Minh Chien Vu
BigCodeBench: Benchmarking Code Generation with Diverse Function Calls and Complex Instructions
Jun 26, 2024



Abstract:Automated software engineering has been greatly empowered by the recent advances in Large Language Models (LLMs) for programming. While current benchmarks have shown that LLMs can perform various software engineering tasks like human developers, the majority of their evaluations are limited to short and self-contained algorithmic tasks. Solving challenging and practical programming tasks requires the capability of utilizing diverse function calls as tools to efficiently implement functionalities like data analysis and web development. In addition, using multiple tools to solve a task needs compositional reasoning by accurately understanding complex instructions. Fulfilling both of these characteristics can pose a great challenge for LLMs. To assess how well LLMs can solve challenging and practical programming tasks, we introduce Bench, a benchmark that challenges LLMs to invoke multiple function calls as tools from 139 libraries and 7 domains for 1,140 fine-grained programming tasks. To evaluate LLMs rigorously, each programming task encompasses 5.6 test cases with an average branch coverage of 99%. In addition, we propose a natural-language-oriented variant of Bench, Benchi, that automatically transforms the original docstrings into short instructions only with essential information. Our extensive evaluation of 60 LLMs shows that LLMs are not yet capable of following complex instructions to use function calls precisely, with scores up to 60%, significantly lower than the human performance of 97%. The results underscore the need for further advancements in this area.
The BigScience ROOTS Corpus: A 1.6TB Composite Multilingual Dataset
Mar 07, 2023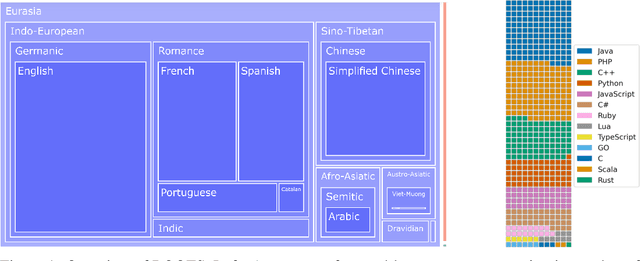

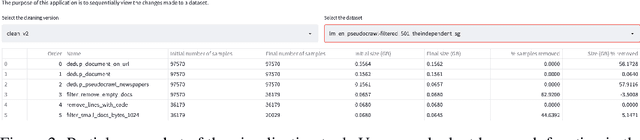

Abstract:As language models grow ever larger, the need for large-scale high-quality text datasets has never been more pressing, especially in multilingual settings. The BigScience workshop, a 1-year international and multidisciplinary initiative, was formed with the goal of researching and training large language models as a values-driven undertaking, putting issues of ethics, harm, and governance in the foreground. This paper documents the data creation and curation efforts undertaken by BigScience to assemble the Responsible Open-science Open-collaboration Text Sources (ROOTS) corpus, a 1.6TB dataset spanning 59 languages that was used to train the 176-billion-parameter BigScience Large Open-science Open-access Multilingual (BLOOM) language model. We further release a large initial subset of the corpus and analyses thereof, and hope to empower large-scale monolingual and multilingual modeling projects with both the data and the processing tools, as well as stimulate research around this large multilingual corpus.
BLOOM: A 176B-Parameter Open-Access Multilingual Language Model
Nov 09, 2022Abstract:Large language models (LLMs) have been shown to be able to perform new tasks based on a few demonstrations or natural language instructions. While these capabilities have led to widespread adoption, most LLMs are developed by resource-rich organizations and are frequently kept from the public. As a step towards democratizing this powerful technology, we present BLOOM, a 176B-parameter open-access language model designed and built thanks to a collaboration of hundreds of researchers. BLOOM is a decoder-only Transformer language model that was trained on the ROOTS corpus, a dataset comprising hundreds of sources in 46 natural and 13 programming languages (59 in total). We find that BLOOM achieves competitive performance on a wide variety of benchmarks, with stronger results after undergoing multitask prompted finetuning. To facilitate future research and applications using LLMs, we publicly release our models and code under the Responsible AI License.
BigBIO: A Framework for Data-Centric Biomedical Natural Language Processing
Jun 30, 2022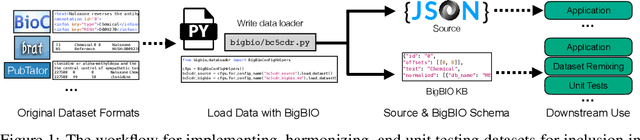
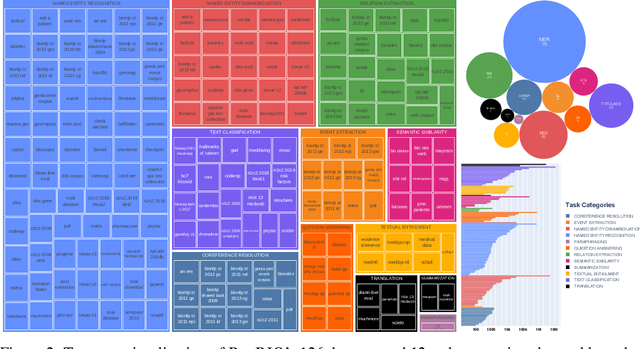
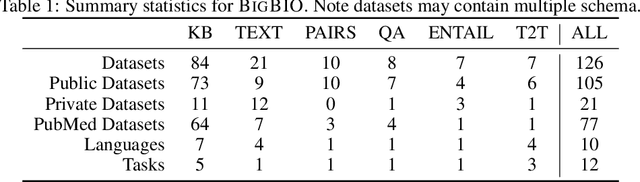

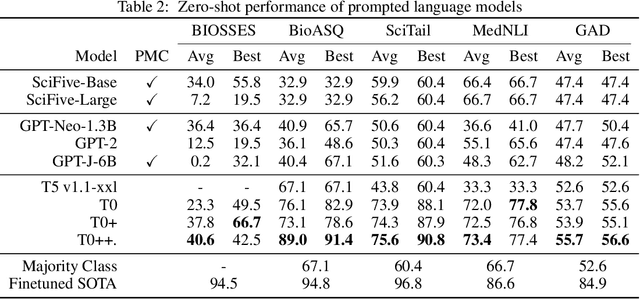
Abstract:Training and evaluating language models increasingly requires the construction of meta-datasets --diverse collections of curated data with clear provenance. Natural language prompting has recently lead to improved zero-shot generalization by transforming existing, supervised datasets into a diversity of novel pretraining tasks, highlighting the benefits of meta-dataset curation. While successful in general-domain text, translating these data-centric approaches to biomedical language modeling remains challenging, as labeled biomedical datasets are significantly underrepresented in popular data hubs. To address this challenge, we introduce BigBIO a community library of 126+ biomedical NLP datasets, currently covering 12 task categories and 10+ languages. BigBIO facilitates reproducible meta-dataset curation via programmatic access to datasets and their metadata, and is compatible with current platforms for prompt engineering and end-to-end few/zero shot language model evaluation. We discuss our process for task schema harmonization, data auditing, contribution guidelines, and outline two illustrative use cases: zero-shot evaluation of biomedical prompts and large-scale, multi-task learning. BigBIO is an ongoing community effort and is available at https://github.com/bigscience-workshop/biomedical
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge