Julio Silva-Rodríguez
Full Conformal Adaptation of Medical Vision-Language Models
Jun 06, 2025



Abstract:Vision-language models (VLMs) pre-trained at large scale have shown unprecedented transferability capabilities and are being progressively integrated into medical image analysis. Although its discriminative potential has been widely explored, its reliability aspect remains overlooked. This work investigates their behavior under the increasingly popular split conformal prediction (SCP) framework, which theoretically guarantees a given error level on output sets by leveraging a labeled calibration set. However, the zero-shot performance of VLMs is inherently limited, and common practice involves few-shot transfer learning pipelines, which cannot absorb the rigid exchangeability assumptions of SCP. To alleviate this issue, we propose full conformal adaptation, a novel setting for jointly adapting and conformalizing pre-trained foundation models, which operates transductively over each test data point using a few-shot adaptation set. Moreover, we complement this framework with SS-Text, a novel training-free linear probe solver for VLMs that alleviates the computational cost of such a transductive approach. We provide comprehensive experiments using 3 different modality-specialized medical VLMs and 9 adaptation tasks. Our framework requires exactly the same data as SCP, and provides consistent relative improvements of up to 27% on set efficiency while maintaining the same coverage guarantees.
Conformal Prediction for Zero-Shot Models
May 30, 2025Abstract:Vision-language models pre-trained at large scale have shown unprecedented adaptability and generalization to downstream tasks. Although its discriminative potential has been widely explored, its reliability and uncertainty are still overlooked. In this work, we investigate the capabilities of CLIP models under the split conformal prediction paradigm, which provides theoretical guarantees to black-box models based on a small, labeled calibration set. In contrast to the main body of literature on conformal predictors in vision classifiers, foundation models exhibit a particular characteristic: they are pre-trained on a one-time basis on an inaccessible source domain, different from the transferred task. This domain drift negatively affects the efficiency of the conformal sets and poses additional challenges. To alleviate this issue, we propose Conf-OT, a transfer learning setting that operates transductive over the combined calibration and query sets. Solving an optimal transport problem, the proposed method bridges the domain gap between pre-training and adaptation without requiring additional data splits but still maintaining coverage guarantees. We comprehensively explore this conformal prediction strategy on a broad span of 15 datasets and three non-conformity scores. Conf-OT provides consistent relative improvements of up to 20% on set efficiency while being 15 times faster than popular transductive approaches.
A Reality Check of Vision-Language Pre-training in Radiology: Have We Progressed Using Text?
Apr 07, 2025Abstract:Vision-language pre-training has recently gained popularity as it allows learning rich feature representations using large-scale data sources. This paradigm has quickly made its way into the medical image analysis community. In particular, there is an impressive amount of recent literature developing vision-language models for radiology. However, the available medical datasets with image-text supervision are scarce, and medical concepts are fine-grained, involving expert knowledge that existing vision-language models struggle to encode. In this paper, we propose to take a prudent step back from the literature and revisit supervised, unimodal pre-training, using fine-grained labels instead. We conduct an extensive comparison demonstrating that unimodal pre-training is highly competitive and better suited to integrating heterogeneous data sources. Our results also question the potential of recent vision-language models for open-vocabulary generalization, which have been evaluated using optimistic experimental settings. Finally, we study novel alternatives to better integrate fine-grained labels and noisy text supervision.
Are foundation models for computer vision good conformal predictors?
Dec 08, 2024



Abstract:Recent advances in self-supervision and constrastive learning have brought the performance of foundation models to unprecedented levels in a variety of tasks. Fueled by this progress, these models are becoming the prevailing approach for a wide array of real-world vision problems, including risk-sensitive and high-stakes applications. However, ensuring safe deployment in these scenarios requires a more comprehensive understanding of their uncertainty modeling capabilities, which has been barely explored. In this work, we delve into the behavior of vision and vision-language foundation models under Conformal Prediction (CP), a statistical framework that provides theoretical guarantees of marginal coverage of the true class. Across extensive experiments including popular vision classification benchmarks, well-known foundation vision models, and three CP methods, our findings reveal that foundation models are well-suited for conformalization procedures, particularly those integrating Vision Transformers. Furthermore, we show that calibrating the confidence predictions of these models leads to efficiency degradation of the conformal set on adaptive CP methods. In contrast, few-shot adaptation to downstream tasks generally enhances conformal scores, where we identify Adapters as a better conformable alternative compared to Prompt Learning strategies. Our empirical study identifies APS as particularly promising in the context of vision foundation models, as it does not violate the marginal coverage property across multiple challenging, yet realistic scenarios.
Few-shot Adaptation of Medical Vision-Language Models
Sep 05, 2024Abstract:Integrating image and text data through multi-modal learning has emerged as a new approach in medical imaging research, following its successful deployment in computer vision. While considerable efforts have been dedicated to establishing medical foundation models and their zero-shot transfer to downstream tasks, the popular few-shot setting remains relatively unexplored. Following on from the currently strong emergence of this setting in computer vision, we introduce the first structured benchmark for adapting medical vision-language models (VLMs) in a strict few-shot regime and investigate various adaptation strategies commonly used in the context of natural images. Furthermore, we evaluate a simple generalization of the linear-probe adaptation baseline, which seeks an optimal blending of the visual prototypes and text embeddings via learnable class-wise multipliers. Surprisingly, such a text-informed linear probe yields competitive performances in comparison to convoluted prompt-learning and adapter-based strategies, while running considerably faster and accommodating the black-box setting. Our extensive experiments span three different medical modalities and specialized foundation models, nine downstream tasks, and several state-of-the-art few-shot adaptation methods. We made our benchmark and code publicly available to trigger further developments in this emergent subject: \url{https://github.com/FereshteShakeri/few-shot-MedVLMs}.
Self-Contrastive Weakly Supervised Learning Framework for Prognostic Prediction Using Whole Slide Images
May 24, 2024



Abstract:We present a pioneering investigation into the application of deep learning techniques to analyze histopathological images for addressing the substantial challenge of automated prognostic prediction. Prognostic prediction poses a unique challenge as the ground truth labels are inherently weak, and the model must anticipate future events that are not directly observable in the image. To address this challenge, we propose a novel three-part framework comprising of a convolutional network based tissue segmentation algorithm for region of interest delineation, a contrastive learning module for feature extraction, and a nested multiple instance learning classification module. Our study explores the significance of various regions of interest within the histopathological slides and exploits diverse learning scenarios. The pipeline is initially validated on artificially generated data and a simpler diagnostic task. Transitioning to prognostic prediction, tasks become more challenging. Employing bladder cancer as use case, our best models yield an AUC of 0.721 and 0.678 for recurrence and treatment outcome prediction respectively.
Attention to detail: inter-resolution knowledge distillation
Jan 11, 2024Abstract:The development of computer vision solutions for gigapixel images in digital pathology is hampered by significant computational limitations due to the large size of whole slide images. In particular, digitizing biopsies at high resolutions is a time-consuming process, which is necessary due to the worsening results from the decrease in image detail. To alleviate this issue, recent literature has proposed using knowledge distillation to enhance the model performance at reduced image resolutions. In particular, soft labels and features extracted at the highest magnification level are distilled into a model that takes lower-magnification images as input. However, this approach fails to transfer knowledge about the most discriminative image regions in the classification process, which may be lost when the resolution is decreased. In this work, we propose to distill this information by incorporating attention maps during training. In particular, our formulation leverages saliency maps of the target class via grad-CAMs, which guides the lower-resolution Student model to match the Teacher distribution by minimizing the l2 distance between them. Comprehensive experiments on prostate histology image grading demonstrate that the proposed approach substantially improves the model performance across different image resolutions compared to previous literature.
HistoColAi: An Open-Source Web Platform for Collaborative Digital Histology Image Annotation with AI-Driven Predictive Integration
Jul 11, 2023Abstract:Digital pathology has become a standard in the pathology workflow due to its many benefits. These include the level of detail of the whole slide images generated and the potential immediate sharing of cases between hospitals. Recent advances in deep learning-based methods for image analysis make them of potential aid in digital pathology. However, a major limitation in developing computer-aided diagnostic systems for pathology is the lack of an intuitive and open web application for data annotation. This paper proposes a web service that efficiently provides a tool to visualize and annotate digitized histological images. In addition, to show and validate the tool, in this paper we include a use case centered on the diagnosis of spindle cell skin neoplasm for multiple annotators. A usability study of the tool is also presented, showing the feasibility of the developed tool.
Transductive few-shot adapters for medical image segmentation
Mar 29, 2023
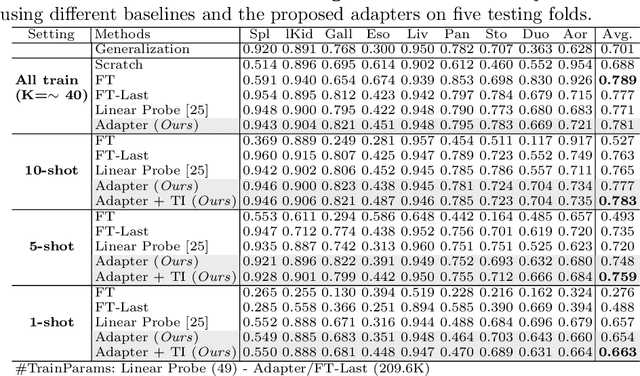


Abstract:With the recent raise of foundation models in computer vision and NLP, the pretrain-and-adapt strategy, where a large-scale model is fine-tuned on downstream tasks, is gaining popularity. However, traditional fine-tuning approaches may still require significant resources and yield sub-optimal results when the labeled data of the target task is scarce. This is especially the case in clinical settings. To address this challenge, we formalize few-shot efficient fine-tuning (FSEFT), a novel and realistic setting for medical image segmentation. Furthermore, we introduce a novel parameter-efficient fine-tuning strategy tailored to medical image segmentation, with (a) spatial adapter modules that are more appropriate for dense prediction tasks; and (b) a constrained transductive inference, which leverages task-specific prior knowledge. Our comprehensive experiments on a collection of public CT datasets for organ segmentation reveal the limitations of standard fine-tuning methods in few-shot scenarios, point to the potential of vision adapters and transductive inference, and confirm the suitability of foundation models.
Challenging mitosis detection algorithms: Global labels allow centroid localization
Nov 30, 2022Abstract:Mitotic activity is a crucial proliferation biomarker for the diagnosis and prognosis of different types of cancers. Nevertheless, mitosis counting is a cumbersome process for pathologists, prone to low reproducibility, due to the large size of augmented biopsy slides, the low density of mitotic cells, and pattern heterogeneity. To improve reproducibility, deep learning methods have been proposed in the last years using convolutional neural networks. However, these methods have been hindered by the process of data labelling, which usually solely consist of the mitosis centroids. Therefore, current literature proposes complex algorithms with multiple stages to refine the labels at pixel level, and to reduce the number of false positives. In this work, we propose to avoid complex scenarios, and we perform the localization task in a weakly supervised manner, using only image-level labels on patches. The results obtained on the publicly available TUPAC16 dataset are competitive with state-of-the-art methods, using only one training phase. Our method achieves an F1-score of 0.729 and challenges the efficiency of previous methods, which required multiple stages and strong mitosis location information.
* Presented at IDEAL 2022
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge