Hehuan Ma
GRAM-TDI: adaptive multimodal representation learning for drug target interaction prediction
Sep 26, 2025Abstract:Drug target interaction (DTI) prediction is a cornerstone of computational drug discovery, enabling rational design, repurposing, and mechanistic insights. While deep learning has advanced DTI modeling, existing approaches primarily rely on SMILES protein pairs and fail to exploit the rich multimodal information available for small molecules and proteins. We introduce GRAMDTI, a pretraining framework that integrates multimodal molecular and protein inputs into unified representations. GRAMDTI extends volume based contrastive learning to four modalities, capturing higher-order semantic alignment beyond conventional pairwise approaches. To handle modality informativeness, we propose adaptive modality dropout, dynamically regulating each modality's contribution during pre-training. Additionally, IC50 activity measurements, when available, are incorporated as weak supervision to ground representations in biologically meaningful interaction strengths. Experiments on four publicly available datasets demonstrate that GRAMDTI consistently outperforms state of the art baselines. Our results highlight the benefits of higher order multimodal alignment, adaptive modality utilization, and auxiliary supervision for robust and generalizable DTI prediction.
TRIDENT: Tri-Modal Molecular Representation Learning with Taxonomic Annotations and Local Correspondence
Jun 26, 2025Abstract:Molecular property prediction aims to learn representations that map chemical structures to functional properties. While multimodal learning has emerged as a powerful paradigm to learn molecular representations, prior works have largely overlooked textual and taxonomic information of molecules for representation learning. We introduce TRIDENT, a novel framework that integrates molecular SMILES, textual descriptions, and taxonomic functional annotations to learn rich molecular representations. To achieve this, we curate a comprehensive dataset of molecule-text pairs with structured, multi-level functional annotations. Instead of relying on conventional contrastive loss, TRIDENT employs a volume-based alignment objective to jointly align tri-modal features at the global level, enabling soft, geometry-aware alignment across modalities. Additionally, TRIDENT introduces a novel local alignment objective that captures detailed relationships between molecular substructures and their corresponding sub-textual descriptions. A momentum-based mechanism dynamically balances global and local alignment, enabling the model to learn both broad functional semantics and fine-grained structure-function mappings. TRIDENT achieves state-of-the-art performance on 11 downstream tasks, demonstrating the value of combining SMILES, textual, and taxonomic functional annotations for molecular property prediction.
Leveraging Gait Patterns as Biomarkers: An attention-guided Deep Multiple Instance Learning Network for Scoliosis Classification
Apr 04, 2025Abstract:Scoliosis is a spinal curvature disorder that is difficult to detect early and can compress the chest cavity, impacting respiratory function and cardiac health. Especially for adolescents, delayed detection and treatment result in worsening compression. Traditional scoliosis detection methods heavily rely on clinical expertise, and X-ray imaging poses radiation risks, limiting large-scale early screening. We propose an Attention-Guided Deep Multi-Instance Learning method (Gait-MIL) to effectively capture discriminative features from gait patterns, which is inspired by ScoNet-MT's pioneering use of gait patterns for scoliosis detection. We evaluate our method on the first large-scale dataset based on gait patterns for scoliosis classification. The results demonstrate that our study improves the performance of using gait as a biomarker for scoliosis detection, significantly enhances detection accuracy for the particularly challenging Neutral cases, where subtle indicators are often overlooked. Our Gait-MIL also performs robustly in imbalanced scenarios, making it a promising tool for large-scale scoliosis screening.
MLLM4PUE: Toward Universal Embeddings in Computational Pathology through Multimodal LLMs
Feb 11, 2025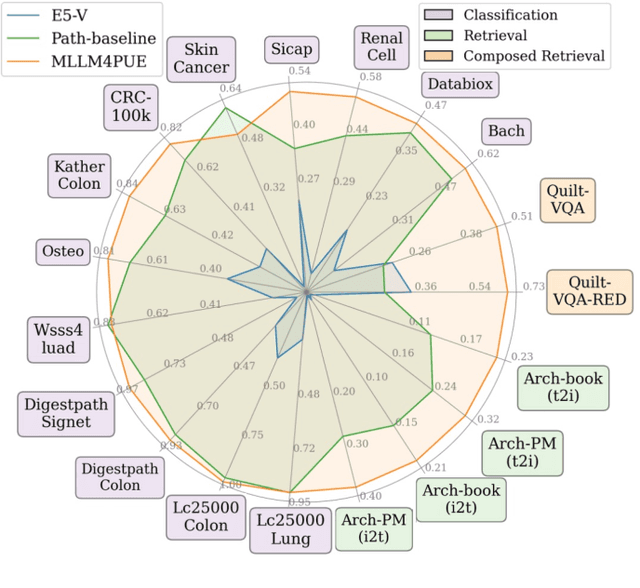
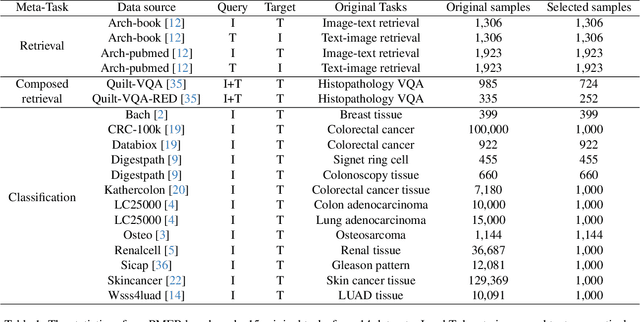
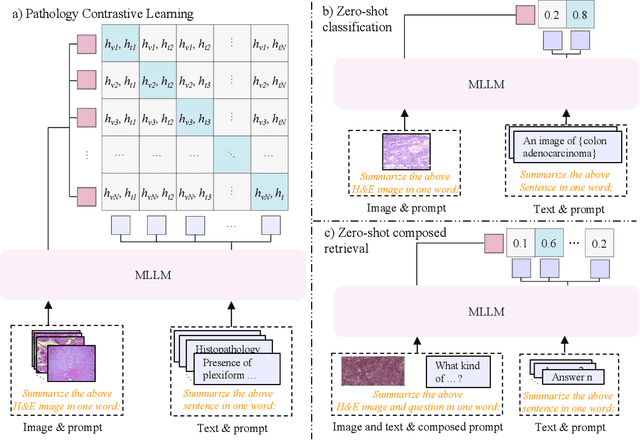
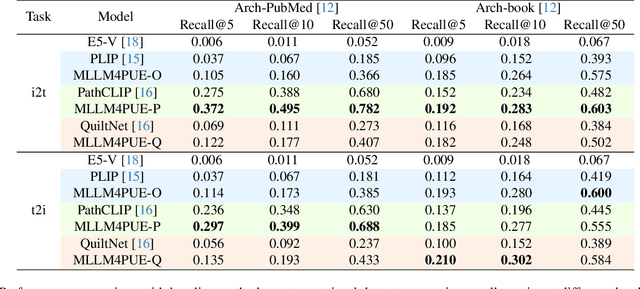
Abstract:Pathology plays a critical role in diagnosing a wide range of diseases, yet existing approaches often rely heavily on task-specific models trained on extensive, well-labeled datasets. These methods face sustainability challenges due to the diversity of pathologies and the labor-intensive nature of data collection. To address these limitations, we highlight the need for universal multimodal embeddings that can support multiple downstream tasks. Previous approaches often involve fine-tuning CLIP-based models, which handle images and text separately, limiting their ability to capture complex multimodal relationships. Additionally, these models are evaluated across diverse datasets without a unified benchmark for assessing multimodal embeddings in pathology. To address these challenges, we propose MLLM4PUE, a novel framework that leverages Multimodal Large Language Models (MLLMs) to generate Pathology Universal Embeddings. The MLLM4PUE framework not only facilitates robust integration of images and text but also enhances understanding and fusion capabilities across various tasks. We further introduce the Pathology Multimodal Embedding Benchmark (PMEB), a comprehensive benchmark designed to assess the quality of pathology multimodal embeddings. PMEB comprises 15 original tasks drawn from 14 datasets, organized into three meta-tasks: retrieval, classification, and composed retrieval. Experimental results demonstrate the superiority of MLLM4PUE, illustrating MLLM-based models can effectively support a wide range of downstream tasks and unify the research direction for foundation models in pathology.
GoBERT: Gene Ontology Graph Informed BERT for Universal Gene Function Prediction
Jan 03, 2025Abstract:Exploring the functions of genes and gene products is crucial to a wide range of fields, including medical research, evolutionary biology, and environmental science. However, discovering new functions largely relies on expensive and exhaustive wet lab experiments. Existing methods of automatic function annotation or prediction mainly focus on protein function prediction with sequence, 3D-structures or protein family information. In this study, we propose to tackle the gene function prediction problem by exploring Gene Ontology graph and annotation with BERT (GoBERT) to decipher the underlying relationships among gene functions. Our proposed novel function prediction task utilizes existing functions as inputs and generalizes the function prediction to gene and gene products. Specifically, two pre-train tasks are designed to jointly train GoBERT to capture both explicit and implicit relations of functions. Neighborhood prediction is a self-supervised multi-label classification task that captures the explicit function relations. Specified masking and recovering task helps GoBERT in finding implicit patterns among functions. The pre-trained GoBERT possess the ability to predict novel functions for various gene and gene products based on known functional annotations. Extensive experiments, biological case studies, and ablation studies are conducted to demonstrate the superiority of our proposed GoBERT.
Compositional Image Retrieval via Instruction-Aware Contrastive Learning
Dec 07, 2024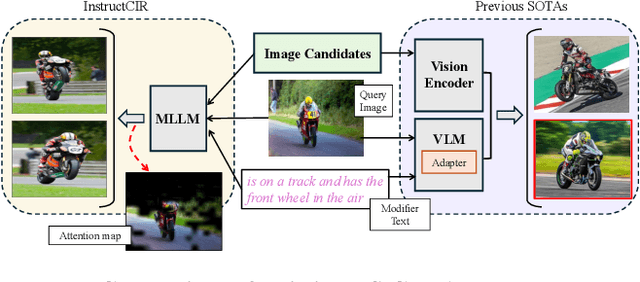
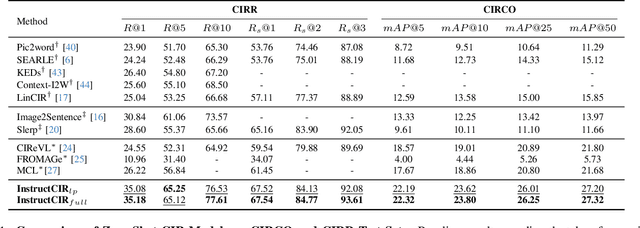
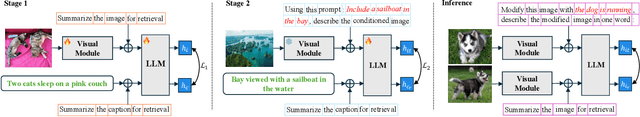
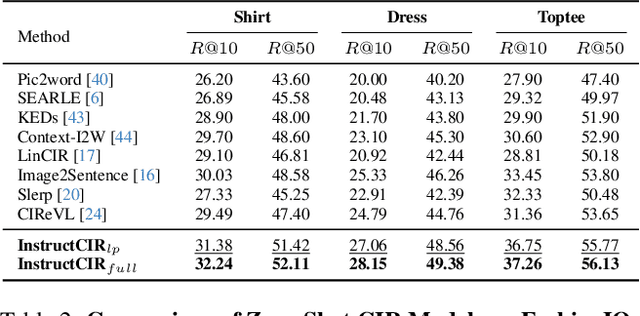
Abstract:Composed Image Retrieval (CIR) involves retrieving a target image based on a composed query of an image paired with text that specifies modifications or changes to the visual reference. CIR is inherently an instruction-following task, as the model needs to interpret and apply modifications to the image. In practice, due to the scarcity of annotated data in downstream tasks, Zero-Shot CIR (ZS-CIR) is desirable. While existing ZS-CIR models based on CLIP have shown promising results, their capability in interpreting and following modification instructions remains limited. Some research attempts to address this by incorporating Large Language Models (LLMs). However, these approaches still face challenges in effectively integrating multimodal information and instruction understanding. To tackle above challenges, we propose a novel embedding method utilizing an instruction-tuned Multimodal LLM (MLLM) to generate composed representation, which significantly enhance the instruction following capability for a comprehensive integration between images and instructions. Nevertheless, directly applying MLLMs introduces a new challenge since MLLMs are primarily designed for text generation rather than embedding extraction as required in CIR. To address this, we introduce a two-stage training strategy to efficiently learn a joint multimodal embedding space and further refining the ability to follow modification instructions by tuning the model in a triplet dataset similar to the CIR format. Extensive experiments on four public datasets: FashionIQ, CIRR, GeneCIS, and CIRCO demonstrates the superior performance of our model, outperforming state-of-the-art baselines by a significant margin. Codes are available at the GitHub repository.
PathM3: A Multimodal Multi-Task Multiple Instance Learning Framework for Whole Slide Image Classification and Captioning
Mar 13, 2024Abstract:In the field of computational histopathology, both whole slide images (WSIs) and diagnostic captions provide valuable insights for making diagnostic decisions. However, aligning WSIs with diagnostic captions presents a significant challenge. This difficulty arises from two main factors: 1) Gigapixel WSIs are unsuitable for direct input into deep learning models, and the redundancy and correlation among the patches demand more attention; and 2) Authentic WSI diagnostic captions are extremely limited, making it difficult to train an effective model. To overcome these obstacles, we present PathM3, a multimodal, multi-task, multiple instance learning (MIL) framework for WSI classification and captioning. PathM3 adapts a query-based transformer to effectively align WSIs with diagnostic captions. Given that histopathology visual patterns are redundantly distributed across WSIs, we aggregate each patch feature with MIL method that considers the correlations among instances. Furthermore, our PathM3 overcomes data scarcity in WSI-level captions by leveraging limited WSI diagnostic caption data in the manner of multi-task joint learning. Extensive experiments with improved classification accuracy and caption generation demonstrate the effectiveness of our method on both WSI classification and captioning task.
Segment Any Cell: A SAM-based Auto-prompting Fine-tuning Framework for Nuclei Segmentation
Jan 24, 2024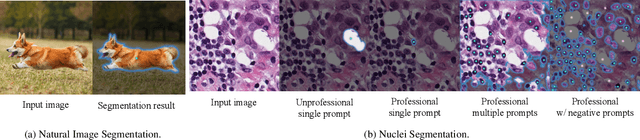


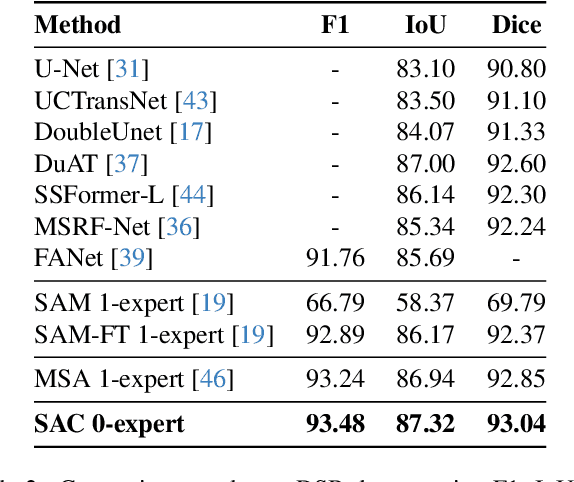
Abstract:In the rapidly evolving field of AI research, foundational models like BERT and GPT have significantly advanced language and vision tasks. The advent of pretrain-prompting models such as ChatGPT and Segmentation Anything Model (SAM) has further revolutionized image segmentation. However, their applications in specialized areas, particularly in nuclei segmentation within medical imaging, reveal a key challenge: the generation of high-quality, informative prompts is as crucial as applying state-of-the-art (SOTA) fine-tuning techniques on foundation models. To address this, we introduce Segment Any Cell (SAC), an innovative framework that enhances SAM specifically for nuclei segmentation. SAC integrates a Low-Rank Adaptation (LoRA) within the attention layer of the Transformer to improve the fine-tuning process, outperforming existing SOTA methods. It also introduces an innovative auto-prompt generator that produces effective prompts to guide segmentation, a critical factor in handling the complexities of nuclei segmentation in biomedical imaging. Our extensive experiments demonstrate the superiority of SAC in nuclei segmentation tasks, proving its effectiveness as a tool for pathologists and researchers. Our contributions include a novel prompt generation strategy, automated adaptability for diverse segmentation tasks, the innovative application of Low-Rank Attention Adaptation in SAM, and a versatile framework for semantic segmentation challenges.
ChatGraph: Interpretable Text Classification by Converting ChatGPT Knowledge to Graphs
May 03, 2023



Abstract:ChatGPT, as a recently launched large language model (LLM), has shown superior performance in various natural language processing (NLP) tasks. However, two major limitations hinder its potential applications: (1) the inflexibility of finetuning on downstream tasks and (2) the lack of interpretability in the decision-making process. To tackle these limitations, we propose a novel framework that leverages the power of ChatGPT for specific tasks, such as text classification, while improving its interpretability. The proposed framework conducts a knowledge graph extraction task to extract refined and structural knowledge from the raw data using ChatGPT. The rich knowledge is then converted into a graph, which is further used to train an interpretable linear classifier to make predictions. To evaluate the effectiveness of our proposed method, we conduct experiments on four datasets. The result shows that our method can significantly improve the performance compared to directly utilizing ChatGPT for text classification tasks. And our method provides a more transparent decision-making process compared with previous text classification methods.
Exploring Robustness of Unsupervised Domain Adaptation in Semantic Segmentation
May 23, 2021



Abstract:Recent studies imply that deep neural networks are vulnerable to adversarial examples -- inputs with a slight but intentional perturbation are incorrectly classified by the network. Such vulnerability makes it risky for some security-related applications (e.g., semantic segmentation in autonomous cars) and triggers tremendous concerns on the model reliability. For the first time, we comprehensively evaluate the robustness of existing UDA methods and propose a robust UDA approach. It is rooted in two observations: (i) the robustness of UDA methods in semantic segmentation remains unexplored, which pose a security concern in this field; and (ii) although commonly used self-supervision (e.g., rotation and jigsaw) benefits image tasks such as classification and recognition, they fail to provide the critical supervision signals that could learn discriminative representation for segmentation tasks. These observations motivate us to propose adversarial self-supervision UDA (or ASSUDA) that maximizes the agreement between clean images and their adversarial examples by a contrastive loss in the output space. Extensive empirical studies on commonly used benchmarks demonstrate that ASSUDA is resistant to adversarial attacks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge