Hongshuai Wang
PolyConf: Unlocking Polymer Conformation Generation through Hierarchical Generative Models
Apr 11, 2025Abstract:Polymer conformation generation is a critical task that enables atomic-level studies of diverse polymer materials. While significant advances have been made in designing various conformation generation methods for small molecules and proteins, these methods struggle to generate polymer conformations due to polymers' unique structural characteristics. The scarcity of polymer conformation datasets further limits progress, making this promising area largely unexplored. In this work, we propose PolyConf, a pioneering tailored polymer conformation generation method that leverages hierarchical generative models to unlock new possibilities for this task. Specifically, we decompose the polymer conformation into a series of local conformations (i.e., the conformations of its repeating units), generating these local conformations through an autoregressive model. We then generate corresponding orientation transformations via a diffusion model to assemble these local conformations into the complete polymer conformation. Moreover, we develop the first benchmark with a high-quality polymer conformation dataset derived from molecular dynamics simulations to boost related research in this area. The comprehensive evaluation demonstrates that PolyConf consistently generates high-quality polymer conformations, facilitating advancements in polymer modeling and simulation.
SciAssess: Benchmarking LLM Proficiency in Scientific Literature Analysis
Mar 15, 2024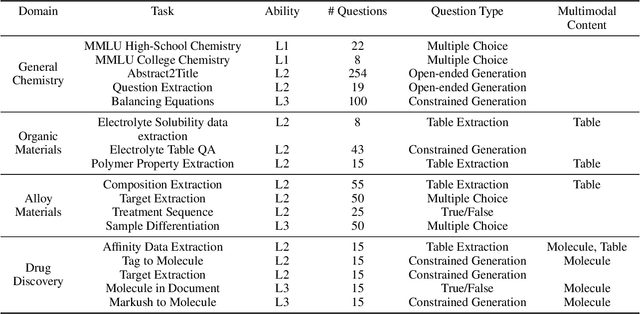
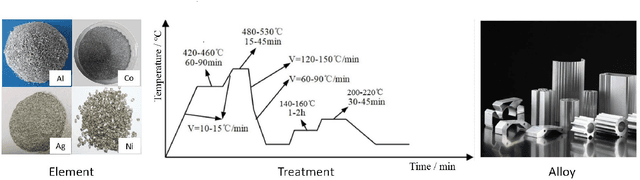
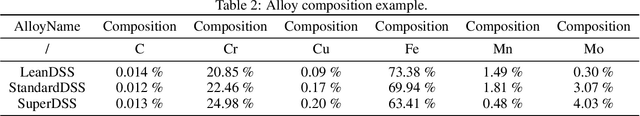
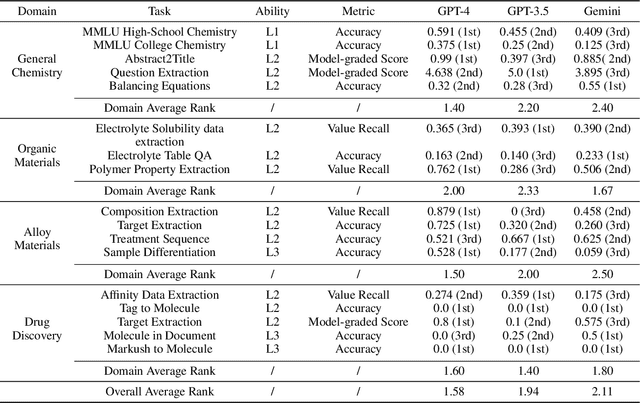
Abstract:Recent breakthroughs in Large Language Models (LLMs) have revolutionized natural language understanding and generation, igniting a surge of interest in leveraging these technologies in the field of scientific literature analysis. Existing benchmarks, however, inadequately evaluate the proficiency of LLMs in scientific literature analysis, especially in scenarios involving complex comprehension and multimodal data. In response, we introduced SciAssess, a benchmark tailored for the in-depth analysis of scientific literature, crafted to provide a thorough assessment of LLMs' efficacy. SciAssess focuses on evaluating LLMs' abilities in memorization, comprehension, and analysis within the context of scientific literature analysis. It includes representative tasks from diverse scientific fields, such as general chemistry, organic materials, and alloy materials. And rigorous quality control measures ensure its reliability in terms of correctness, anonymization, and copyright compliance. SciAssess evaluates leading LLMs, including GPT-4, GPT-3.5, and Gemini, identifying their strengths and aspects for improvement and supporting the ongoing development of LLM applications in scientific literature analysis. SciAssess and its resources are made available at https://sci-assess.github.io, offering a valuable tool for advancing LLM capabilities in scientific literature analysis.
Uni-SMART: Universal Science Multimodal Analysis and Research Transformer
Mar 15, 2024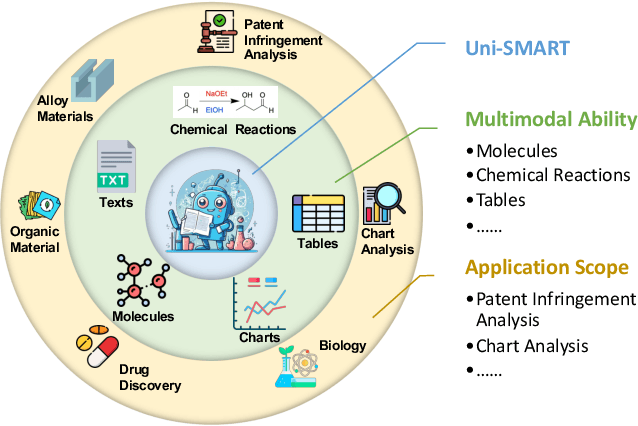
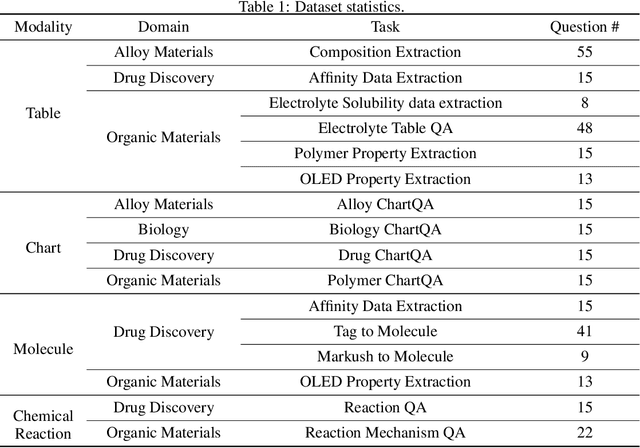
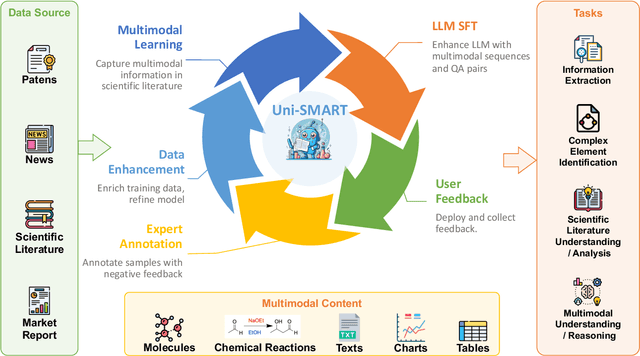
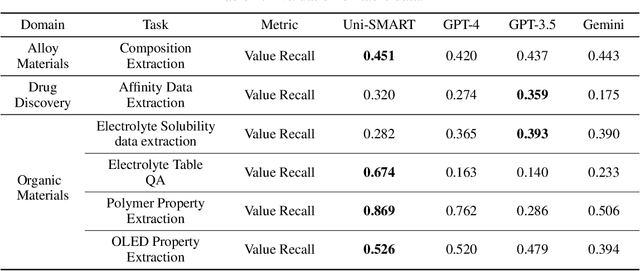
Abstract:In scientific research and its application, scientific literature analysis is crucial as it allows researchers to build on the work of others. However, the fast growth of scientific knowledge has led to a massive increase in scholarly articles, making in-depth literature analysis increasingly challenging and time-consuming. The emergence of Large Language Models (LLMs) has offered a new way to address this challenge. Known for their strong abilities in summarizing texts, LLMs are seen as a potential tool to improve the analysis of scientific literature. However, existing LLMs have their own limits. Scientific literature often includes a wide range of multimodal elements, such as molecular structure, tables, and charts, which are hard for text-focused LLMs to understand and analyze. This issue points to the urgent need for new solutions that can fully understand and analyze multimodal content in scientific literature. To answer this demand, we present Uni-SMART (Universal Science Multimodal Analysis and Research Transformer), an innovative model designed for in-depth understanding of multimodal scientific literature. Through rigorous quantitative evaluation across several domains, Uni-SMART demonstrates superior performance over leading text-focused LLMs. Furthermore, our exploration extends to practical applications, including patent infringement detection and nuanced analysis of charts. These applications not only highlight Uni-SMART's adaptability but also its potential to revolutionize how we interact with scientific literature.
End-to-End Crystal Structure Prediction from Powder X-Ray Diffraction
Jan 08, 2024



Abstract:Powder X-ray diffraction (PXRD) is a crucial means for crystal structure determination. Such determination often involves external database matching to find a structural analogue and Rietveld refinement to obtain finer structure. However, databases may be incomplete and Rietveld refinement often requires intensive trial-and-error efforts from trained experimentalists, which remains ineffective in practice. To settle these issues, we propose XtalNet, the first end-to-end deep learning-based framework capable of ab initio generation of crystal structures that accurately match given PXRD patterns. The model employs contrastive learning and Diffusion-based conditional generation to enable the simultaneous execution of two tasks: crystal structure retrieval based on PXRD patterns and conditional structure generations. To validate the effectiveness of XtalNet, we curate a much more challenging and practical dataset hMOF-100, XtalNet performs well on this dataset, reaching 96.3\% top-10 hit ratio on the database retrieval task and 95.0\% top-10 match rate on the ranked structure generation task.
Uni-QSAR: an Auto-ML Tool for Molecular Property Prediction
Apr 24, 2023



Abstract:Recently deep learning based quantitative structure-activity relationship (QSAR) models has shown surpassing performance than traditional methods for property prediction tasks in drug discovery. However, most DL based QSAR models are restricted to limited labeled data to achieve better performance, and also are sensitive to model scale and hyper-parameters. In this paper, we propose Uni-QSAR, a powerful Auto-ML tool for molecule property prediction tasks. Uni-QSAR combines molecular representation learning (MRL) of 1D sequential tokens, 2D topology graphs, and 3D conformers with pretraining models to leverage rich representation from large-scale unlabeled data. Without any manual fine-tuning or model selection, Uni-QSAR outperforms SOTA in 21/22 tasks of the Therapeutic Data Commons (TDC) benchmark under designed parallel workflow, with an average performance improvement of 6.09\%. Furthermore, we demonstrate the practical usefulness of Uni-QSAR in drug discovery domains.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge