Zhiyi Peng
Multi-phase Liver Tumor Segmentation with Spatial Aggregation and Uncertain Region Inpainting
Aug 05, 2021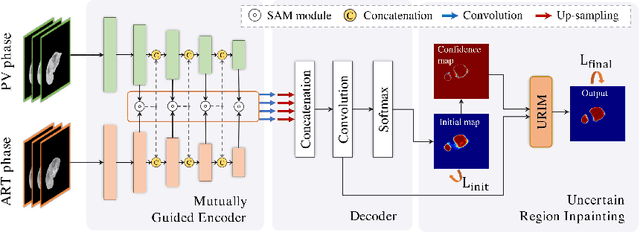
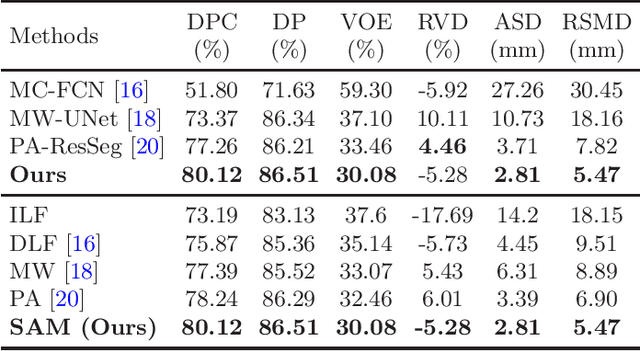
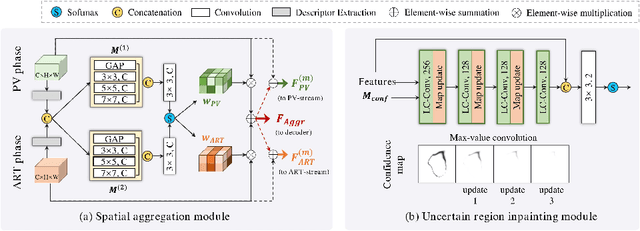
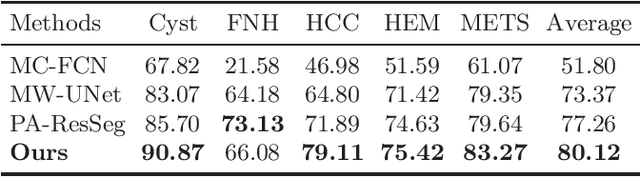
Abstract:Multi-phase computed tomography (CT) images provide crucial complementary information for accurate liver tumor segmentation (LiTS). State-of-the-art multi-phase LiTS methods usually fused cross-phase features through phase-weighted summation or channel-attention based concatenation. However, these methods ignored the spatial (pixel-wise) relationships between different phases, hence leading to insufficient feature integration. In addition, the performance of existing methods remains subject to the uncertainty in segmentation, which is particularly acute in tumor boundary regions. In this work, we propose a novel LiTS method to adequately aggregate multi-phase information and refine uncertain region segmentation. To this end, we introduce a spatial aggregation module (SAM), which encourages per-pixel interactions between different phases, to make full use of cross-phase information. Moreover, we devise an uncertain region inpainting module (URIM) to refine uncertain pixels using neighboring discriminative features. Experiments on an in-house multi-phase CT dataset of focal liver lesions (MPCT-FLLs) demonstrate that our method achieves promising liver tumor segmentation and outperforms state-of-the-arts.
Automatic CT Segmentation from Bounding Box Annotations using Convolutional Neural Networks
Jun 09, 2021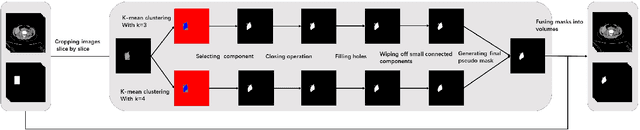
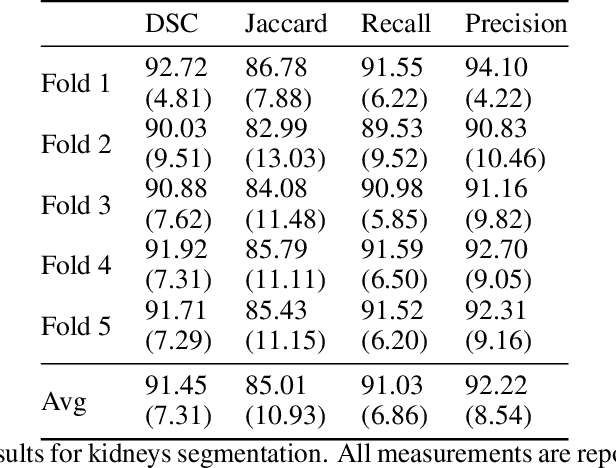
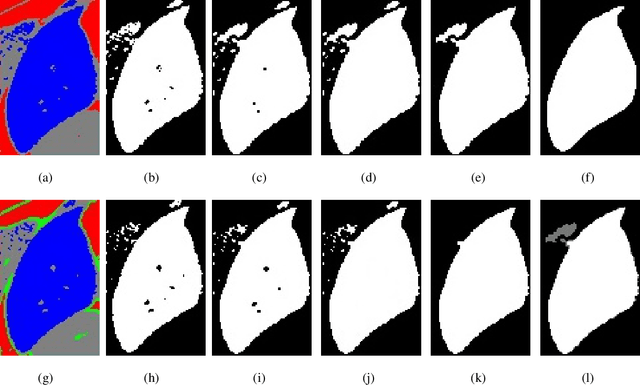
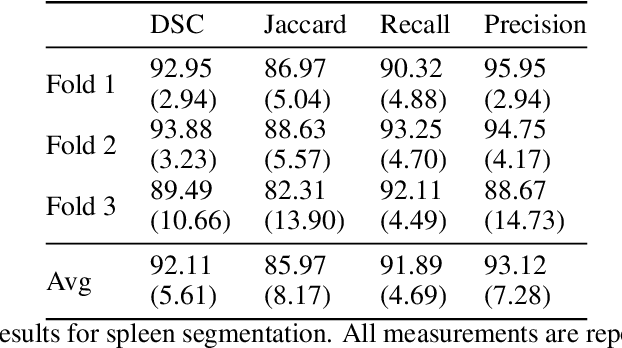
Abstract:Accurate segmentation for medical images is important for clinical diagnosis. Existing automatic segmentation methods are mainly based on fully supervised learning and have an extremely high demand for precise annotations, which are very costly and time-consuming to obtain. To address this problem, we proposed an automatic CT segmentation method based on weakly supervised learning, by which one could train an accurate segmentation model only with weak annotations in the form of bounding boxes. The proposed method is composed of two steps: 1) generating pseudo masks with bounding box annotations by k-means clustering, and 2) iteratively training a 3D U-Net convolutional neural network as a segmentation model. Some data pre-processing methods are used to improve performance. The method was validated on four datasets containing three types of organs with a total of 627 CT volumes. For liver, spleen and kidney segmentation, it achieved an accuracy of 95.19%, 92.11%, and 91.45%, respectively. Experimental results demonstrate that our method is accurate, efficient, and suitable for clinical use.
Graph-based Pyramid Global Context Reasoning with a Saliency-aware Projection for COVID-19 Lung Infections Segmentation
Mar 07, 2021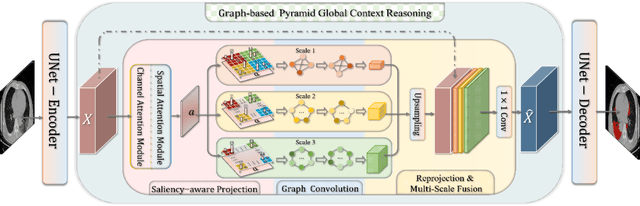
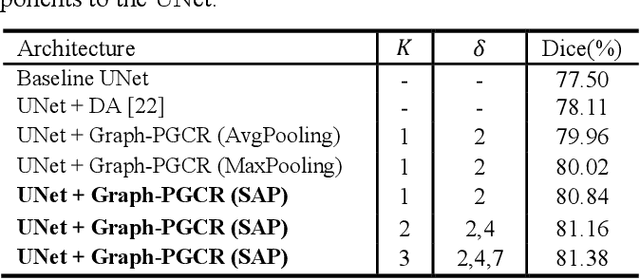
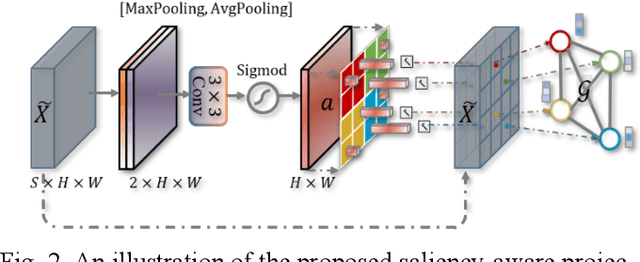
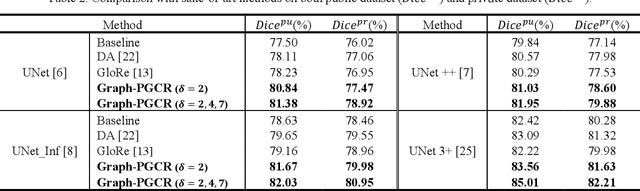
Abstract:Coronavirus Disease 2019 (COVID-19) has rapidly spread in 2020, emerging a mass of studies for lung infection segmentation from CT images. Though many methods have been proposed for this issue, it is a challenging task because of infections of various size appearing in different lobe zones. To tackle these issues, we propose a Graph-based Pyramid Global Context Reasoning (Graph-PGCR) module, which is capable of modeling long-range dependencies among disjoint infections as well as adapt size variation. We first incorporate graph convolution to exploit long-term contextual information from multiple lobe zones. Different from previous average pooling or maximum object probability, we propose a saliency-aware projection mechanism to pick up infection-related pixels as a set of graph nodes. After graph reasoning, the relation-aware features are reversed back to the original coordinate space for the down-stream tasks. We further construct multiple graphs with different sampling rates to handle the size variation problem. To this end, distinct multi-scale long-range contextual patterns can be captured. Our Graph-PGCR module is plug-and-play, which can be integrated into any architecture to improve its performance. Experiments demonstrated that the proposed method consistently boost the performance of state-of-the-art backbone architectures on both of public and our private COVID-19 datasets.
PA-ResSeg: A Phase Attention Residual Network for Liver Tumor Segmentation from Multi-phase CT Images
Feb 27, 2021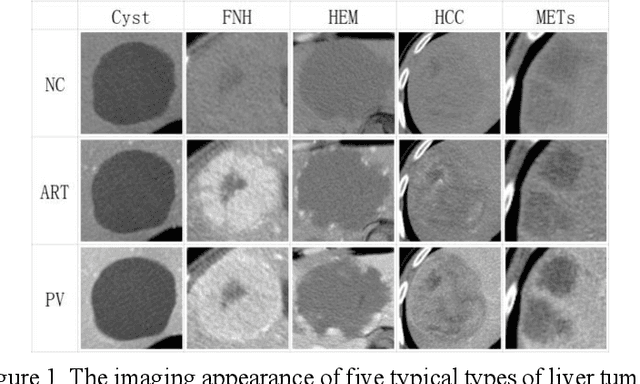
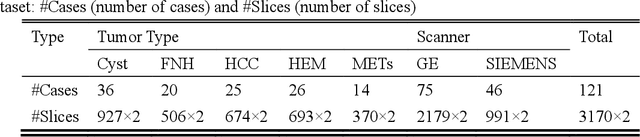
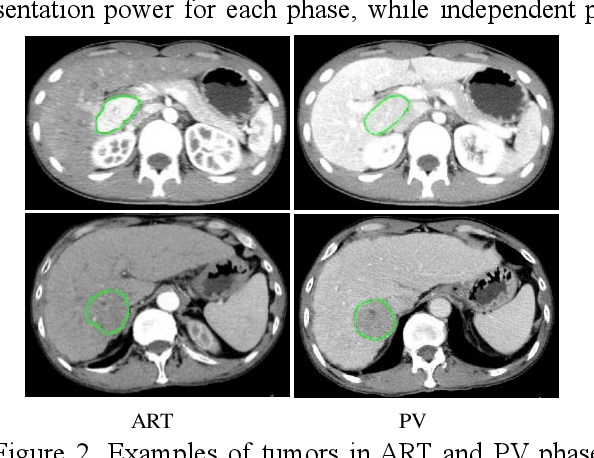
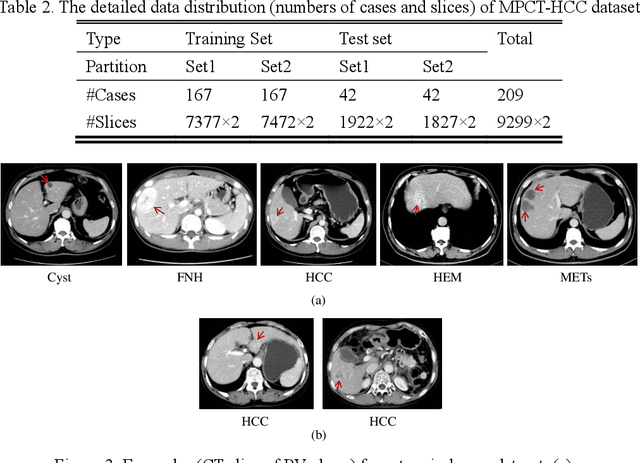
Abstract:In this paper, we propose a phase attention residual network (PA-ResSeg) to model multi-phase features for accurate liver tumor segmentation, in which a phase attention (PA) is newly proposed to additionally exploit the images of arterial (ART) phase to facilitate the segmentation of portal venous (PV) phase. The PA block consists of an intra-phase attention (Intra-PA) module and an inter-phase attention (Inter-PA) module to capture channel-wise self-dependencies and cross-phase interdependencies, respectively. Thus it enables the network to learn more representative multi-phase features by refining the PV features according to the channel dependencies and recalibrating the ART features based on the learned interdependencies between phases. We propose a PA-based multi-scale fusion (MSF) architecture to embed the PA blocks in the network at multiple levels along the encoding path to fuse multi-scale features from multi-phase images. Moreover, a 3D boundary-enhanced loss (BE-loss) is proposed for training to make the network more sensitive to boundaries. To evaluate the performance of our proposed PA-ResSeg, we conducted experiments on a multi-phase CT dataset of focal liver lesions (MPCT-FLLs). Experimental results show the effectiveness of the proposed method by achieving a dice per case (DPC) of 0.77.87, a dice global (DG) of 0.8682, a volumetric overlap error (VOE) of 0.3328 and a relative volume difference (RVD) of 0.0443 on the MPCT-FLLs. Furthermore, to validate the effectiveness and robustness of PA-ResSeg, we conducted extra experiments on another multi-phase liver tumor dataset and obtained a DPC of 0.8290, a DG of 0.9132, a VOE of 0.2637 and a RVD of 0.0163. The proposed method shows its robustness and generalization capability in different datasets and different backbones.
Automatic 3D liver location and segmentation via convolutional neural networks and graph cut
May 10, 2016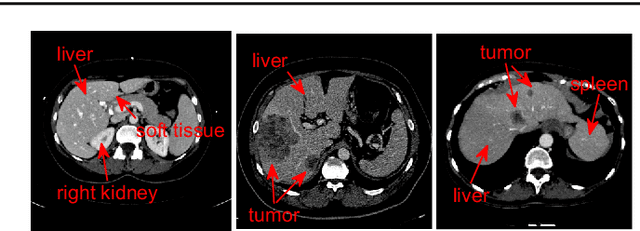
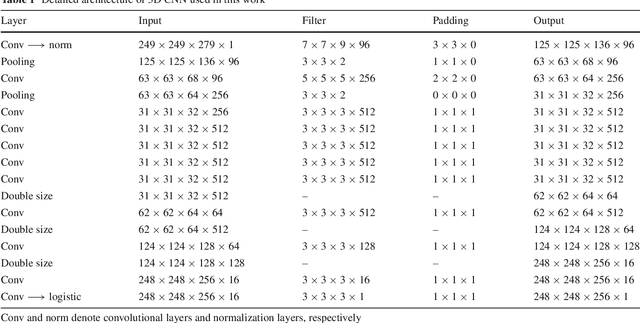
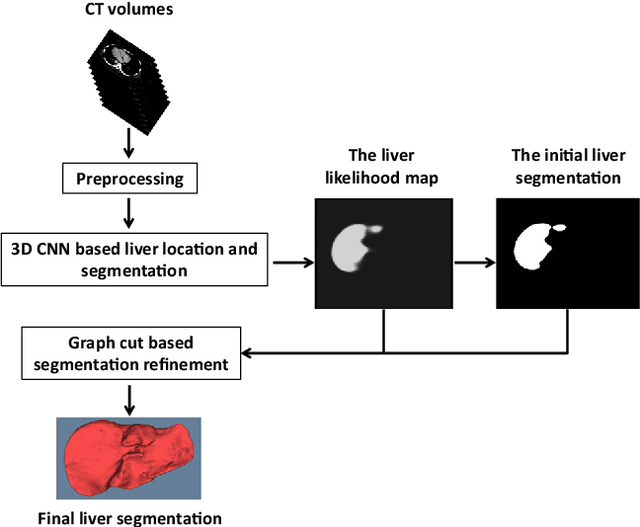
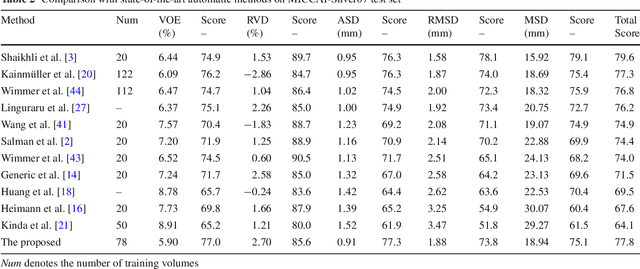
Abstract:Purpose Segmentation of the liver from abdominal computed tomography (CT) image is an essential step in some computer assisted clinical interventions, such as surgery planning for living donor liver transplant (LDLT), radiotherapy and volume measurement. In this work, we develop a deep learning algorithm with graph cut refinement to automatically segment liver in CT scans. Methods The proposed method consists of two main steps: (i) simultaneously liver detection and probabilistic segmentation using 3D convolutional neural networks (CNNs); (ii) accuracy refinement of initial segmentation with graph cut and the previously learned probability map. Results The proposed approach was validated on forty CT volumes taken from two public databases MICCAI-Sliver07 and 3Dircadb. For the MICCAI-Sliver07 test set, the calculated mean ratios of volumetric overlap error (VOE), relative volume difference (RVD), average symmetric surface distance (ASD), root mean square symmetric surface distance (RMSD) and maximum symmetric surface distance (MSD) are 5.9%, 2.7%, 0.91%, 1.88 mm, and 18.94 mm, respectively. In the case of 20 3Dircadb data, the calculated mean ratios of VOE, RVD, ASD, RMSD and MSD are 9.36%, 0.97%, 1.89%, 4.15 mm and 33.14 mm, respectively. Conclusion The proposed method is fully automatic without any user interaction. Quantitative results reveal that the proposed approach is efficient and accurate for hepatic volume estimation in a clinical setup. The high correlation between the automatic and manual references shows that the proposed method can be good enough to replace the time-consuming and non-reproducible manual segmentation method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge