Linda Du
3D-GMIC: an efficient deep neural network to find small objects in large 3D images
Oct 16, 2022
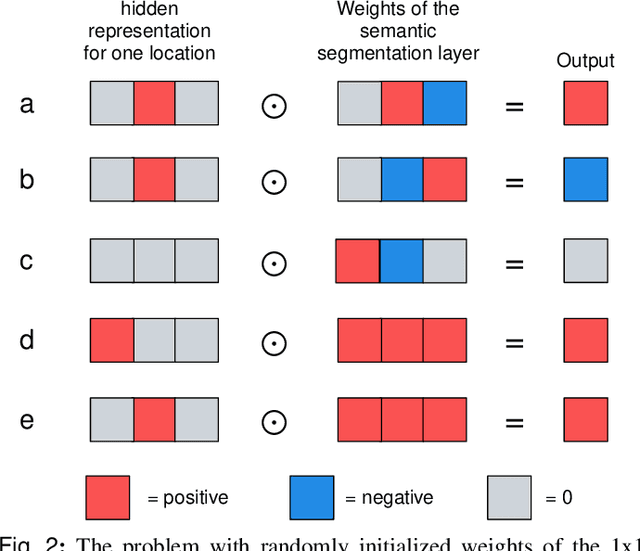
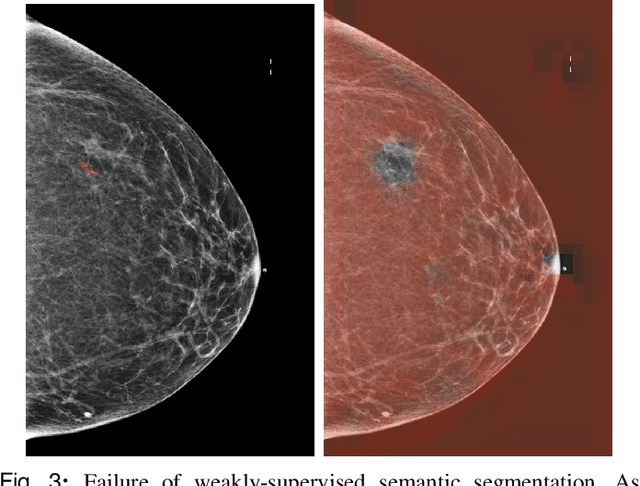
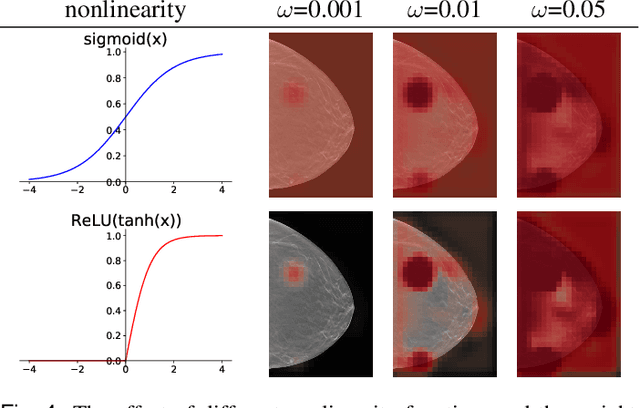
Abstract:3D imaging enables a more accurate diagnosis by providing spatial information about organ anatomy. However, using 3D images to train AI models is computationally challenging because they consist of tens or hundreds of times more pixels than their 2D counterparts. To train with high-resolution 3D images, convolutional neural networks typically resort to downsampling them or projecting them to two dimensions. In this work, we propose an effective alternative, a novel neural network architecture that enables computationally efficient classification of 3D medical images in their full resolution. Compared to off-the-shelf convolutional neural networks, 3D-GMIC uses 77.98%-90.05% less GPU memory and 91.23%-96.02% less computation. While our network is trained only with image-level labels, without segmentation labels, it explains its classification predictions by providing pixel-level saliency maps. On a dataset collected at NYU Langone Health, including 85,526 patients with full-field 2D mammography (FFDM), synthetic 2D mammography, and 3D mammography (DBT), our model, the 3D Globally-Aware Multiple Instance Classifier (3D-GMIC), achieves a breast-wise AUC of 0.831 (95% CI: 0.769-0.887) in classifying breasts with malignant findings using DBT images. As DBT and 2D mammography capture different information, averaging predictions on 2D and 3D mammography together leads to a diverse ensemble with an improved breast-wise AUC of 0.841 (95% CI: 0.768-0.895). Our model generalizes well to an external dataset from Duke University Hospital, achieving an image-wise AUC of 0.848 (95% CI: 0.798-0.896) in classifying DBT images with malignant findings.
Differences between human and machine perception in medical diagnosis
Nov 28, 2020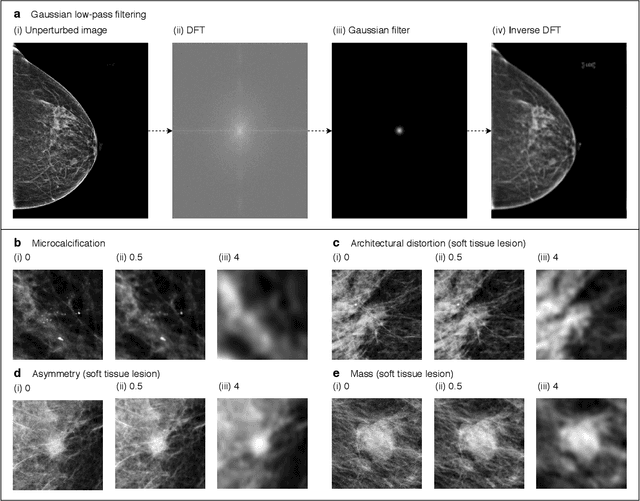
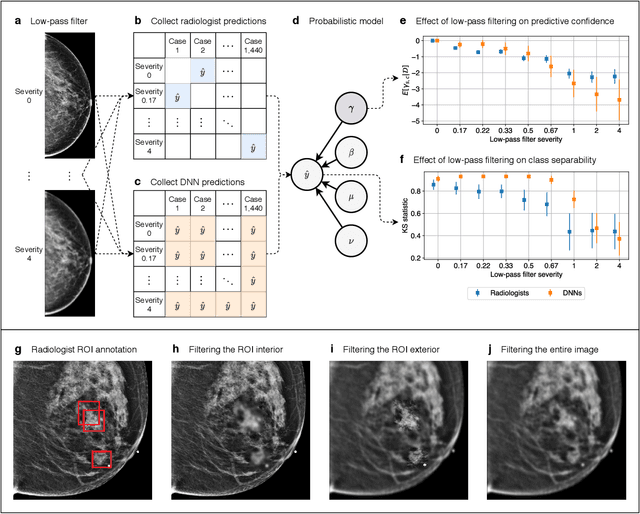
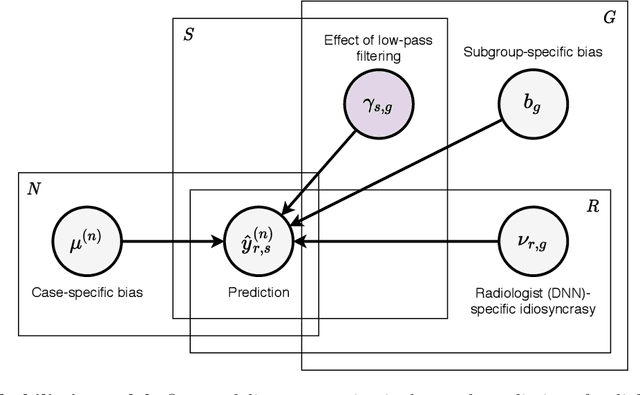
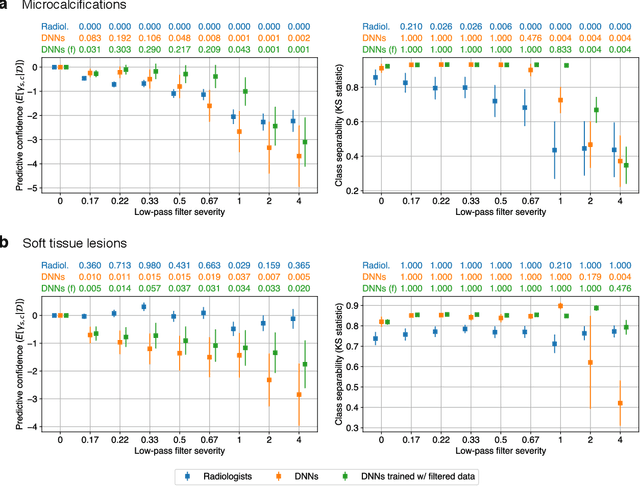
Abstract:Deep neural networks (DNNs) show promise in image-based medical diagnosis, but cannot be fully trusted since their performance can be severely degraded by dataset shifts to which human perception remains invariant. If we can better understand the differences between human and machine perception, we can potentially characterize and mitigate this effect. We therefore propose a framework for comparing human and machine perception in medical diagnosis. The two are compared with respect to their sensitivity to the removal of clinically meaningful information, and to the regions of an image deemed most suspicious. Drawing inspiration from the natural image domain, we frame both comparisons in terms of perturbation robustness. The novelty of our framework is that separate analyses are performed for subgroups with clinically meaningful differences. We argue that this is necessary in order to avert Simpson's paradox and draw correct conclusions. We demonstrate our framework with a case study in breast cancer screening, and reveal significant differences between radiologists and DNNs. We compare the two with respect to their robustness to Gaussian low-pass filtering, performing a subgroup analysis on microcalcifications and soft tissue lesions. For microcalcifications, DNNs use a separate set of high frequency components than radiologists, some of which lie outside the image regions considered most suspicious by radiologists. These features run the risk of being spurious, but if not, could represent potential new biomarkers. For soft tissue lesions, the divergence between radiologists and DNNs is even starker, with DNNs relying heavily on spurious high frequency components ignored by radiologists. Importantly, this deviation in soft tissue lesions was only observable through subgroup analysis, which highlights the importance of incorporating medical domain knowledge into our comparison framework.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge