Zhongzhao Teng
on behalf of the AIX-COVNET collaboration
Advancing COVID-19 Diagnosis with Privacy-Preserving Collaboration in Artificial Intelligence
Nov 18, 2021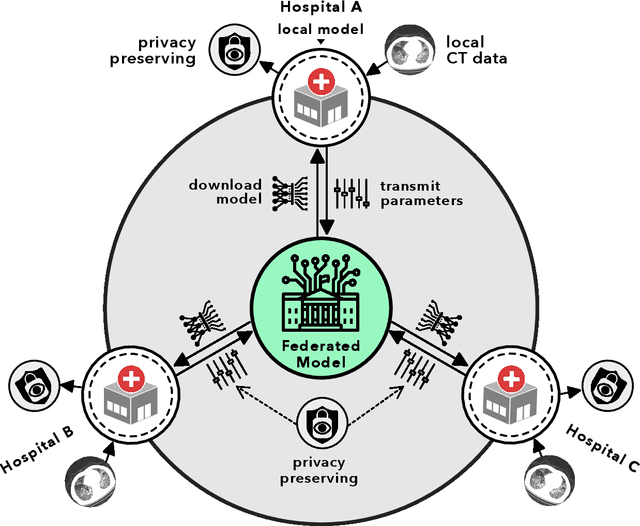
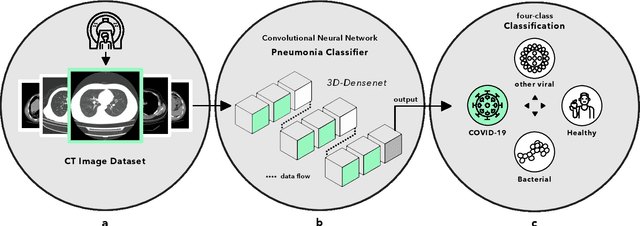
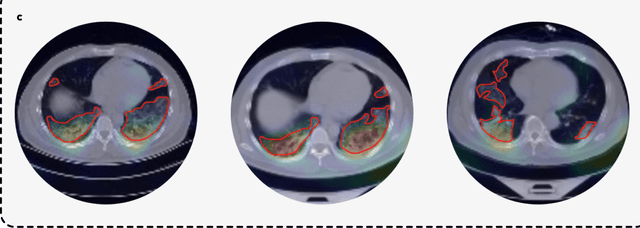
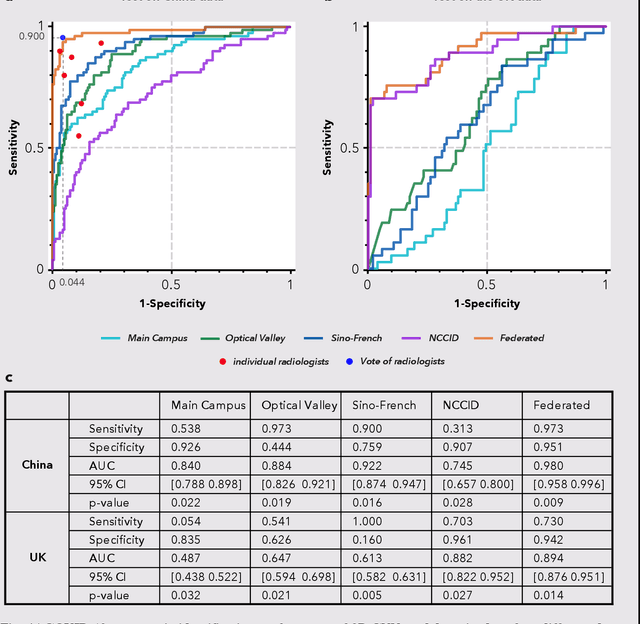
Abstract:Artificial intelligence (AI) provides a promising substitution for streamlining COVID-19 diagnoses. However, concerns surrounding security and trustworthiness impede the collection of large-scale representative medical data, posing a considerable challenge for training a well-generalised model in clinical practices. To address this, we launch the Unified CT-COVID AI Diagnostic Initiative (UCADI), where the AI model can be distributedly trained and independently executed at each host institution under a federated learning framework (FL) without data sharing. Here we show that our FL model outperformed all the local models by a large yield (test sensitivity /specificity in China: 0.973/0.951, in the UK: 0.730/0.942), achieving comparable performance with a panel of professional radiologists. We further evaluated the model on the hold-out (collected from another two hospitals leaving out the FL) and heterogeneous (acquired with contrast materials) data, provided visual explanations for decisions made by the model, and analysed the trade-offs between the model performance and the communication costs in the federated training process. Our study is based on 9,573 chest computed tomography scans (CTs) from 3,336 patients collected from 23 hospitals located in China and the UK. Collectively, our work advanced the prospects of utilising federated learning for privacy-preserving AI in digital health.
Machine learning for COVID-19 detection and prognostication using chest radiographs and CT scans: a systematic methodological review
Sep 01, 2020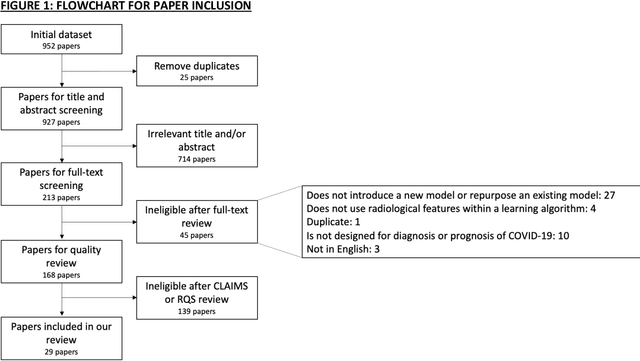
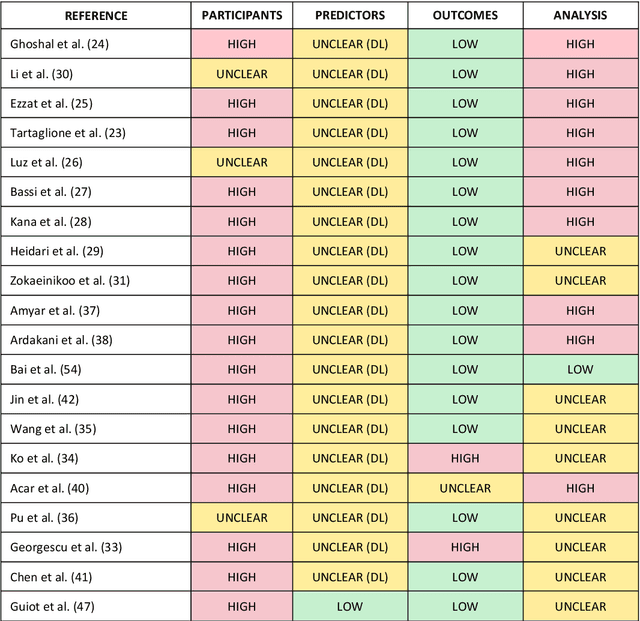
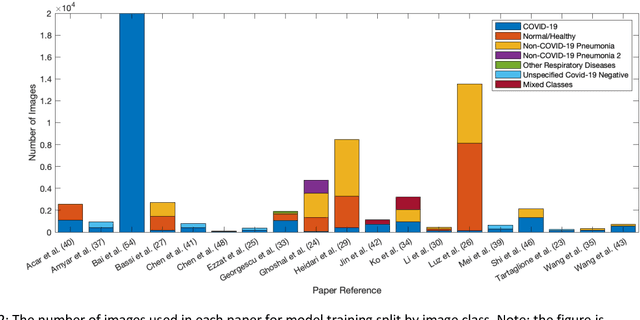
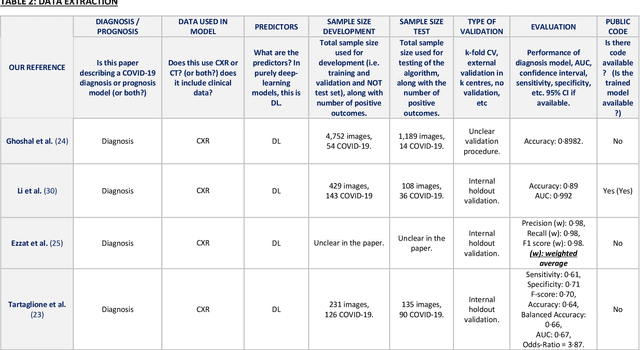
Abstract:Background: Machine learning methods offer great potential for fast and accurate detection and prognostication of COVID-19 from standard-of-care chest radiographs (CXR) and computed tomography (CT) images. In this systematic review we critically evaluate the machine learning methodologies employed in the rapidly growing literature. Methods: In this systematic review we reviewed EMBASE via OVID, MEDLINE via PubMed, bioRxiv, medRxiv and arXiv for published papers and preprints uploaded from Jan 1, 2020 to June 24, 2020. Studies which consider machine learning models for the diagnosis or prognosis of COVID-19 from CXR or CT images were included. A methodology quality review of each paper was performed against established benchmarks to ensure the review focusses only on high-quality reproducible papers. This study is registered with PROSPERO [CRD42020188887]. Interpretation: Our review finds that none of the developed models discussed are of potential clinical use due to methodological flaws and underlying biases. This is a major weakness, given the urgency with which validated COVID-19 models are needed. Typically, we find that the documentation of a model's development is not sufficient to make the results reproducible and therefore of 168 candidate papers only 29 are deemed to be reproducible and subsequently considered in this review. We therefore encourage authors to use established machine learning checklists to ensure sufficient documentation is made available, and to follow the PROBAST (prediction model risk of bias assessment tool) framework to determine the underlying biases in their model development process and to mitigate these where possible. This is key to safe clinical implementation which is urgently needed.
Efficient Deep Representation Learning by Adaptive Latent Space Sampling
Apr 12, 2020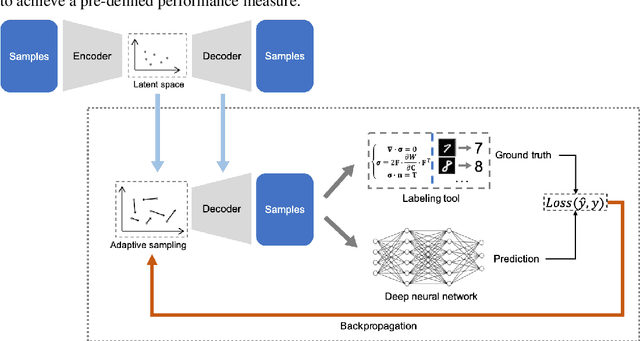
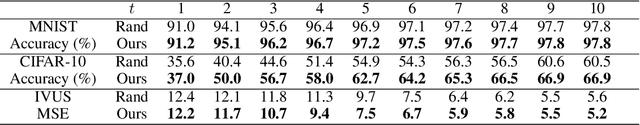

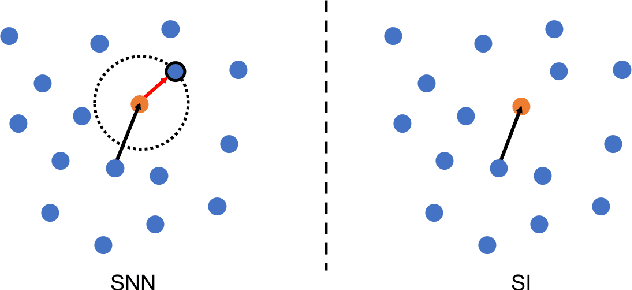
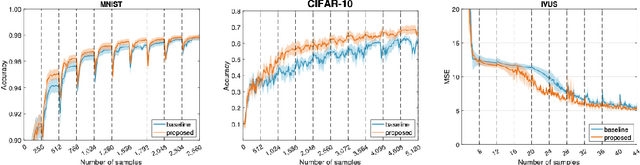
Abstract:Supervised deep learning requires a large amount of training samples with annotations (e.g. label class for classification task, pixel- or voxel-wised label map for segmentation tasks), which are expensive and time-consuming to obtain. During the training of a deep neural network, the annotated samples are fed into the network in a mini-batch way, where they are often regarded of equal importance. However, some of the samples may become less informative during training, as the magnitude of the gradient start to vanish for these samples. In the meantime, other samples of higher utility or hardness may be more demanded for the training process to proceed and require more exploitation. To address the challenges of expensive annotations and loss of sample informativeness, here we propose a novel training framework which adaptively selects informative samples that are fed to the training process. The adaptive selection or sampling is performed based on a hardness-aware strategy in the latent space constructed by a generative model. To evaluate the proposed training framework, we perform experiments on three different datasets, including MNIST and CIFAR-10 for image classification task and a medical image dataset IVUS for biophysical simulation task. On all three datasets, the proposed framework outperforms a random sampling method, which demonstrates the effectiveness of proposed framework.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge