Ujas Parikh
3D-GMIC: an efficient deep neural network to find small objects in large 3D images
Oct 16, 2022
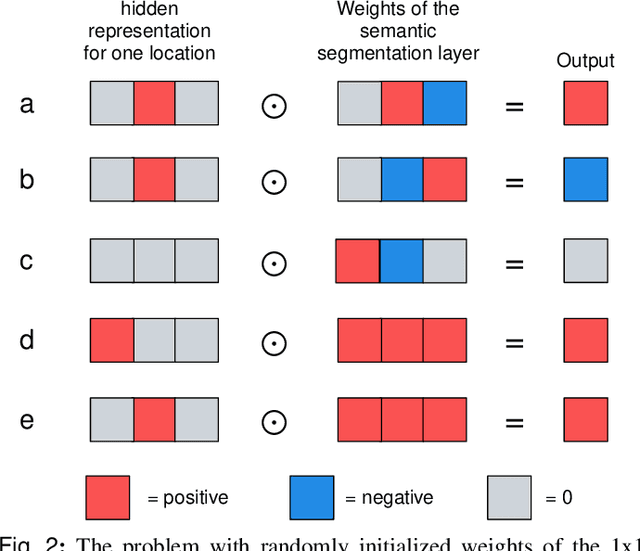
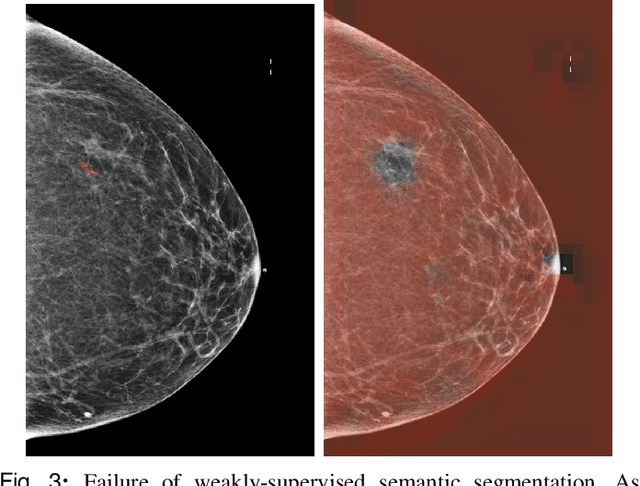
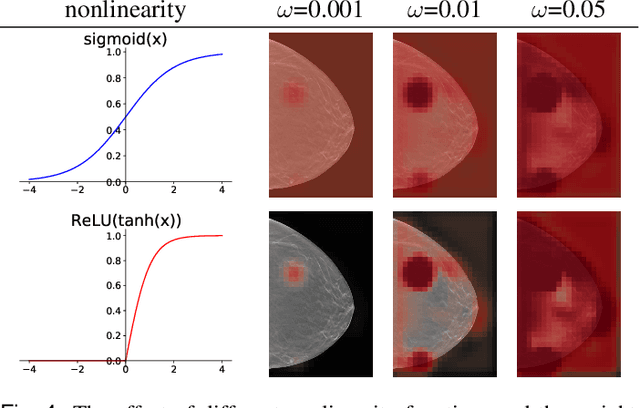
Abstract:3D imaging enables a more accurate diagnosis by providing spatial information about organ anatomy. However, using 3D images to train AI models is computationally challenging because they consist of tens or hundreds of times more pixels than their 2D counterparts. To train with high-resolution 3D images, convolutional neural networks typically resort to downsampling them or projecting them to two dimensions. In this work, we propose an effective alternative, a novel neural network architecture that enables computationally efficient classification of 3D medical images in their full resolution. Compared to off-the-shelf convolutional neural networks, 3D-GMIC uses 77.98%-90.05% less GPU memory and 91.23%-96.02% less computation. While our network is trained only with image-level labels, without segmentation labels, it explains its classification predictions by providing pixel-level saliency maps. On a dataset collected at NYU Langone Health, including 85,526 patients with full-field 2D mammography (FFDM), synthetic 2D mammography, and 3D mammography (DBT), our model, the 3D Globally-Aware Multiple Instance Classifier (3D-GMIC), achieves a breast-wise AUC of 0.831 (95% CI: 0.769-0.887) in classifying breasts with malignant findings using DBT images. As DBT and 2D mammography capture different information, averaging predictions on 2D and 3D mammography together leads to a diverse ensemble with an improved breast-wise AUC of 0.841 (95% CI: 0.768-0.895). Our model generalizes well to an external dataset from Duke University Hospital, achieving an image-wise AUC of 0.848 (95% CI: 0.798-0.896) in classifying DBT images with malignant findings.
Deep Neural Networks Improve Radiologists' Performance in Breast Cancer Screening
Mar 20, 2019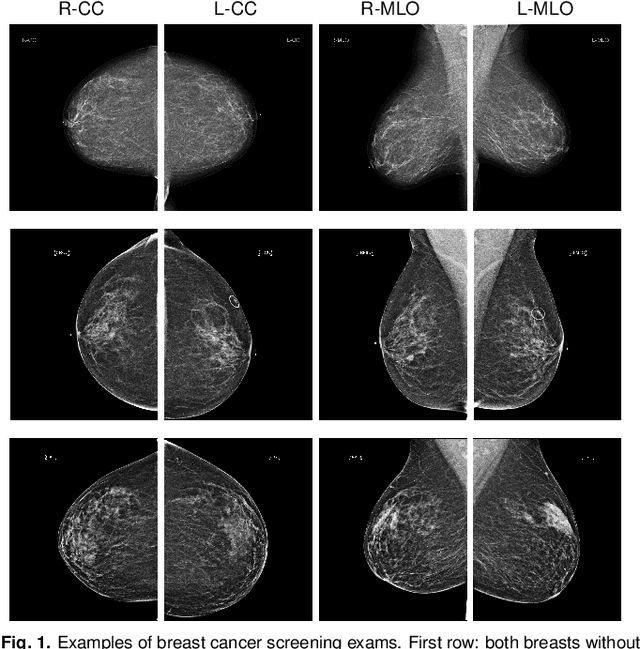
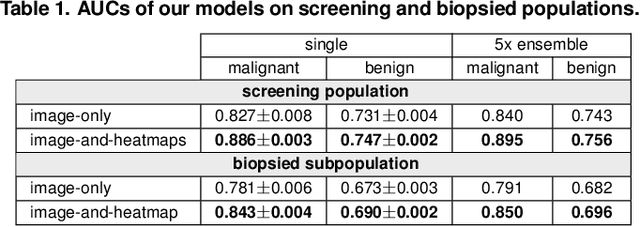
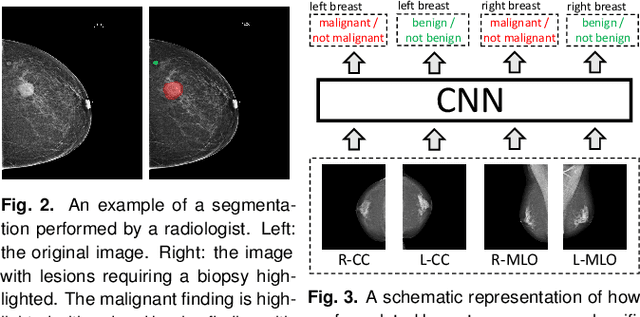
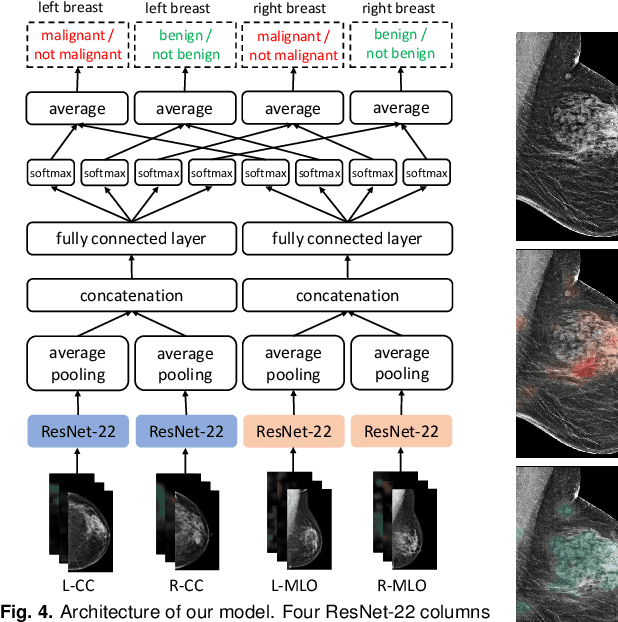
Abstract:We present a deep convolutional neural network for breast cancer screening exam classification, trained and evaluated on over 200,000 exams (over 1,000,000 images). Our network achieves an AUC of 0.895 in predicting whether there is a cancer in the breast, when tested on the screening population. We attribute the high accuracy of our model to a two-stage training procedure, which allows us to use a very high-capacity patch-level network to learn from pixel-level labels alongside a network learning from macroscopic breast-level labels. To validate our model, we conducted a reader study with 14 readers, each reading 720 screening mammogram exams, and find our model to be as accurate as experienced radiologists when presented with the same data. Finally, we show that a hybrid model, averaging probability of malignancy predicted by a radiologist with a prediction of our neural network, is more accurate than either of the two separately. To better understand our results, we conduct a thorough analysis of our network's performance on different subpopulations of the screening population, model design, training procedure, errors, and properties of its internal representations.
High-Resolution Breast Cancer Screening with Multi-View Deep Convolutional Neural Networks
Jun 28, 2018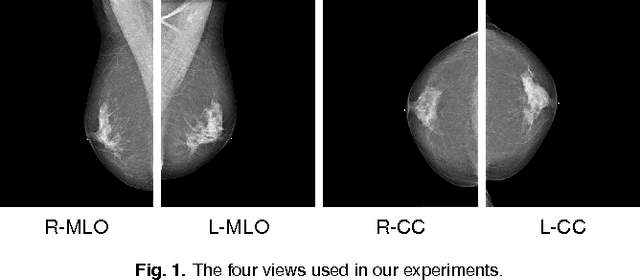
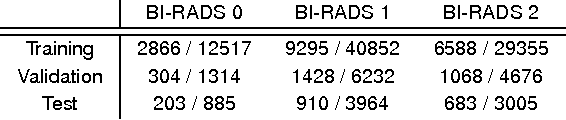
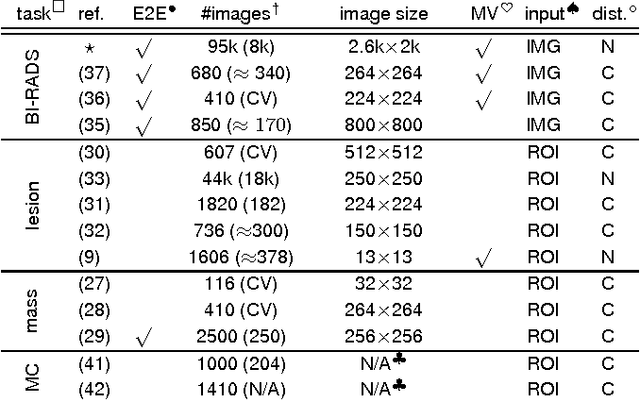
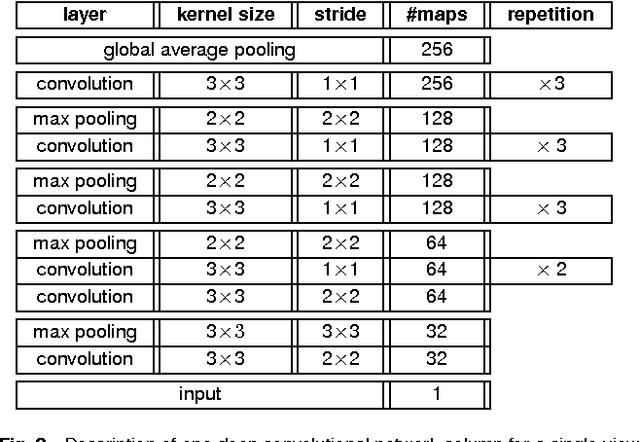

Abstract:Advances in deep learning for natural images have prompted a surge of interest in applying similar techniques to medical images. The majority of the initial attempts focused on replacing the input of a deep convolutional neural network with a medical image, which does not take into consideration the fundamental differences between these two types of images. Specifically, fine details are necessary for detection in medical images, unlike in natural images where coarse structures matter most. This difference makes it inadequate to use the existing network architectures developed for natural images, because they work on heavily downscaled images to reduce the memory requirements. This hides details necessary to make accurate predictions. Additionally, a single exam in medical imaging often comes with a set of views which must be fused in order to reach a correct conclusion. In our work, we propose to use a multi-view deep convolutional neural network that handles a set of high-resolution medical images. We evaluate it on large-scale mammography-based breast cancer screening (BI-RADS prediction) using 886,000 images. We focus on investigating the impact of the training set size and image size on the prediction accuracy. Our results highlight that performance increases with the size of training set, and that the best performance can only be achieved using the original resolution. In the reader study, performed on a random subset of the test set, we confirmed the efficacy of our model, which achieved performance comparable to a committee of radiologists when presented with the same data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge