Martin Vallières
for the Image Biomarker Standardisation Initiative
Unraveling Radiomics Complexity: Strategies for Optimal Simplicity in Predictive Modeling
Jul 05, 2024



Abstract:Background: The high dimensionality of radiomic feature sets, the variability in radiomic feature types and potentially high computational requirements all underscore the need for an effective method to identify the smallest set of predictive features for a given clinical problem. Purpose: Develop a methodology and tools to identify and explain the smallest set of predictive radiomic features. Materials and Methods: 89,714 radiomic features were extracted from five cancer datasets: low-grade glioma, meningioma, non-small cell lung cancer (NSCLC), and two renal cell carcinoma cohorts (n=2104). Features were categorized by computational complexity into morphological, intensity, texture, linear filters, and nonlinear filters. Models were trained and evaluated on each complexity level using the area under the curve (AUC). The most informative features were identified, and their importance was explained. The optimal complexity level and associated most informative features were identified using systematic statistical significance analyses and a false discovery avoidance procedure, respectively. Their predictive importance was explained using a novel tree-based method. Results: MEDimage, a new open-source tool, was developed to facilitate radiomic studies. Morphological features were optimal for MRI-based meningioma (AUC: 0.65) and low-grade glioma (AUC: 0.68). Intensity features were optimal for CECT-based renal cell carcinoma (AUC: 0.82) and CT-based NSCLC (AUC: 0.76). Texture features were optimal for MRI-based renal cell carcinoma (AUC: 0.72). Tuning the Hounsfield unit range improved results for CECT-based renal cell carcinoma (AUC: 0.86). Conclusion: Our proposed methodology and software can estimate the optimal radiomics complexity level for specific medical outcomes, potentially simplifying the use of radiomics in predictive modeling across various contexts.
Federated Learning Enables Big Data for Rare Cancer Boundary Detection
Apr 25, 2022Abstract:Although machine learning (ML) has shown promise in numerous domains, there are concerns about generalizability to out-of-sample data. This is currently addressed by centrally sharing ample, and importantly diverse, data from multiple sites. However, such centralization is challenging to scale (or even not feasible) due to various limitations. Federated ML (FL) provides an alternative to train accurate and generalizable ML models, by only sharing numerical model updates. Here we present findings from the largest FL study to-date, involving data from 71 healthcare institutions across 6 continents, to generate an automatic tumor boundary detector for the rare disease of glioblastoma, utilizing the largest dataset of such patients ever used in the literature (25,256 MRI scans from 6,314 patients). We demonstrate a 33% improvement over a publicly trained model to delineate the surgically targetable tumor, and 23% improvement over the tumor's entire extent. We anticipate our study to: 1) enable more studies in healthcare informed by large and diverse data, ensuring meaningful results for rare diseases and underrepresented populations, 2) facilitate further quantitative analyses for glioblastoma via performance optimization of our consensus model for eventual public release, and 3) demonstrate the effectiveness of FL at such scale and task complexity as a paradigm shift for multi-site collaborations, alleviating the need for data sharing.
Overview of the HECKTOR Challenge at MICCAI 2021: Automatic Head and Neck Tumor Segmentation and Outcome Prediction in PET/CT Images
Jan 11, 2022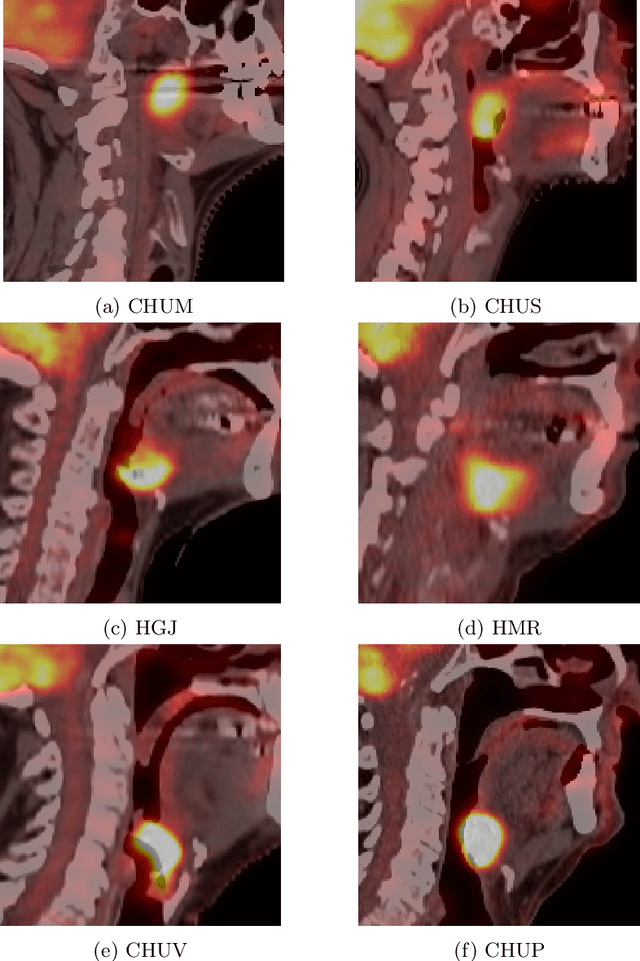
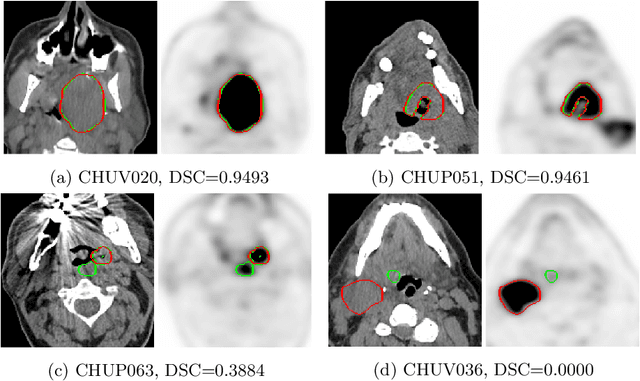
Abstract:This paper presents an overview of the second edition of the HEad and neCK TumOR (HECKTOR) challenge, organized as a satellite event of the 24th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI) 2021. The challenge is composed of three tasks related to the automatic analysis of PET/CT images for patients with Head and Neck cancer (H&N), focusing on the oropharynx region. Task 1 is the automatic segmentation of H&N primary Gross Tumor Volume (GTVt) in FDG-PET/CT images. Task 2 is the automatic prediction of Progression Free Survival (PFS) from the same FDG-PET/CT. Finally, Task 3 is the same as Task 2 with ground truth GTVt annotations provided to the participants. The data were collected from six centers for a total of 325 images, split into 224 training and 101 testing cases. The interest in the challenge was highlighted by the important participation with 103 registered teams and 448 result submissions. The best methods obtained a Dice Similarity Coefficient (DSC) of 0.7591 in the first task, and a Concordance index (C-index) of 0.7196 and 0.6978 in Tasks 2 and 3, respectively. In all tasks, simplicity of the approach was found to be key to ensure generalization performance. The comparison of the PFS prediction performance in Tasks 2 and 3 suggests that providing the GTVt contour was not crucial to achieve best results, which indicates that fully automatic methods can be used. This potentially obviates the need for GTVt contouring, opening avenues for reproducible and large scale radiomics studies including thousands potential subjects.
Standardised convolutional filtering for radiomics
Jun 09, 2020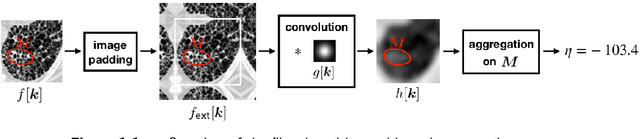
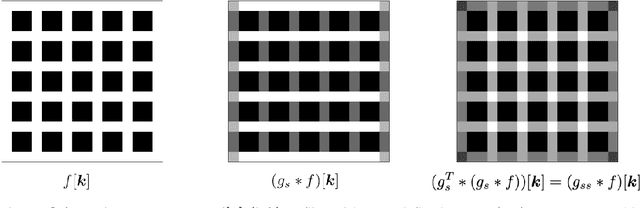
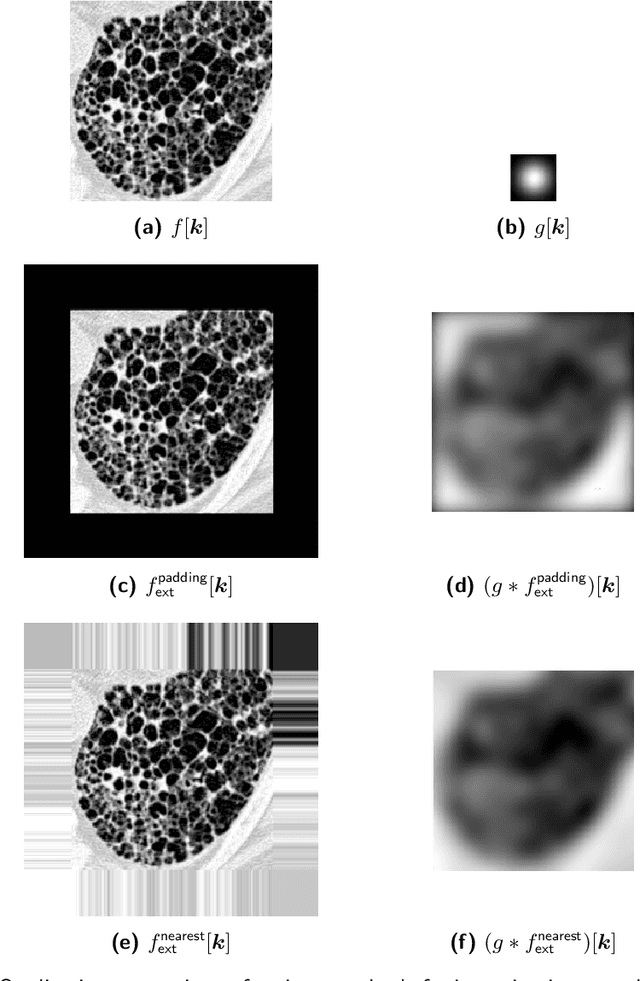


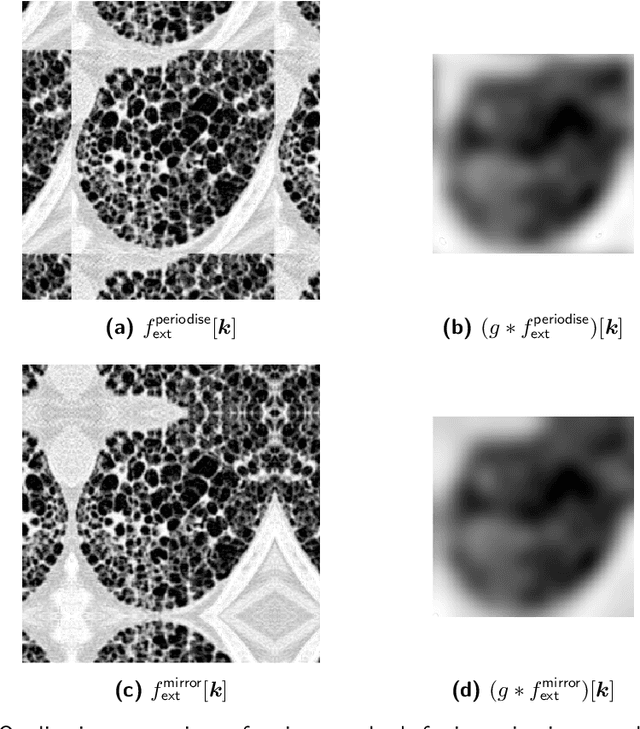
Abstract:The Image Biomarker Standardisation Initiative (IBSI) aims to improve reproducibility of radiomics studies by standardising the computational process of extracting image biomarkers (features) from images. We have previously established reference values for 169 commonly used features, created a standard radiomics image processing scheme, and developed reporting guidelines for radiomic studies. However, several aspects are not standardised. Here we present a preliminary version of a reference manual on the use of convolutional image filters in radiomics. Filters, such as wavelets or Laplacian of Gaussian filters, play an important part in emphasising specific image characteristics such as edges and blobs. Features derived from filter response maps have been found to be poorly reproducible. This reference manual forms the basis of ongoing work on standardising convolutional filters in radiomics, and will be updated as this work progresses.
Image biomarker standardisation initiative
Sep 17, 2018
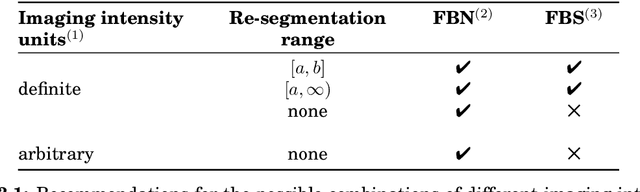
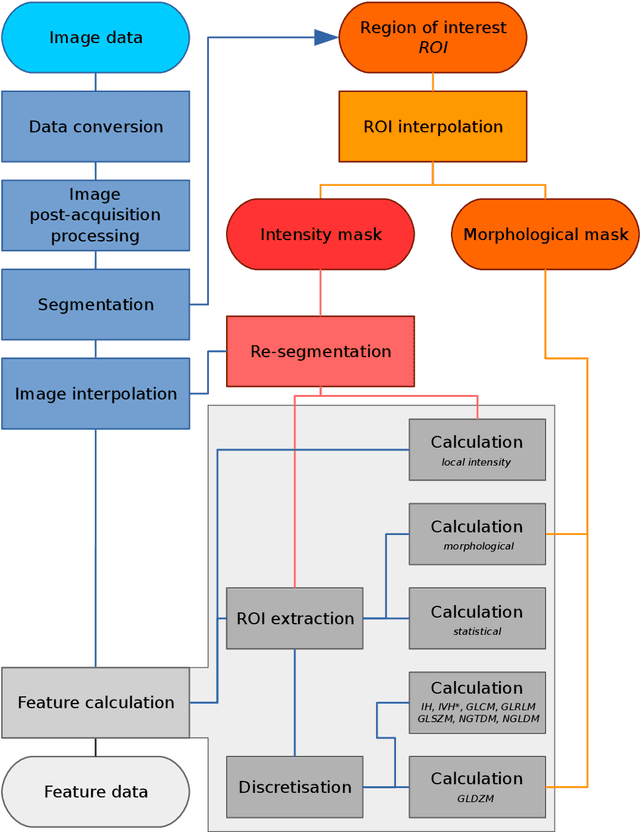

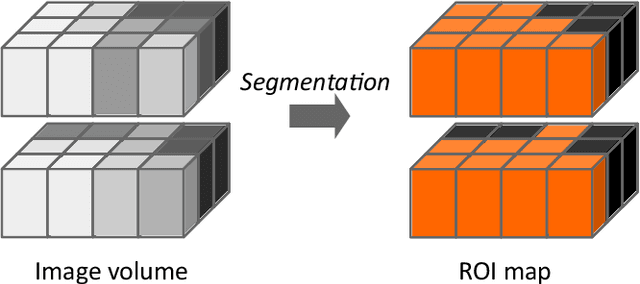
Abstract:The image biomarker standardisation initiative (IBSI) is an independent international collaboration which works towards standardising the extraction of image biomarkers from acquired imaging for the purpose of high-throughput quantitative image analysis (radiomics). Lack of reproducibility and validation of high-throughput quantitative image analysis studies is considered to be a major challenge for the field. Part of this challenge lies in the scantiness of consensus-based guidelines and definitions for the process of translating acquired imaging into high-throughput image biomarkers. The IBSI therefore seeks to provide image biomarker nomenclature and definitions, benchmark data sets, and benchmark values to verify image processing and image biomarker calculations, as well as reporting guidelines, for high-throughput image analysis.
Radiomics strategies for risk assessment of tumour failure in head-and-neck cancer
Mar 24, 2017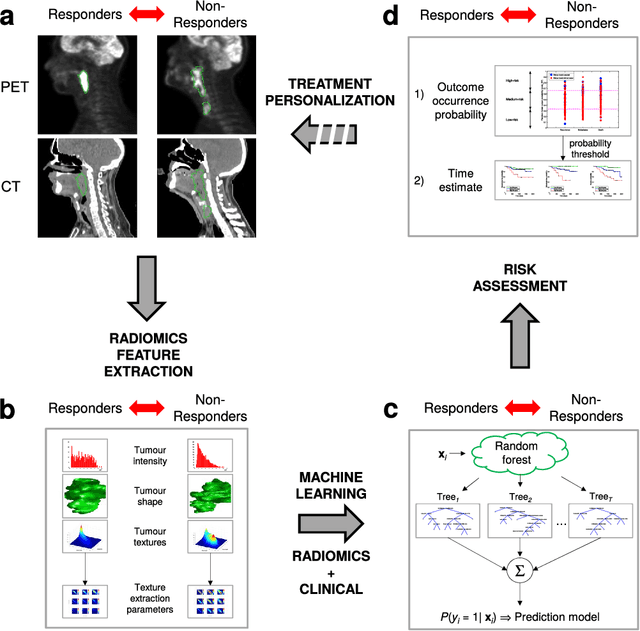
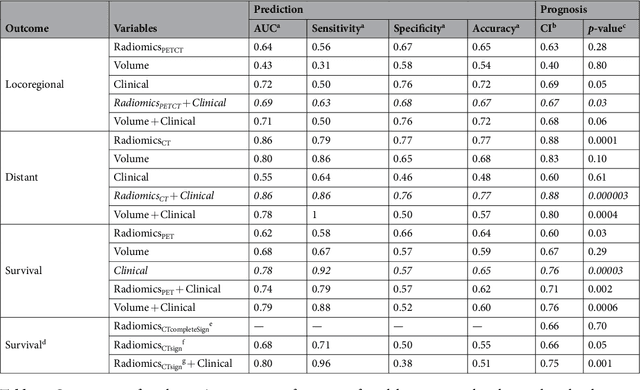
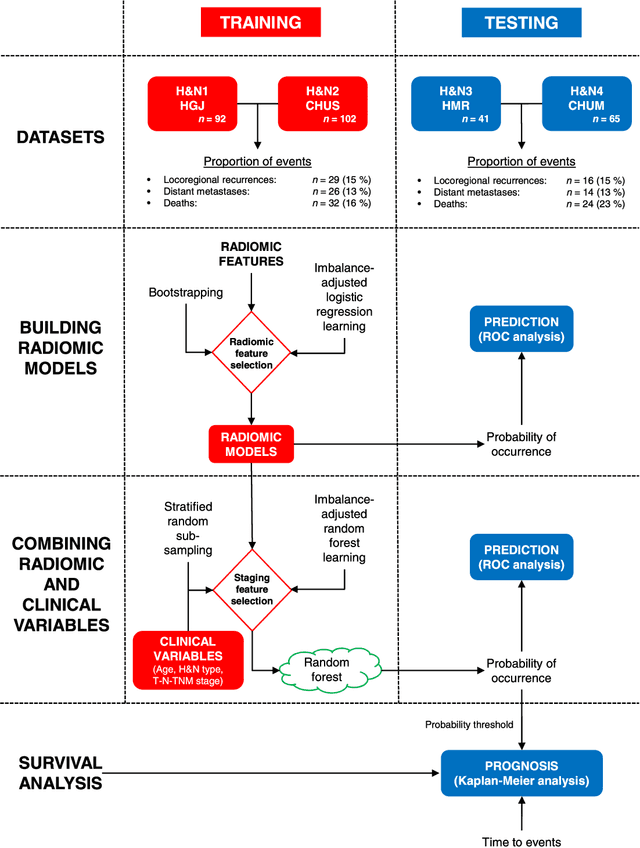
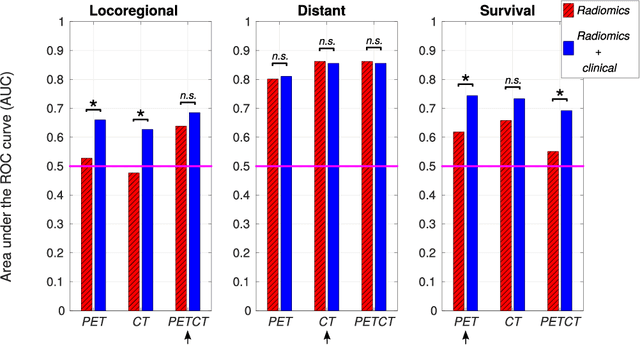
Abstract:Quantitative extraction of high-dimensional mineable data from medical images is a process known as radiomics. Radiomics is foreseen as an essential prognostic tool for cancer risk assessment and the quantification of intratumoural heterogeneity. In this work, 1615 radiomic features (quantifying tumour image intensity, shape, texture) extracted from pre-treatment FDG-PET and CT images of 300 patients from four different cohorts were analyzed for the risk assessment of locoregional recurrences (LR) and distant metastases (DM) in head-and-neck cancer. Prediction models combining radiomic and clinical variables were constructed via random forests and imbalance-adjustment strategies using two of the four cohorts. Independent validation of the prediction and prognostic performance of the models was carried out on the other two cohorts (LR: AUC = 0.69 and CI = 0.67; DM: AUC = 0.86 and CI = 0.88). Furthermore, the results obtained via Kaplan-Meier analysis demonstrated the potential of radiomics for assessing the risk of specific tumour outcomes using multiple stratification groups. This could have important clinical impact, notably by allowing for a better personalization of chemo-radiation treatments for head-and-neck cancer patients from different risk groups.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge