Kunal Nagpal
Cameron
Efficient and generalizable prediction of molecular alterations in multiple cancer cohorts using H&E whole slide images
Jul 22, 2024Abstract:Molecular testing of tumor samples for targetable biomarkers is restricted by a lack of standardization, turnaround-time, cost, and tissue availability across cancer types. Additionally, targetable alterations of low prevalence may not be tested in routine workflows. Algorithms that predict DNA alterations from routinely generated hematoxylin and eosin (H&E)-stained images could prioritize samples for confirmatory molecular testing. Costs and the necessity of a large number of samples containing mutations limit approaches that train individual algorithms for each alteration. In this work, models were trained for simultaneous prediction of multiple DNA alterations from H&E images using a multi-task approach. Compared to biomarker-specific models, this approach performed better on average, with pronounced gains for rare mutations. The models reasonably generalized to independent temporal-holdout, externally-stained, and multi-site TCGA test sets. Additionally, whole slide image embeddings derived using multi-task models demonstrated strong performance in downstream tasks that were not a part of training. Overall, this is a promising approach to develop clinically useful algorithms that provide multiple actionable predictions from a single slide.
Prediction of MET Overexpression in Non-Small Cell Lung Adenocarcinomas from Hematoxylin and Eosin Images
Oct 12, 2023Abstract:MET protein overexpression is a targetable event in non-small cell lung cancer (NSCLC) and is the subject of active drug development. Challenges in identifying patients for these therapies include lack of access to validated testing, such as standardized immunohistochemistry (IHC) assessment, and consumption of valuable tissue for a single gene/protein assay. Development of pre-screening algorithms using routinely available digitized hematoxylin and eosin (H&E)-stained slides to predict MET overexpression could promote testing for those who will benefit most. While assessment of MET expression using IHC is currently not routinely performed in NSCLC, next-generation sequencing is common and in some cases includes RNA expression panel testing. In this work, we leveraged a large database of matched H&E slides and RNA expression data to train a weakly supervised model to predict MET RNA overexpression directly from H&E images. This model was evaluated on an independent holdout test set of 300 over-expressed and 289 normal patients, demonstrating an ROC-AUC of 0.70 (95th percentile interval: 0.66 - 0.74) with stable performance characteristics across different patient clinical variables and robust to synthetic noise on the test set. These results suggest that H&E-based predictive models could be useful to prioritize patients for confirmatory testing of MET protein or MET gene expression status.
Development and Validation of a Deep Learning-Based Microsatellite Instability Predictor from Prostate Cancer Whole-Slide Images
Oct 12, 2023Abstract:Microsatellite instability-high (MSI-H) is a tumor agnostic biomarker for immune checkpoint inhibitor therapy. However, MSI status is not routinely tested in prostate cancer, in part due to low prevalence and assay cost. As such, prediction of MSI status from hematoxylin and eosin (H&E) stained whole-slide images (WSIs) could identify prostate cancer patients most likely to benefit from confirmatory testing and becoming eligible for immunotherapy. Prostate biopsies and surgical resections from de-identified records of consecutive prostate cancer patients referred to our institution were analyzed. Their MSI status was determined by next generation sequencing. Patients before a cutoff date were split into an algorithm development set (n=4015, MSI-H 1.8%) and a paired validation set (n=173, MSI-H 19.7%) that consisted of two serial sections from each sample, one stained and scanned internally and the other at an external site. Patients after the cutoff date formed the temporal validation set (n=1350, MSI-H 2.3%). Attention-based multiple instance learning models were trained to predict MSI-H from H&E WSIs. The MSI-H predictor achieved area under the receiver operating characteristic curve values of 0.78 (95% CI [0.69-0.86]), 0.72 (95% CI [0.63-0.81]), and 0.72 (95% CI [0.62-0.82]) on the internally prepared, externally prepared, and temporal validation sets, respectively. While MSI-H status is significantly correlated with Gleason score, the model remained predictive within each Gleason score subgroup. In summary, we developed and validated an AI-based MSI-H diagnostic model on a large real-world cohort of routine H&E slides, which effectively generalized to externally stained and scanned samples and a temporally independent validation cohort. This algorithm has the potential to direct prostate cancer patients toward immunotherapy and to identify MSI-H cases secondary to Lynch syndrome.
Predicting Prostate Cancer-Specific Mortality with A.I.-based Gleason Grading
Nov 25, 2020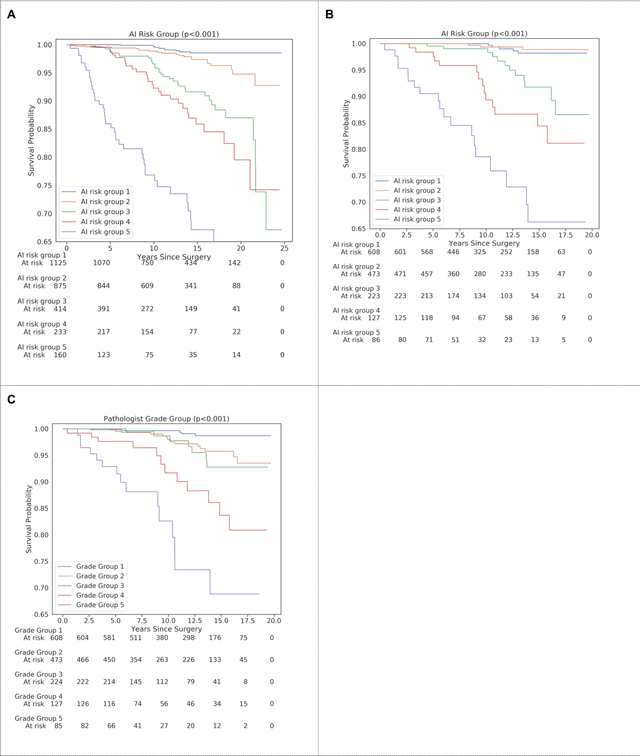
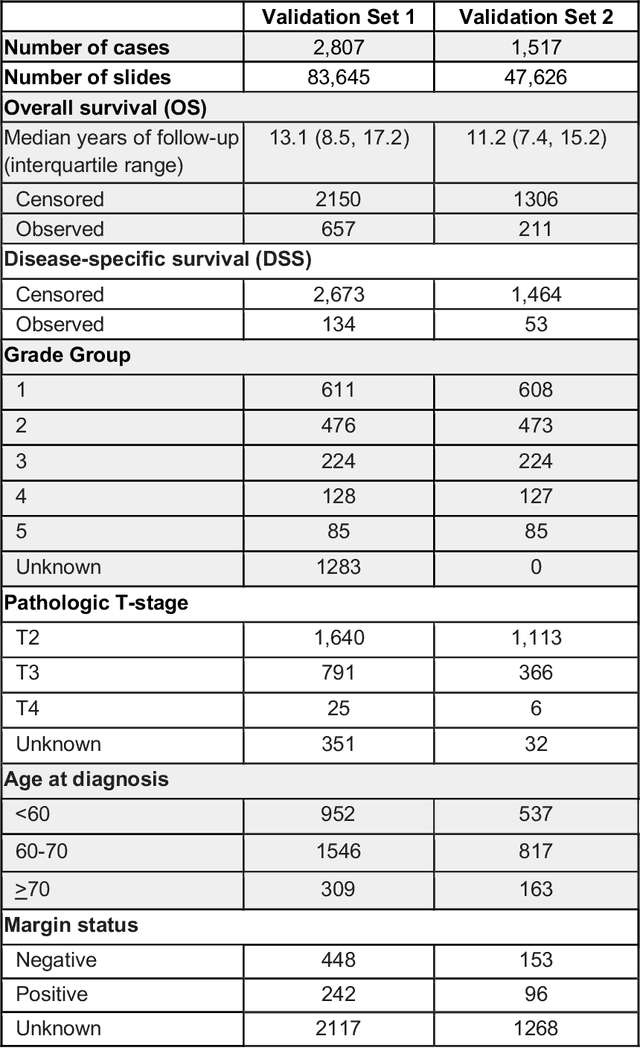
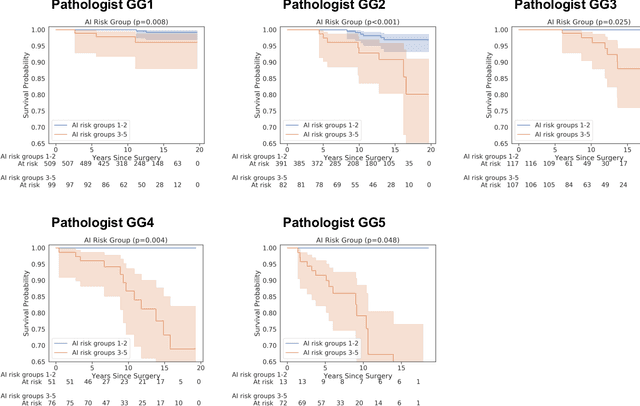
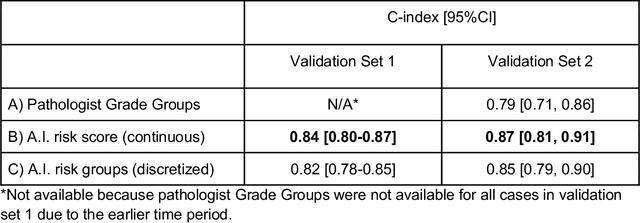
Abstract:Gleason grading of prostate cancer is an important prognostic factor but suffers from poor reproducibility, particularly among non-subspecialist pathologists. Although artificial intelligence (A.I.) tools have demonstrated Gleason grading on-par with expert pathologists, it remains an open question whether A.I. grading translates to better prognostication. In this study, we developed a system to predict prostate-cancer specific mortality via A.I.-based Gleason grading and subsequently evaluated its ability to risk-stratify patients on an independent retrospective cohort of 2,807 prostatectomy cases from a single European center with 5-25 years of follow-up (median: 13, interquartile range 9-17). The A.I.'s risk scores produced a C-index of 0.84 (95%CI 0.80-0.87) for prostate cancer-specific mortality. Upon discretizing these risk scores into risk groups analogous to pathologist Grade Groups (GG), the A.I. had a C-index of 0.82 (95%CI 0.78-0.85). On the subset of cases with a GG in the original pathology report (n=1,517), the A.I.'s C-indices were 0.87 and 0.85 for continuous and discrete grading, respectively, compared to 0.79 (95%CI 0.71-0.86) for GG obtained from the reports. These represent improvements of 0.08 (95%CI 0.01-0.15) and 0.07 (95%CI 0.00-0.14) respectively. Our results suggest that A.I.-based Gleason grading can lead to effective risk-stratification and warrants further evaluation for improving disease management.
Microscope 2.0: An Augmented Reality Microscope with Real-time Artificial Intelligence Integration
Dec 04, 2018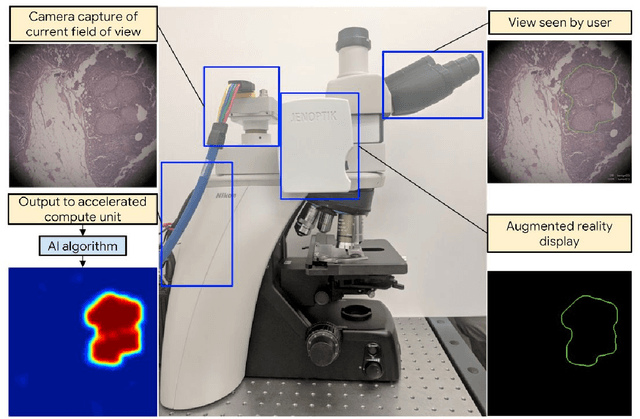
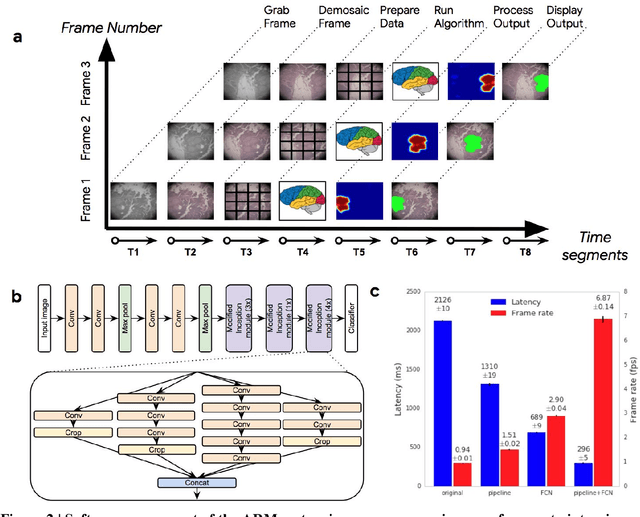
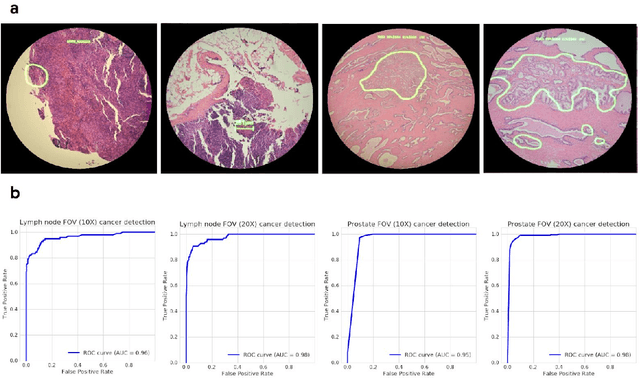
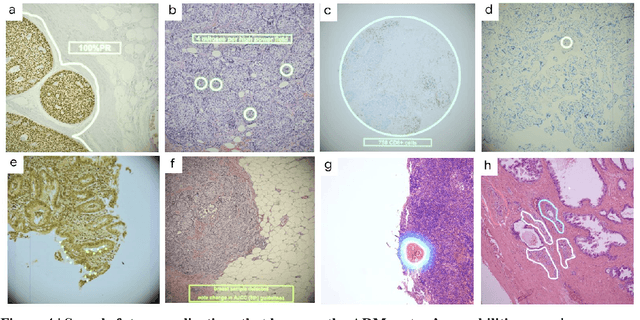
Abstract:The brightfield microscope is instrumental in the visual examination of both biological and physical samples at sub-millimeter scales. One key clinical application has been in cancer histopathology, where the microscopic assessment of the tissue samples is used for the diagnosis and staging of cancer and thus guides clinical therapy. However, the interpretation of these samples is inherently subjective, resulting in significant diagnostic variability. Moreover, in many regions of the world, access to pathologists is severely limited due to lack of trained personnel. In this regard, Artificial Intelligence (AI) based tools promise to improve the access and quality of healthcare. However, despite significant advances in AI research, integration of these tools into real-world cancer diagnosis workflows remains challenging because of the costs of image digitization and difficulties in deploying AI solutions. Here we propose a cost-effective solution to the integration of AI: the Augmented Reality Microscope (ARM). The ARM overlays AI-based information onto the current view of the sample through the optical pathway in real-time, enabling seamless integration of AI into the regular microscopy workflow. We demonstrate the utility of ARM in the detection of lymph node metastases in breast cancer and the identification of prostate cancer with a latency that supports real-time workflows. We anticipate that ARM will remove barriers towards the use of AI in microscopic analysis and thus improve the accuracy and efficiency of cancer diagnosis. This approach is applicable to other microscopy tasks and AI algorithms in the life sciences and beyond.
Development and Validation of a Deep Learning Algorithm for Improving Gleason Scoring of Prostate Cancer
Nov 15, 2018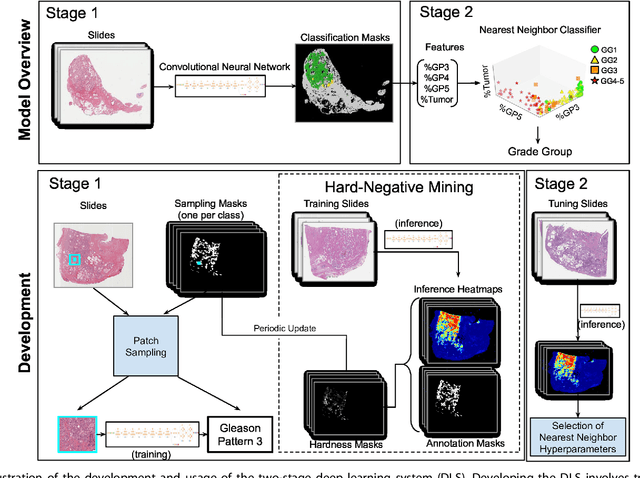
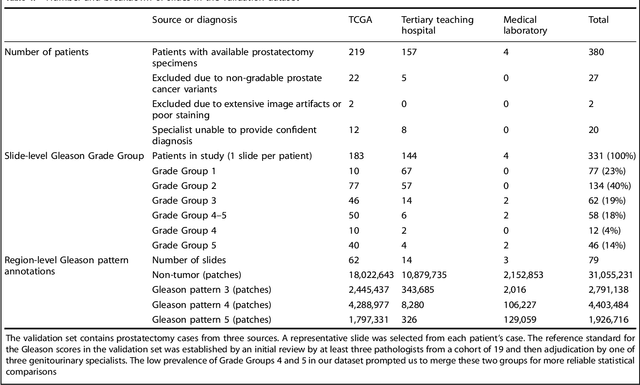
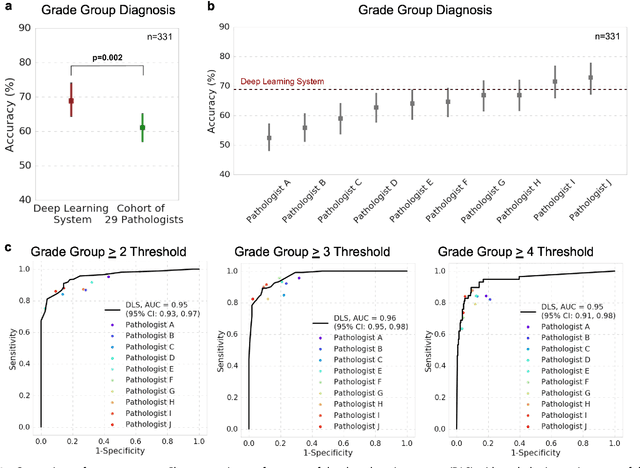
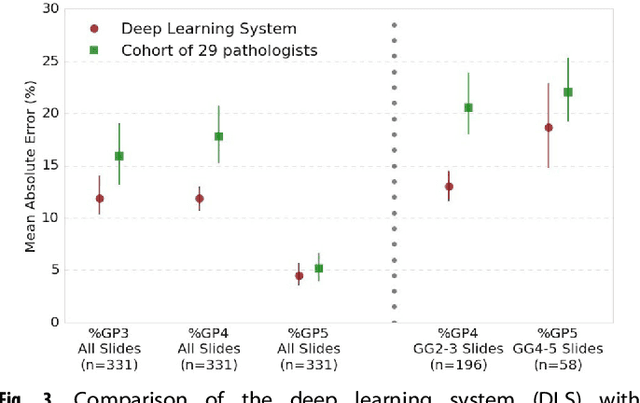
Abstract:For prostate cancer patients, the Gleason score is one of the most important prognostic factors, potentially determining treatment independent of the stage. However, Gleason scoring is based on subjective microscopic examination of tumor morphology and suffers from poor reproducibility. Here we present a deep learning system (DLS) for Gleason scoring whole-slide images of prostatectomies. Our system was developed using 112 million pathologist-annotated image patches from 1,226 slides, and evaluated on an independent validation dataset of 331 slides, where the reference standard was established by genitourinary specialist pathologists. On the validation dataset, the mean accuracy among 29 general pathologists was 0.61. The DLS achieved a significantly higher diagnostic accuracy of 0.70 (p=0.002) and trended towards better patient risk stratification in correlations to clinical follow-up data. Our approach could improve the accuracy of Gleason scoring and subsequent therapy decisions, particularly where specialist expertise is unavailable. The DLS also goes beyond the current Gleason system to more finely characterize and quantitate tumor morphology, providing opportunities for refinement of the Gleason system itself.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge