Paul-Henry Cournède
SoC: Semantic Orthogonal Calibration for Test-Time Prompt Tuning
Jan 13, 2026Abstract:With the increasing adoption of vision-language models (VLMs) in critical decision-making systems such as healthcare or autonomous driving, the calibration of their uncertainty estimates becomes paramount. Yet, this dimension has been largely underexplored in the VLM test-time prompt-tuning (TPT) literature, which has predominantly focused on improving their discriminative performance. Recent state-of-the-art advocates for enforcing full orthogonality over pairs of text prompt embeddings to enhance separability, and therefore calibration. Nevertheless, as we theoretically show in this work, the inherent gradients from fully orthogonal constraints will strongly push semantically related classes away, ultimately making the model overconfident. Based on our findings, we propose Semantic Orthogonal Calibration (SoC), a Huber-based regularizer that enforces smooth prototype separation while preserving semantic proximity, thereby improving calibration compared to prior orthogonality-based approaches. Across a comprehensive empirical validation, we demonstrate that SoC consistently improves calibration performance, while also maintaining competitive discriminative capabilities.
PVeRA: Probabilistic Vector-Based Random Matrix Adaptation
Dec 08, 2025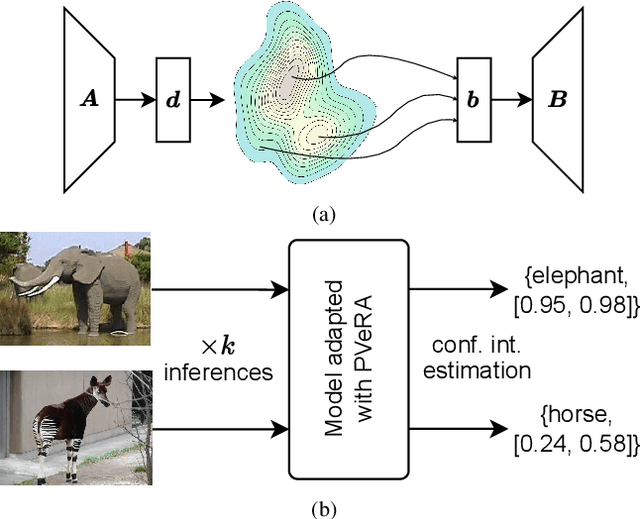
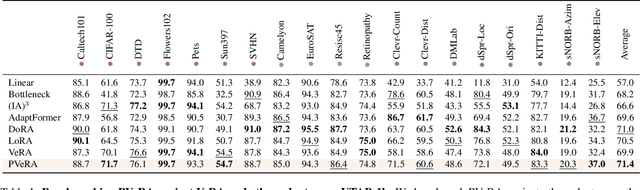
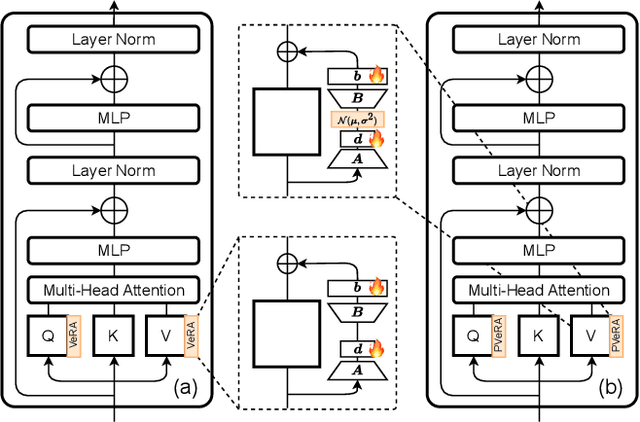

Abstract:Large foundation models have emerged in the last years and are pushing performance boundaries for a variety of tasks. Training or even finetuning such models demands vast datasets and computational resources, which are often scarce and costly. Adaptation methods provide a computationally efficient solution to address these limitations by allowing such models to be finetuned on small amounts of data and computing power. This is achieved by appending new trainable modules to frozen backbones with only a fraction of the trainable parameters and fitting only these modules on novel tasks. Recently, the VeRA adapter was shown to excel in parameter-efficient adaptations by utilizing a pair of frozen random low-rank matrices shared across all layers. In this paper, we propose PVeRA, a probabilistic version of the VeRA adapter, which modifies the low-rank matrices of VeRA in a probabilistic manner. This modification naturally allows handling inherent ambiguities in the input and allows for different sampling configurations during training and testing. A comprehensive evaluation was performed on the VTAB-1k benchmark and seven adapters, with PVeRA outperforming VeRA and other adapters. Our code for training models with PVeRA and benchmarking all adapters is available https://github.com/leofillioux/pvera.
Full Conformal Adaptation of Medical Vision-Language Models
Jun 06, 2025



Abstract:Vision-language models (VLMs) pre-trained at large scale have shown unprecedented transferability capabilities and are being progressively integrated into medical image analysis. Although its discriminative potential has been widely explored, its reliability aspect remains overlooked. This work investigates their behavior under the increasingly popular split conformal prediction (SCP) framework, which theoretically guarantees a given error level on output sets by leveraging a labeled calibration set. However, the zero-shot performance of VLMs is inherently limited, and common practice involves few-shot transfer learning pipelines, which cannot absorb the rigid exchangeability assumptions of SCP. To alleviate this issue, we propose full conformal adaptation, a novel setting for jointly adapting and conformalizing pre-trained foundation models, which operates transductively over each test data point using a few-shot adaptation set. Moreover, we complement this framework with SS-Text, a novel training-free linear probe solver for VLMs that alleviates the computational cost of such a transductive approach. We provide comprehensive experiments using 3 different modality-specialized medical VLMs and 9 adaptation tasks. Our framework requires exactly the same data as SCP, and provides consistent relative improvements of up to 27% on set efficiency while maintaining the same coverage guarantees.
Are foundation models for computer vision good conformal predictors?
Dec 08, 2024



Abstract:Recent advances in self-supervision and constrastive learning have brought the performance of foundation models to unprecedented levels in a variety of tasks. Fueled by this progress, these models are becoming the prevailing approach for a wide array of real-world vision problems, including risk-sensitive and high-stakes applications. However, ensuring safe deployment in these scenarios requires a more comprehensive understanding of their uncertainty modeling capabilities, which has been barely explored. In this work, we delve into the behavior of vision and vision-language foundation models under Conformal Prediction (CP), a statistical framework that provides theoretical guarantees of marginal coverage of the true class. Across extensive experiments including popular vision classification benchmarks, well-known foundation vision models, and three CP methods, our findings reveal that foundation models are well-suited for conformalization procedures, particularly those integrating Vision Transformers. Furthermore, we show that calibrating the confidence predictions of these models leads to efficiency degradation of the conformal set on adaptive CP methods. In contrast, few-shot adaptation to downstream tasks generally enhances conformal scores, where we identify Adapters as a better conformable alternative compared to Prompt Learning strategies. Our empirical study identifies APS as particularly promising in the context of vision foundation models, as it does not violate the marginal coverage property across multiple challenging, yet realistic scenarios.
Causal Contrastive Learning for Counterfactual Regression Over Time
Jun 01, 2024Abstract:Estimating treatment effects over time holds significance in various domains, including precision medicine, epidemiology, economy, and marketing. This paper introduces a unique approach to counterfactual regression over time, emphasizing long-term predictions. Distinguishing itself from existing models like Causal Transformer, our approach highlights the efficacy of employing RNNs for long-term forecasting, complemented by Contrastive Predictive Coding (CPC) and Information Maximization (InfoMax). Emphasizing efficiency, we avoid the need for computationally expensive transformers. Leveraging CPC, our method captures long-term dependencies in the presence of time-varying confounders. Notably, recent models have disregarded the importance of invertible representation, compromising identification assumptions. To remedy this, we employ the InfoMax principle, maximizing a lower bound of mutual information between sequence data and its representation. Our method achieves state-of-the-art counterfactual estimation results using both synthetic and real-world data, marking the pioneering incorporation of Contrastive Predictive Encoding in causal inference.
Causal Dynamic Variational Autoencoder for Counterfactual Regression in Longitudinal Data
Oct 16, 2023Abstract:Estimating treatment effects over time is relevant in many real-world applications, such as precision medicine, epidemiology, economy, and marketing. Many state-of-the-art methods either assume the observations of all confounders or seek to infer the unobserved ones. We take a different perspective by assuming unobserved risk factors, i.e., adjustment variables that affect only the sequence of outcomes. Under unconfoundedness, we target the Individual Treatment Effect (ITE) estimation with unobserved heterogeneity in the treatment response due to missing risk factors. We address the challenges posed by time-varying effects and unobserved adjustment variables. Led by theoretical results over the validity of the learned adjustment variables and generalization bounds over the treatment effect, we devise Causal DVAE (CDVAE). This model combines a Dynamic Variational Autoencoder (DVAE) framework with a weighting strategy using propensity scores to estimate counterfactual responses. The CDVAE model allows for accurate estimation of ITE and captures the underlying heterogeneity in longitudinal data. Evaluations of our model show superior performance over state-of-the-art models.
Spatio-Temporal Analysis of Patient-Derived Organoid Videos Using Deep Learning for the Prediction of Drug Efficacy
Aug 28, 2023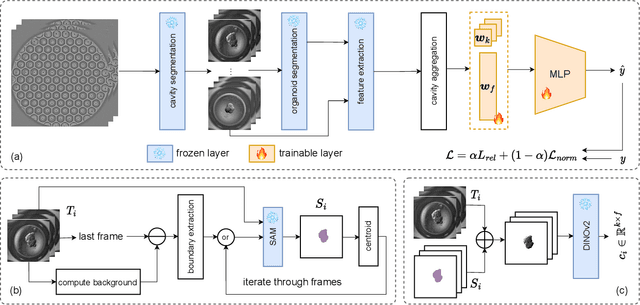

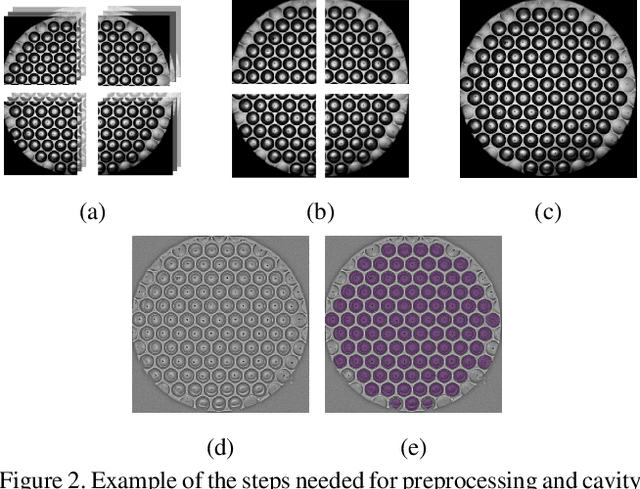

Abstract:Over the last ten years, Patient-Derived Organoids (PDOs) emerged as the most reliable technology to generate ex-vivo tumor avatars. PDOs retain the main characteristics of their original tumor, making them a system of choice for pre-clinical and clinical studies. In particular, PDOs are attracting interest in the field of Functional Precision Medicine (FPM), which is based upon an ex-vivo drug test in which living tumor cells (such as PDOs) from a specific patient are exposed to a panel of anti-cancer drugs. Currently, the Adenosine Triphosphate (ATP) based cell viability assay is the gold standard test to assess the sensitivity of PDOs to drugs. The readout is measured at the end of the assay from a global PDO population and therefore does not capture single PDO responses and does not provide time resolution of drug effect. To this end, in this study, we explore for the first time the use of powerful large foundation models for the automatic processing of PDO data. In particular, we propose a novel imaging-based high-throughput screening method to assess real-time drug efficacy from a time-lapse microscopy video of PDOs. The recently proposed SAM algorithm for segmentation and DINOv2 model are adapted in a comprehensive pipeline for processing PDO microscopy frames. Moreover, an attention mechanism is proposed for fusing temporal and spatial features in a multiple instance learning setting to predict ATP. We report better results than other non-time-resolved methods, indicating that the temporality of data is an important factor for the prediction of ATP. Extensive ablations shed light on optimizing the experimental setting and automating the prediction both in real-time and for forecasting.
Structured State Space Models for Multiple Instance Learning in Digital Pathology
Jun 27, 2023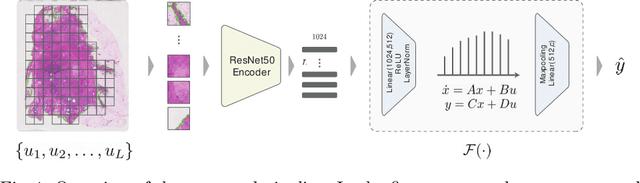
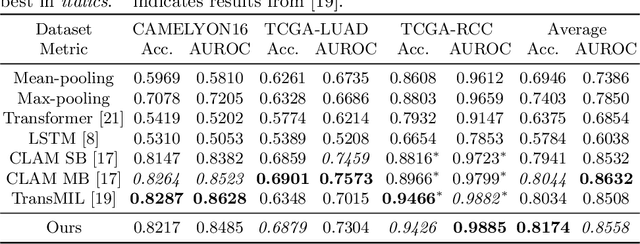

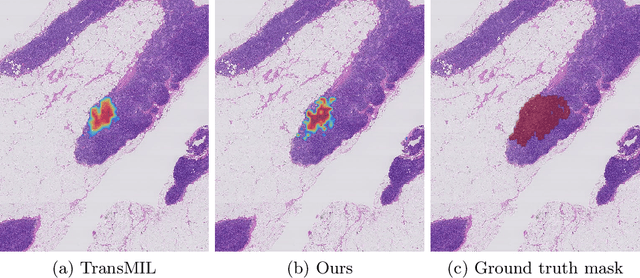
Abstract:Multiple instance learning is an ideal mode of analysis for histopathology data, where vast whole slide images are typically annotated with a single global label. In such cases, a whole slide image is modelled as a collection of tissue patches to be aggregated and classified. Common models for performing this classification include recurrent neural networks and transformers. Although powerful compression algorithms, such as deep pre-trained neural networks, are used to reduce the dimensionality of each patch, the sequences arising from whole slide images remain excessively long, routinely containing tens of thousands of patches. Structured state space models are an emerging alternative for sequence modelling, specifically designed for the efficient modelling of long sequences. These models invoke an optimal projection of an input sequence into memory units that compress the entire sequence. In this paper, we propose the use of state space models as a multiple instance learner to a variety of problems in digital pathology. Across experiments in metastasis detection, cancer subtyping, mutation classification, and multitask learning, we demonstrate the competitiveness of this new class of models with existing state of the art approaches. Our code is available at https://github.com/MICS-Lab/s4_digital_pathology.
CustOmics: A versatile deep-learning based strategy for multi-omics integration
Sep 12, 2022
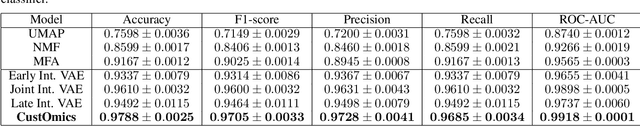
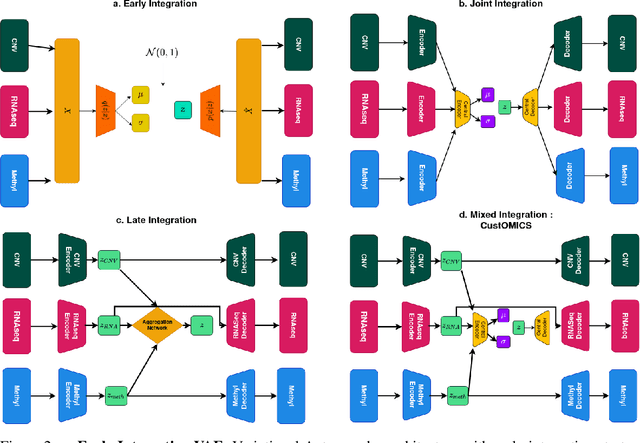

Abstract:Recent advances in high-throughput sequencing technologies have enabled the extraction of multiple features that depict patient samples at diverse and complementary molecular levels. The generation of such data has led to new challenges in computational biology regarding the integration of high-dimensional and heterogeneous datasets that capture the interrelationships between multiple genes and their functions. Thanks to their versatility and ability to learn synthetic latent representations of complex data, deep learning methods offer promising perspectives for integrating multi-omics data. These methods have led to the conception of many original architectures that are primarily based on autoencoder models. However, due to the difficulty of the task, the integration strategy is fundamental to take full advantage of the sources' particularities without losing the global trends. This paper presents a novel strategy to build a customizable autoencoder model that adapts to the dataset used in the case of high-dimensional multi-source integration. We will assess the impact of integration strategies on the latent representation and combine the best strategies to propose a new method, CustOmics (https://github.com/HakimBenkirane/CustOmics). We focus here on the integration of data from multiple omics sources and demonstrate the performance of the proposed method on test cases for several tasks such as classification and survival analysis.
Interpretable cytometry cell-type annotation with flow-based deep generative models
Aug 11, 2022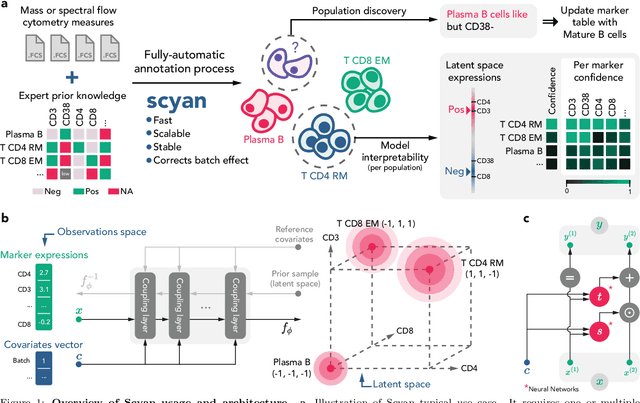
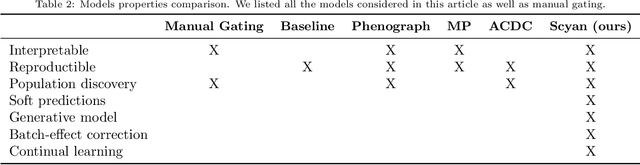
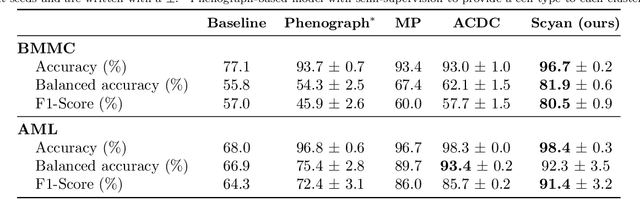
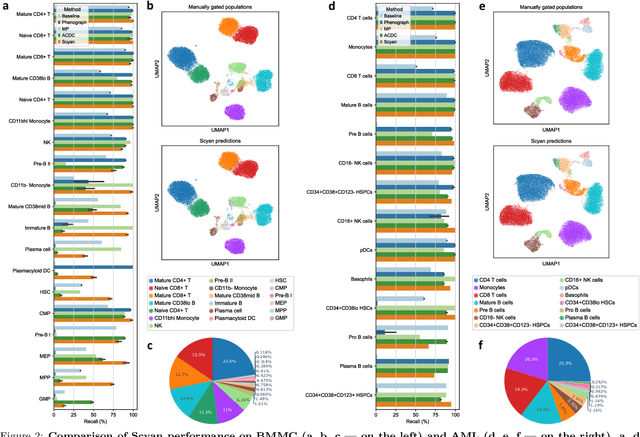

Abstract:Cytometry enables precise single-cell phenotyping within heterogeneous populations. These cell types are traditionally annotated via manual gating, but this method suffers from a lack of reproducibility and sensitivity to batch-effect. Also, the most recent cytometers - spectral flow or mass cytometers - create rich and high-dimensional data whose analysis via manual gating becomes challenging and time-consuming. To tackle these limitations, we introduce Scyan (https://github.com/MICS-Lab/scyan), a Single-cell Cytometry Annotation Network that automatically annotates cell types using only prior expert knowledge about the cytometry panel. We demonstrate that Scyan significantly outperforms the related state-of-the-art models on multiple public datasets while being faster and interpretable. In addition, Scyan overcomes several complementary tasks such as batch-effect removal, debarcoding, and population discovery. Overall, this model accelerates and eases cell population characterisation, quantification, and discovery in cytometry.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge