3D images of the heart. The data is typically acquired volumetrically, so it must be reformatted to generate cardiac long axis and short axis views for diagnostic interpretation. These views may be generated by placing 6 landmarks: the left and right ventricle apex, and the aortic, mitral, pulmonary, and tricuspid valves. In this paper, we propose an automatic method to localize landmarks in order to compute the cardiac views. Our approach consists of first calculating a bounding box that tightly crops the heart, followed by a landmark localization step within this bounded region. Both steps are based on a 3D extension of the recently introduced ENet. We demonstrate that the long and short axis projections computed with our automated method are of equivalent quality to projections created with landmarks placed by an experienced cardiac radiologist, based on a blinded test administered to a different cardiac radiologist.
FastVentricle: Cardiac Segmentation with ENet
Apr 13, 2017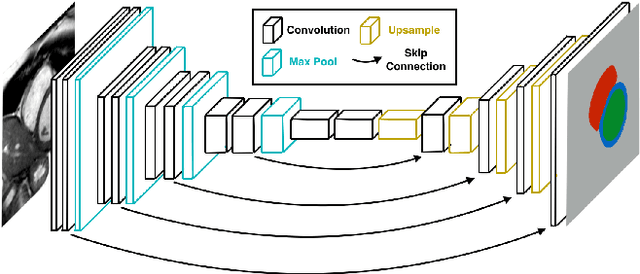
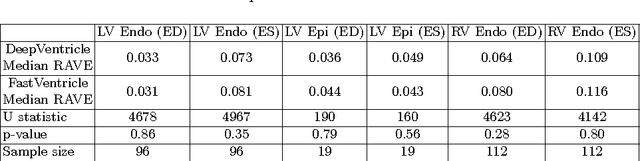
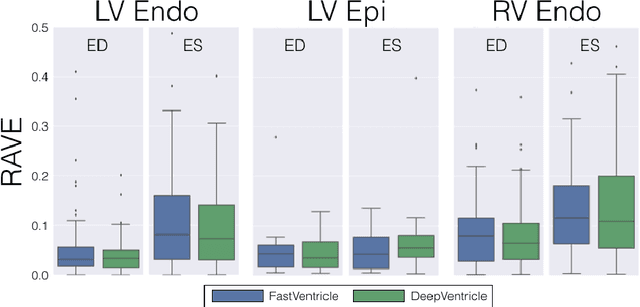
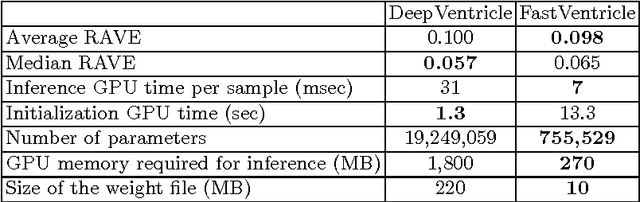
Abstract:Cardiac Magnetic Resonance (CMR) imaging is commonly used to assess cardiac structure and function. One disadvantage of CMR is that post-processing of exams is tedious. Without automation, precise assessment of cardiac function via CMR typically requires an annotator to spend tens of minutes per case manually contouring ventricular structures. Automatic contouring can lower the required time per patient by generating contour suggestions that can be lightly modified by the annotator. Fully convolutional networks (FCNs), a variant of convolutional neural networks, have been used to rapidly advance the state-of-the-art in automated segmentation, which makes FCNs a natural choice for ventricular segmentation. However, FCNs are limited by their computational cost, which increases the monetary cost and degrades the user experience of production systems. To combat this shortcoming, we have developed the FastVentricle architecture, an FCN architecture for ventricular segmentation based on the recently developed ENet architecture. FastVentricle is 4x faster and runs with 6x less memory than the previous state-of-the-art ventricular segmentation architecture while still maintaining excellent clinical accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge