Lingchao Xu
A Global Benchmark of Algorithms for Segmenting Late Gadolinium-Enhanced Cardiac Magnetic Resonance Imaging
May 07, 2020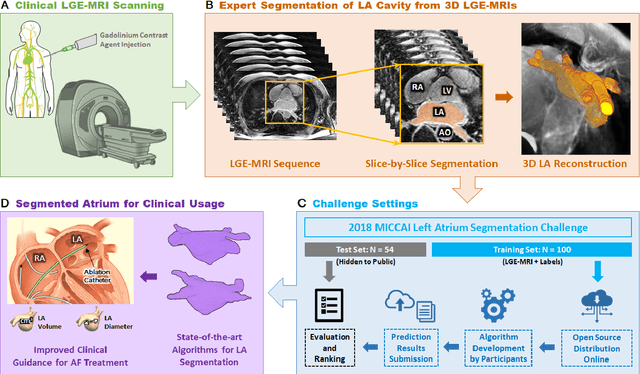
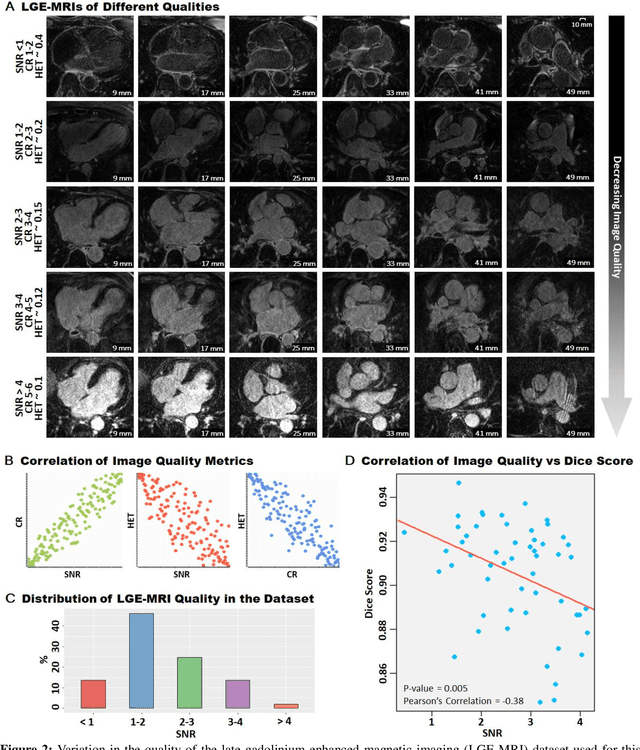
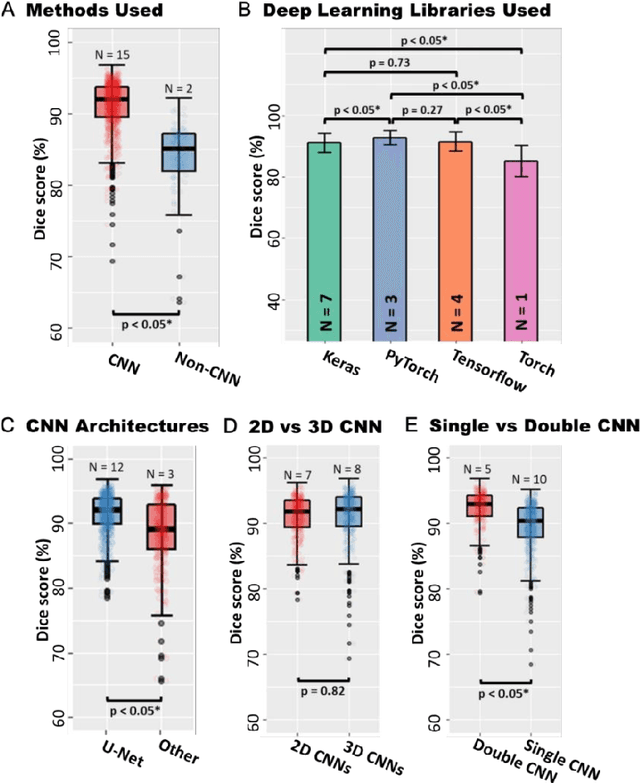
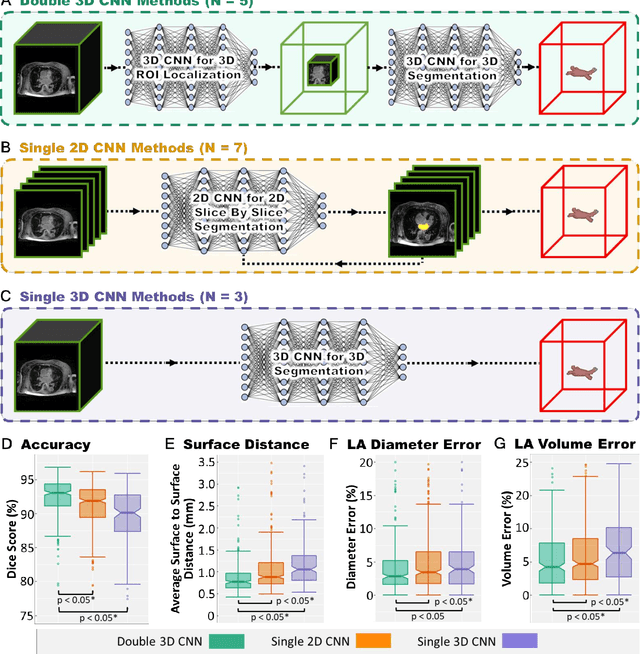
Abstract:Segmentation of cardiac images, particularly late gadolinium-enhanced magnetic resonance imaging (LGE-MRI) widely used for visualizing diseased cardiac structures, is a crucial first step for clinical diagnosis and treatment. However, direct segmentation of LGE-MRIs is challenging due to its attenuated contrast. Since most clinical studies have relied on manual and labor-intensive approaches, automatic methods are of high interest, particularly optimized machine learning approaches. To address this, we organized the "2018 Left Atrium Segmentation Challenge" using 154 3D LGE-MRIs, currently the world's largest cardiac LGE-MRI dataset, and associated labels of the left atrium segmented by three medical experts, ultimately attracting the participation of 27 international teams. In this paper, extensive analysis of the submitted algorithms using technical and biological metrics was performed by undergoing subgroup analysis and conducting hyper-parameter analysis, offering an overall picture of the major design choices of convolutional neural networks (CNNs) and practical considerations for achieving state-of-the-art left atrium segmentation. Results show the top method achieved a dice score of 93.2% and a mean surface to a surface distance of 0.7 mm, significantly outperforming prior state-of-the-art. Particularly, our analysis demonstrated that double, sequentially used CNNs, in which a first CNN is used for automatic region-of-interest localization and a subsequent CNN is used for refined regional segmentation, achieved far superior results than traditional methods and pipelines containing single CNNs. This large-scale benchmarking study makes a significant step towards much-improved segmentation methods for cardiac LGE-MRIs, and will serve as an important benchmark for evaluating and comparing the future works in the field.
Cardiac Segmentation from LGE MRI Using Deep Neural Network Incorporating Shape and Spatial Priors
Jun 25, 2019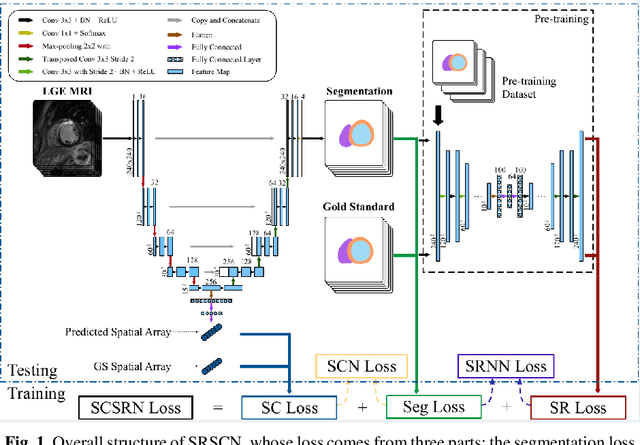
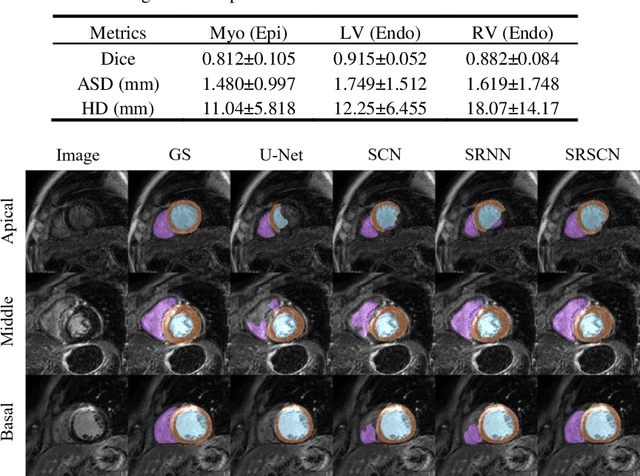
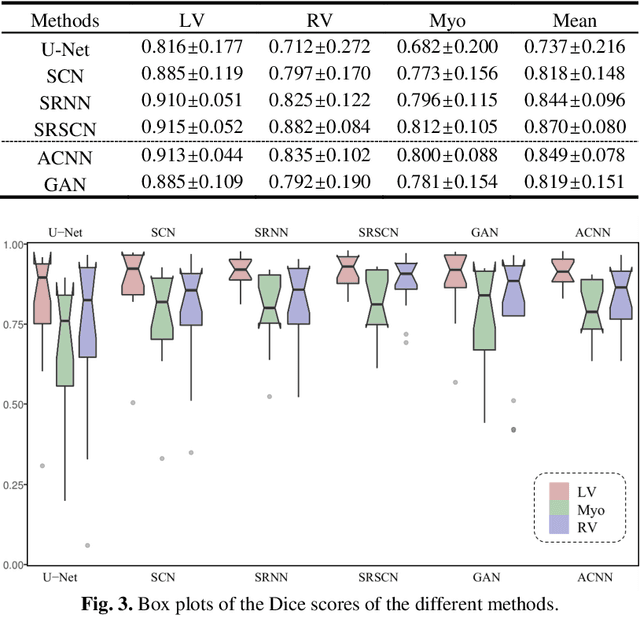
Abstract:Cardiac segmentation from late gadolinium enhancement MRI is an important task in clinics to identify and evaluate the infarction of myocardium. The automatic segmentation is however still challenging, due to the heterogeneous intensity distributions and indistinct boundaries in the images. In this paper, we propose a new method, based on deep neural networks (DNN), for fully automatic segmentation. The proposed network, referred to as SRSCN, comprises a shape reconstruction neural network (SRNN) and a spatial constraint network (SCN). SRNN aims to maintain a realistic shape of the resulting segmentation. It can be pre-trained by a set of label images, and then be embedded into a unified loss function as a regularization term. Hence, no manually designed feature is needed. Furthermore, SCN incorporates the spatial information of the 2D slices. It is formulated and trained with the segmentation network via the multi-task learning strategy. We evaluated the proposed method using 45 patients and compared with two state-of-the-art regularization schemes, i.e., the anatomically constraint neural network and the adversarial neural network. The results show that the proposed SRSCN outperformed the conventional schemes, and obtained a Dice score of 0.758(std=0.227) for myocardial segmentation, which compares with 0.757(std=0.083) from the inter-observer variations.
Atrial Scar Quantification via Multi-scale CNN in the Graph-cuts Framework
Feb 21, 2019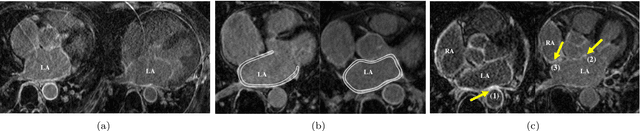
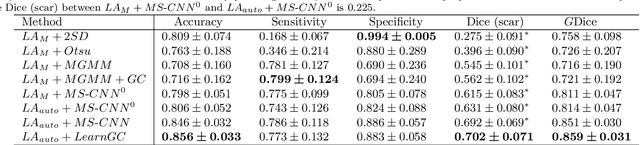
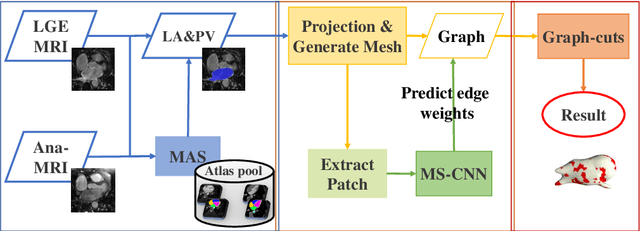

Abstract:Late gadolinium enhancement magnetic resonance imaging (LGE MRI) appears to be a promising alternative for scar assessment in patients with atrial fibrillation (AF). Automating the quantification and analysis of atrial scars can be challenging due to the low image quality. In this work, we propose a fully automated method based on the graph-cuts framework, where the potentials of the graph are learned on a surface mesh of the left atrium (LA) using a multi-scale convolutional neural network (MS-CNN). For validation, we have employed fifty-eight images with manual delineations. MS-CNN, which can efficiently incorporate both the local and global texture information of the images, has been shown to evidently improve the segmentation accuracy of the proposed graph-cuts based method. The segmentation could be further improved when the contribution between the t-link and n-link weights of the graph is balanced. The proposed method achieves a mean accuracy of 0.856 +- 0.033 and mean Dice score of 0.702 +- 0.071 for LA scar quantification. Compared with the conventional methods, which are based on the manual delineation of LA for initialization, our method is fully automatic and has demonstrated significantly better Dice score and accuracy (p < 0.01). The method is promising and can be useful in diagnosis and prognosis of AF.
Atrial scars segmentation via potential learning in the graph-cuts framework
Oct 22, 2018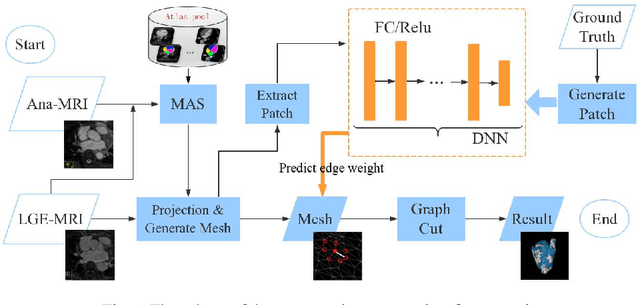
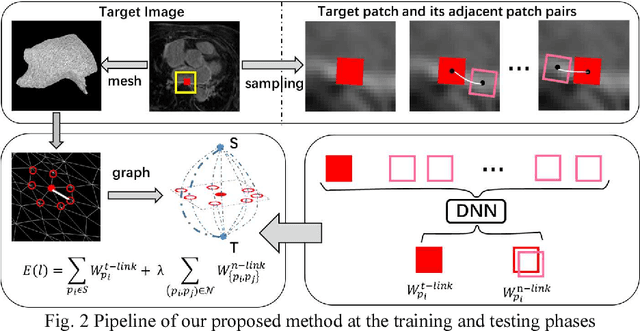
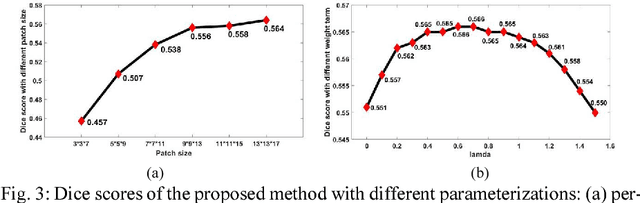
Abstract:Late Gadolinium Enhancement Magnetic Resonance Imaging (LGE MRI) emerged as a routine scan for patients with atrial fibrillation (AF). However, due to the low image quality automating the quantification and analysis of the atrial scars is challenging. In this study, we pro-posed a fully automated method based on the graph-cuts framework, where the potential of the graph is learned on a surface mesh of the left atrium (LA) using an equidistant projection and a Deep Neural Network (DNN). For validation, we employed 100 datasets with manual delineation. The results showed that the performance of the proposed method improved and converged with respect to the increased size of training patches, which provide important features of the structural and texture information learned by the DNN. The segmentation could be further improved when the contribution from the t-link and n-link is balanced, thanks to inter-relationship learned by the DNN for the graph-cuts algorithm. Compared with the published methods which mostly acquired manual delineation of the LA or LA wall, our method is fully automatic and demonstrated evidently better results with statistical significance. Finally, the accuracy of quantifying the scars assessed by the Dice score was 0.570. The results are promising and the method can be useful in diagnosis and prognosis of AF.
Atrial fibrosis quantification based on maximum likelihood estimator of multivariate images
Oct 22, 2018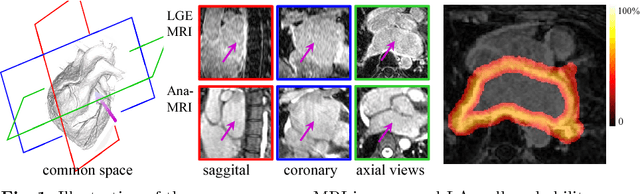
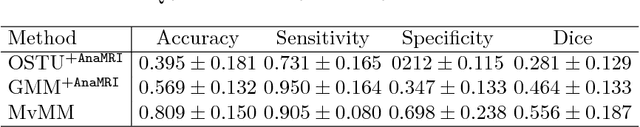

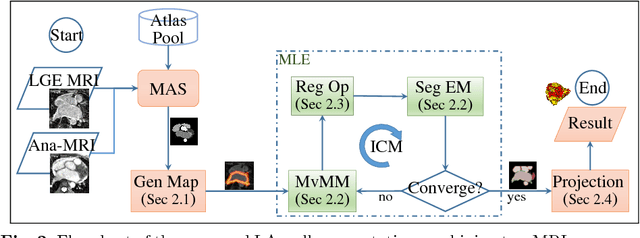
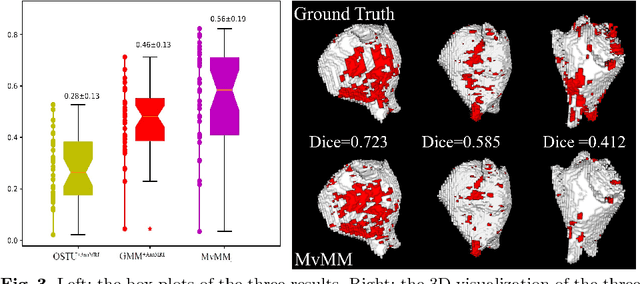
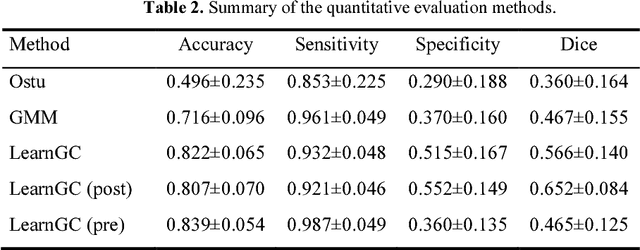
Abstract:We present a fully-automated segmentation and quantification of the left atrial (LA) fibrosis and scars combining two cardiac MRIs, one is the target late gadolinium-enhanced (LGE) image, and the other is an anatomical MRI from the same acquisition session. We formulate the joint distribution of images using a multivariate mixture model (MvMM), and employ the maximum likelihood estimator (MLE) for texture classification of the images simultaneously. The MvMM can also embed transformations assigned to the images to correct the misregistration. The iterated conditional mode algorithm is adopted for optimization. This method first extracts the anatomical shape of the LA, and then estimates a prior probability map. It projects the resulting segmentation onto the LA surface, for quantification and analysis of scarring. We applied the proposed method to 36 clinical data sets and obtained promising results (Accuracy: $0.809\pm .150$, Dice: $0.556\pm.187$). We compared the method with the conventional algorithms and showed an evidently and statistically better performance ($p<0.03$).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge