Brian L. Hill
OrdShap: Feature Position Importance for Sequential Black-Box Models
Jul 16, 2025



Abstract:Sequential deep learning models excel in domains with temporal or sequential dependencies, but their complexity necessitates post-hoc feature attribution methods for understanding their predictions. While existing techniques quantify feature importance, they inherently assume fixed feature ordering - conflating the effects of (1) feature values and (2) their positions within input sequences. To address this gap, we introduce OrdShap, a novel attribution method that disentangles these effects by quantifying how a model's predictions change in response to permuting feature position. We establish a game-theoretic connection between OrdShap and Sanchez-Berganti\~nos values, providing a theoretically grounded approach to position-sensitive attribution. Empirical results from health, natural language, and synthetic datasets highlight OrdShap's effectiveness in capturing feature value and feature position attributions, and provide deeper insight into model behavior.
Zero-shot Medical Event Prediction Using a Generative Pre-trained Transformer on Electronic Health Records
Mar 07, 2025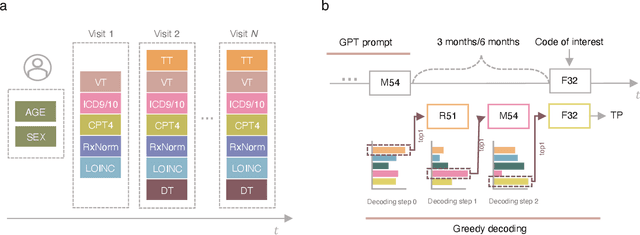
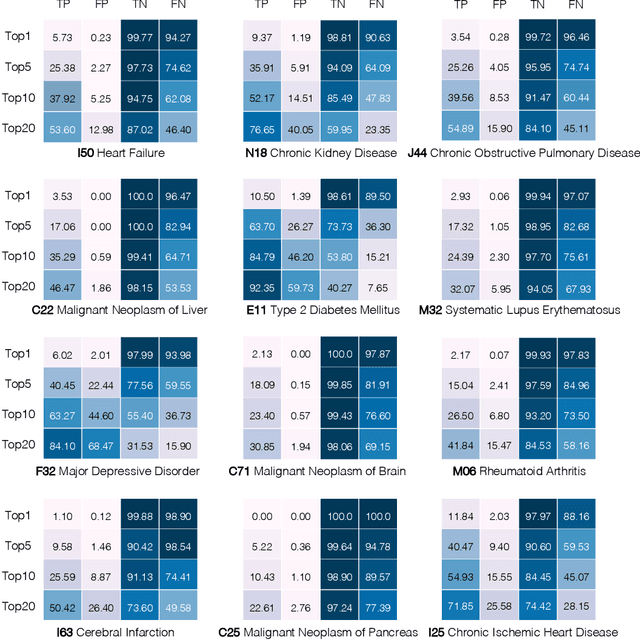
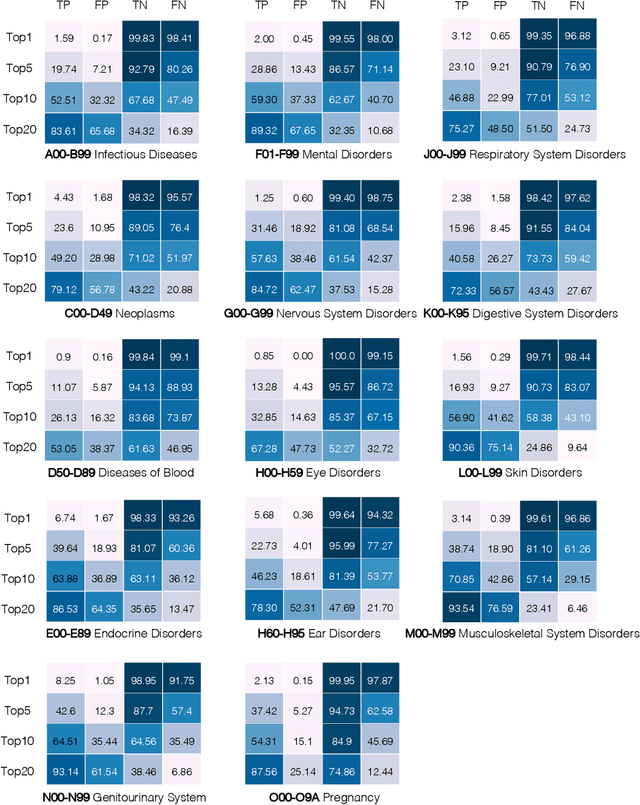
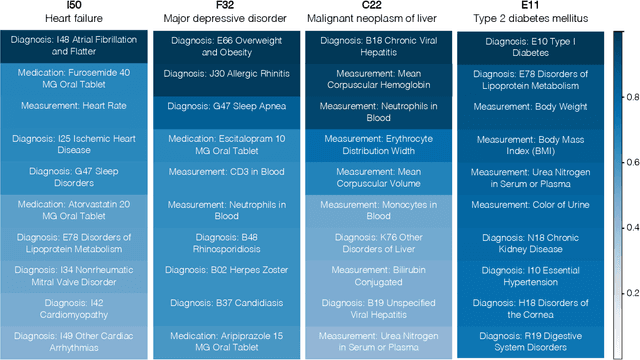
Abstract:Longitudinal data in electronic health records (EHRs) represent an individual`s clinical history through a sequence of codified concepts, including diagnoses, procedures, medications, and laboratory tests. Foundational models, such as generative pre-trained transformers (GPT), can leverage this data to predict future events. While fine-tuning of these models enhances task-specific performance, it is costly, complex, and unsustainable for every target. We show that a foundation model trained on EHRs can perform predictive tasks in a zero-shot manner, eliminating the need for fine-tuning. This study presents the first comprehensive analysis of zero-shot forecasting with GPT-based foundational models in EHRs, introducing a novel pipeline that formulates medical concept prediction as a generative modeling task. Unlike supervised approaches requiring extensive labeled data, our method enables the model to forecast a next medical event purely from a pretraining knowledge. We evaluate performance across multiple time horizons and clinical categories, demonstrating model`s ability to capture latent temporal dependencies and complex patient trajectories without task supervision. Model performance for predicting the next medical concept was evaluated using precision and recall metrics, achieving an average top1 precision of 0.614 and recall of 0.524. For 12 major diagnostic conditions, the model demonstrated strong zero-shot performance, achieving high true positive rates while maintaining low false positives. We demonstrate the power of a foundational EHR GPT model in capturing diverse phenotypes and enabling robust, zero-shot forecasting of clinical outcomes. This capability enhances the versatility of predictive healthcare models and reduces the need for task-specific training, enabling more scalable applications in clinical settings.
Extend and Explain: Interpreting Very Long Language Models
Sep 07, 2022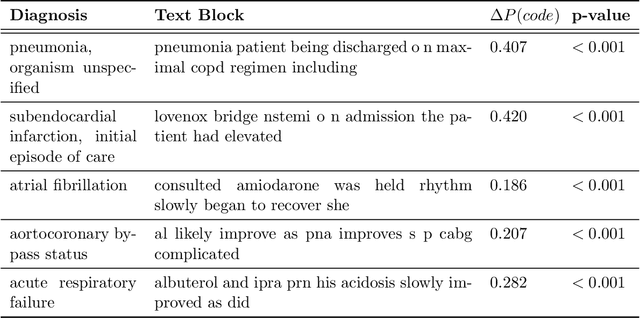
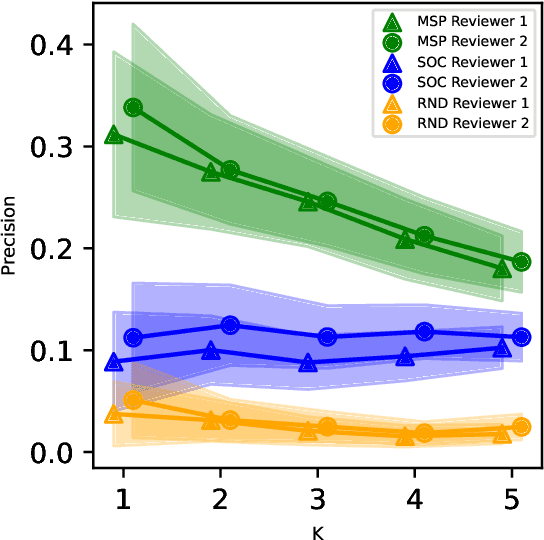
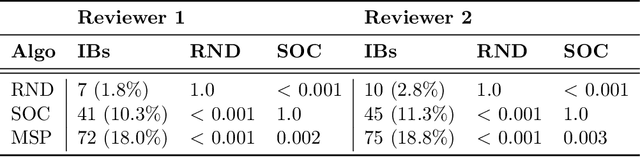
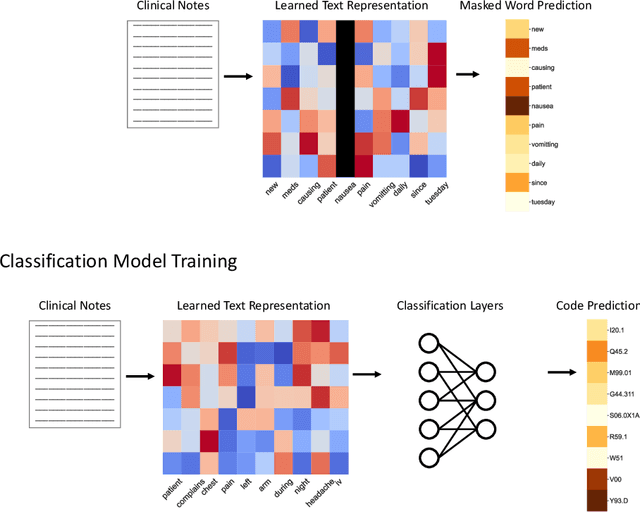
Abstract:While Transformer language models (LMs) are state-of-the-art for information extraction, long text introduces computational challenges requiring suboptimal preprocessing steps or alternative model architectures. Sparse-attention LMs can represent longer sequences, overcoming performance hurdles. However, it remains unclear how to explain predictions from these models, as not all tokens attend to each other in the self-attention layers, and long sequences pose computational challenges for explainability algorithms when runtime depends on document length. These challenges are severe in the medical context where documents can be very long, and machine learning (ML) models must be auditable and trustworthy. We introduce a novel Masked Sampling Procedure (MSP) to identify the text blocks that contribute to a prediction, apply MSP in the context of predicting diagnoses from medical text, and validate our approach with a blind review by two clinicians. Our method identifies about 1.7x more clinically informative text blocks than the previous state-of-the-art, runs up to 100x faster, and is tractable for generating important phrase pairs. MSP is particularly well-suited to long LMs but can be applied to any text classifier. We provide a general implementation of MSP.
SCAMPS: Synthetics for Camera Measurement of Physiological Signals
Jun 08, 2022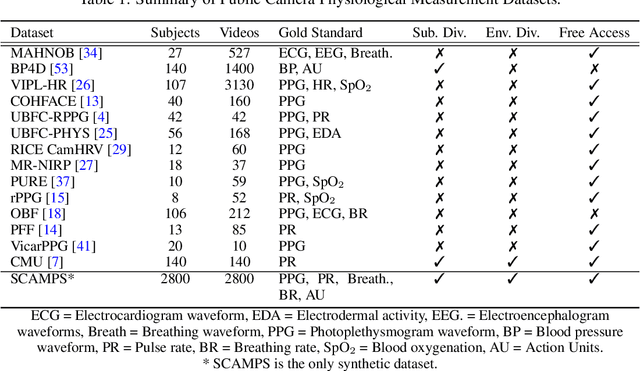
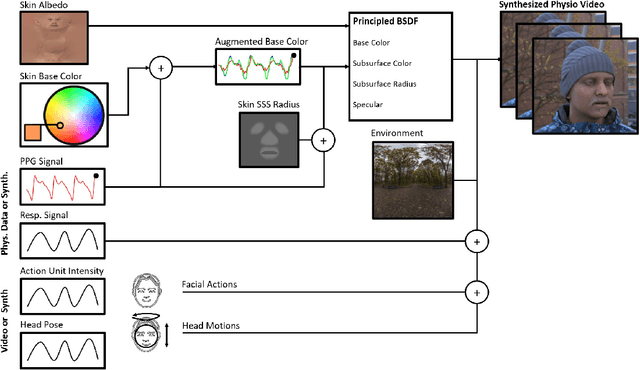
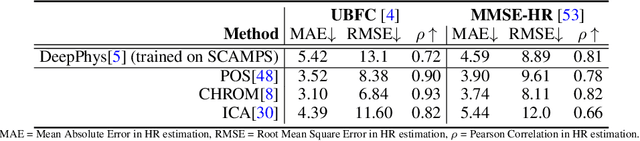

Abstract:The use of cameras and computational algorithms for noninvasive, low-cost and scalable measurement of physiological (e.g., cardiac and pulmonary) vital signs is very attractive. However, diverse data representing a range of environments, body motions, illumination conditions and physiological states is laborious, time consuming and expensive to obtain. Synthetic data have proven a valuable tool in several areas of machine learning, yet are not widely available for camera measurement of physiological states. Synthetic data offer "perfect" labels (e.g., without noise and with precise synchronization), labels that may not be possible to obtain otherwise (e.g., precise pixel level segmentation maps) and provide a high degree of control over variation and diversity in the dataset. We present SCAMPS, a dataset of synthetics containing 2,800 videos (1.68M frames) with aligned cardiac and respiratory signals and facial action intensities. The RGB frames are provided alongside segmentation maps. We provide precise descriptive statistics about the underlying waveforms, including inter-beat interval, heart rate variability, and pulse arrival time. Finally, we present baseline results training on these synthetic data and testing on real-world datasets to illustrate generalizability.
EfficientPhys: Enabling Simple, Fast and Accurate Camera-Based Vitals Measurement
Oct 09, 2021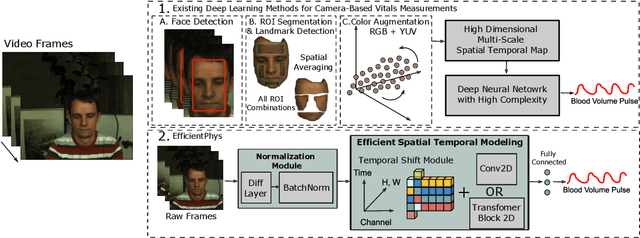
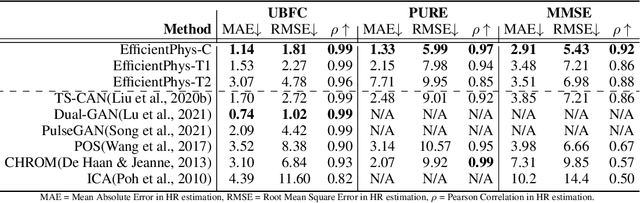


Abstract:Camera-based physiological measurement is a growing field with neural models providing state-the-art-performance. Prior research have explored various ``end-to-end'' models; however these methods still require several preprocessing steps. These additional operations are often non-trivial to implement making replication and deployment difficult and can even have a higher computational budget than the ``core'' network itself. In this paper, we propose two novel and efficient neural models for camera-based physiological measurement called EfficientPhys that remove the need for face detection, segmentation, normalization, color space transformation or any other preprocessing steps. Using an input of raw video frames, our models achieve state-of-the-art accuracy on three public datasets. We show that this is the case whether using a transformer or convolutional backbone. We further evaluate the latency of the proposed networks and show that our most light weight network also achieves a 33% improvement in efficiency.
Learning Higher-Order Dynamics in Video-Based Cardiac Measurement
Oct 07, 2021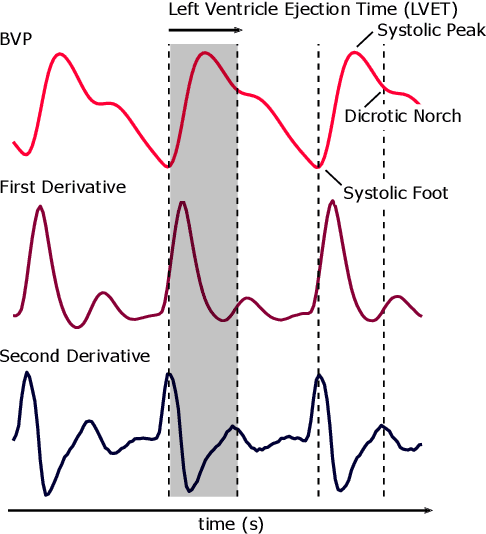
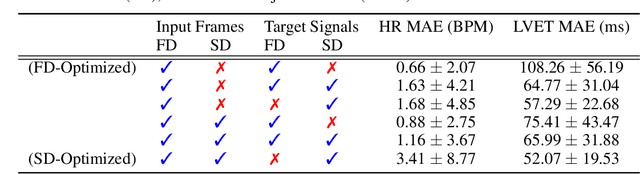
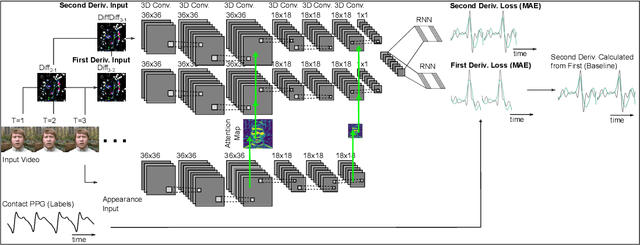
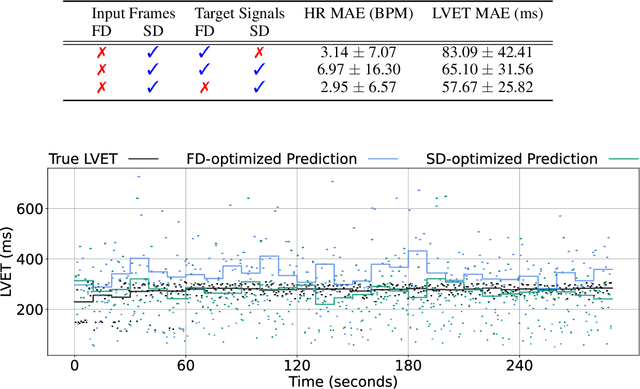
Abstract:Computer vision methods typically optimize for first-order dynamics (e.g., optical flow). However, in many cases the properties of interest are subtle variations in higher-order changes, such as acceleration. This is true in the cardiac pulse, where the second derivative can be used as an indicator of blood pressure and arterial disease. Recent developments in camera-based vital sign measurement have shown that cardiac measurements can be recovered with impressive accuracy from videos; however, the majority of research has focused on extracting summary statistics such as heart rate. Less emphasis has been put on the accuracy of waveform morphology that is necessary for many clinically impactful scenarios. In this work, we provide evidence that higher-order dynamics are better estimated by neural models when explicitly optimized for in the loss function. Furthermore, adding second-derivative inputs also improves performance when estimating second-order dynamics. By incorporating the second derivative of both the input frames and the target vital sign signals into the training procedure, our model is better able to estimate left ventricle ejection time (LVET) intervals.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge