Yuannan Cai
Predicting Prostate Cancer-Specific Mortality with A.I.-based Gleason Grading
Nov 25, 2020
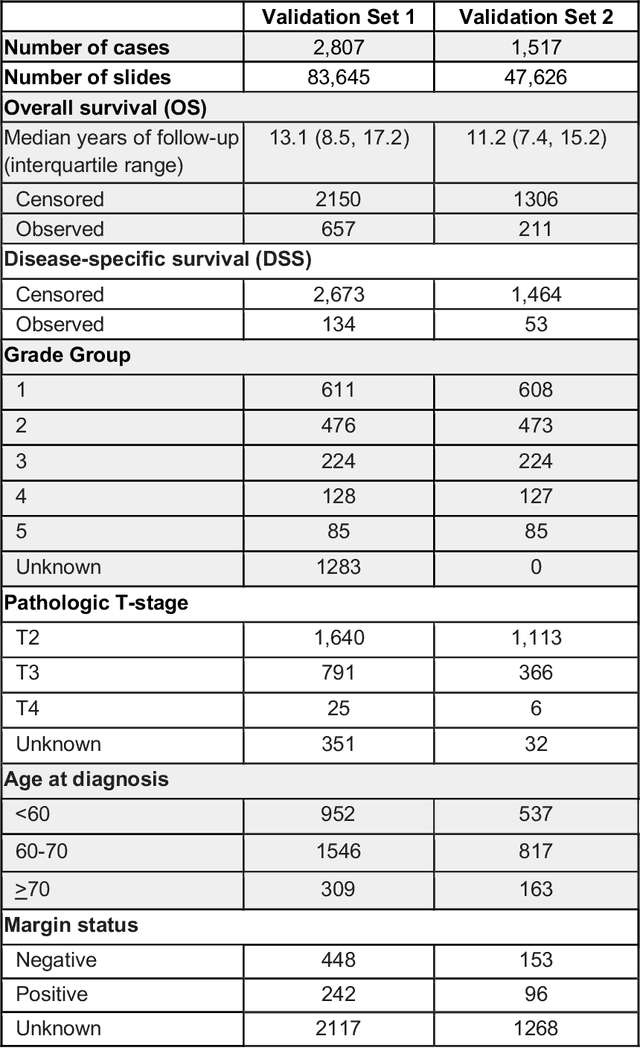
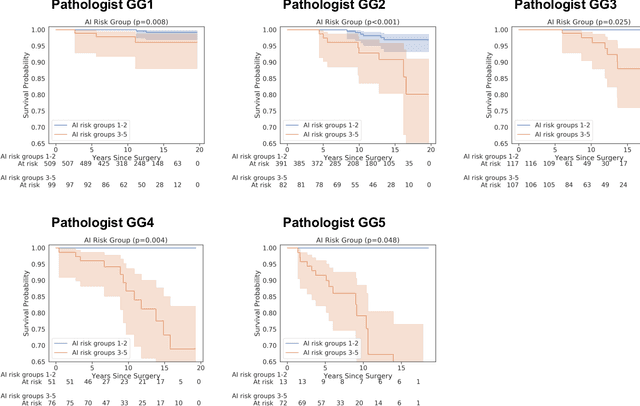
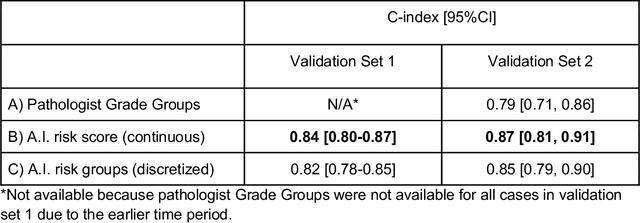
Abstract:Gleason grading of prostate cancer is an important prognostic factor but suffers from poor reproducibility, particularly among non-subspecialist pathologists. Although artificial intelligence (A.I.) tools have demonstrated Gleason grading on-par with expert pathologists, it remains an open question whether A.I. grading translates to better prognostication. In this study, we developed a system to predict prostate-cancer specific mortality via A.I.-based Gleason grading and subsequently evaluated its ability to risk-stratify patients on an independent retrospective cohort of 2,807 prostatectomy cases from a single European center with 5-25 years of follow-up (median: 13, interquartile range 9-17). The A.I.'s risk scores produced a C-index of 0.84 (95%CI 0.80-0.87) for prostate cancer-specific mortality. Upon discretizing these risk scores into risk groups analogous to pathologist Grade Groups (GG), the A.I. had a C-index of 0.82 (95%CI 0.78-0.85). On the subset of cases with a GG in the original pathology report (n=1,517), the A.I.'s C-indices were 0.87 and 0.85 for continuous and discrete grading, respectively, compared to 0.79 (95%CI 0.71-0.86) for GG obtained from the reports. These represent improvements of 0.08 (95%CI 0.01-0.15) and 0.07 (95%CI 0.00-0.14) respectively. Our results suggest that A.I.-based Gleason grading can lead to effective risk-stratification and warrants further evaluation for improving disease management.
Human-centric Metric for Accelerating Pathology Reports Annotation
Nov 12, 2019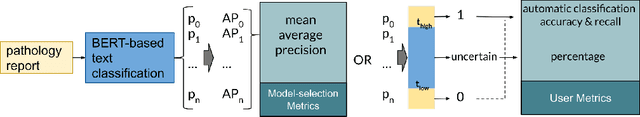
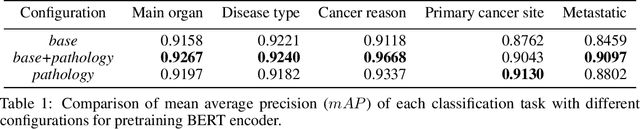
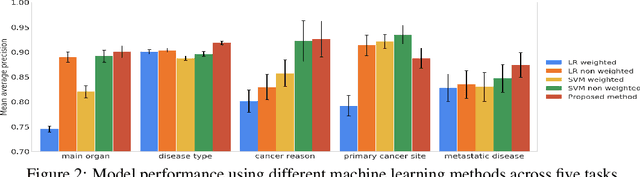
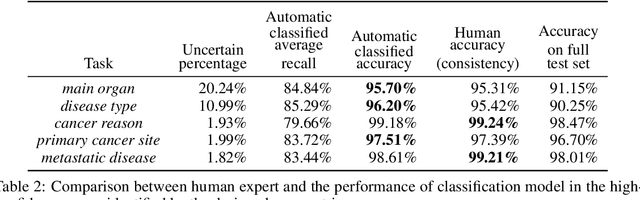
Abstract:Pathology reports contain useful information such as the main involved organ, diagnosis, etc. These information can be identified from the free text reports and used for large-scale statistical analysis or serve as annotation for other modalities such as pathology slides images. However, manual classification for a huge number of reports on multiple tasks is labor-intensive. In this paper, we have developed an automatic text classifier based on BERT and we propose a human-centric metric to evaluate the model. According to the model confidence, we identify low-confidence cases that require further expert annotation and high-confidence cases that are automatically classified. We report the percentage of low-confidence cases and the performance of automatically classified cases. On the high-confidence cases, the model achieves classification accuracy comparable to pathologists. This leads a potential of reducing 80% to 98% of the manual annotation workload.
Multimodal Multitask Representation Learning for Pathology Biobank Metadata Prediction
Sep 17, 2019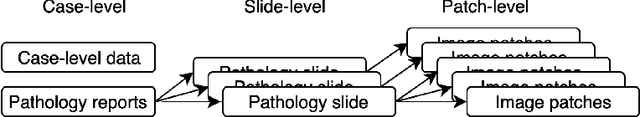
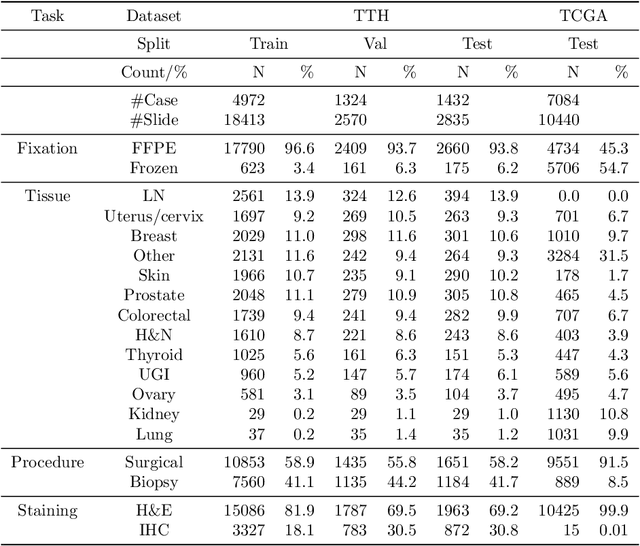


Abstract:Metadata are general characteristics of the data in a well-curated and condensed format, and have been proven to be useful for decision making, knowledge discovery, and also heterogeneous data organization of biobank. Among all data types in the biobank, pathology is the key component of the biobank and also serves as the gold standard of diagnosis. To maximize the utility of biobank and allow the rapid progress of biomedical science, it is essential to organize the data with well-populated pathology metadata. However, manual annotation of such information is tedious and time-consuming. In the study, we develop a multimodal multitask learning framework to predict four major slide-level metadata of pathology images. The framework learns generalizable representations across tissue slides, pathology reports, and case-level structured data. We demonstrate improved performance across all four tasks with the proposed method compared to a single modal single task baseline on two test sets, one external test set from a distinct data source (TCGA) and one internal held-out test set (TTH). In the test sets, the performance improvements on the averaged area under receiver operating characteristic curve across the four tasks are 16.48% and 9.05% on TCGA and TTH, respectively. Such pathology metadata prediction system may be adopted to mitigate the effort of expert annotation and ultimately accelerate the data-driven research by better utilization of the pathology biobank.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge