Francesco Giganti
Collaborative Quantization Embeddings for Intra-Subject Prostate MR Image Registration
Jul 14, 2022
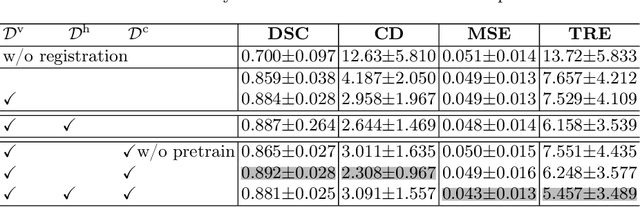
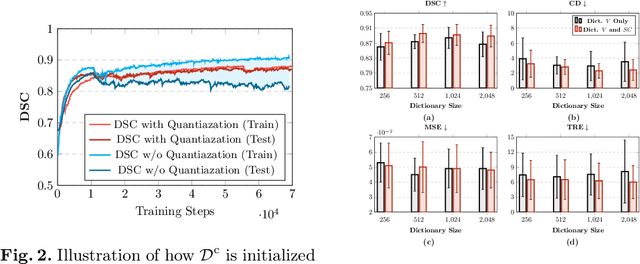
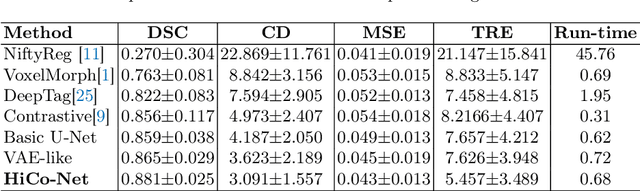
Abstract:Image registration is useful for quantifying morphological changes in longitudinal MR images from prostate cancer patients. This paper describes a development in improving the learning-based registration algorithms, for this challenging clinical application often with highly variable yet limited training data. First, we report that the latent space can be clustered into a much lower dimensional space than that commonly found as bottleneck features at the deep layer of a trained registration network. Based on this observation, we propose a hierarchical quantization method, discretizing the learned feature vectors using a jointly-trained dictionary with a constrained size, in order to improve the generalisation of the registration networks. Furthermore, a novel collaborative dictionary is independently optimised to incorporate additional prior information, such as the segmentation of the gland or other regions of interest, in the latent quantized space. Based on 216 real clinical images from 86 prostate cancer patients, we show the efficacy of both the designed components. Improved registration accuracy was obtained with statistical significance, in terms of both Dice on gland and target registration error on corresponding landmarks, the latter of which achieved 5.46 mm, an improvement of 28.7\% from the baseline without quantization. Experimental results also show that the difference in performance was indeed minimised between training and testing data.
Image quality assessment by overlapping task-specific and task-agnostic measures: application to prostate multiparametric MR images for cancer segmentation
Feb 20, 2022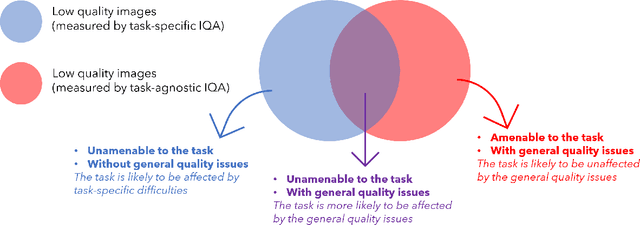
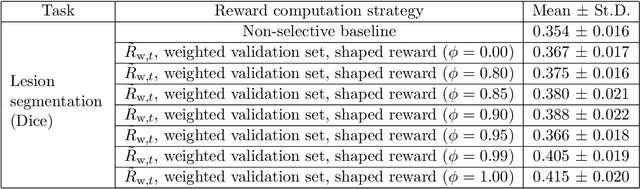


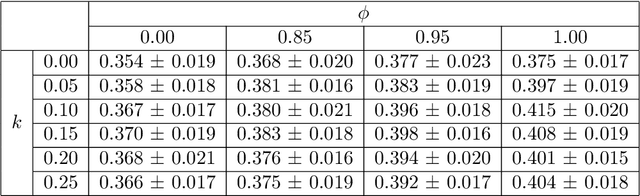
Abstract:Image quality assessment (IQA) in medical imaging can be used to ensure that downstream clinical tasks can be reliably performed. Quantifying the impact of an image on the specific target tasks, also named as task amenability, is needed. A task-specific IQA has recently been proposed to learn an image-amenability-predicting controller simultaneously with a target task predictor. This allows for the trained IQA controller to measure the impact an image has on the target task performance, when this task is performed using the predictor, e.g. segmentation and classification neural networks in modern clinical applications. In this work, we propose an extension to this task-specific IQA approach, by adding a task-agnostic IQA based on auto-encoding as the target task. Analysing the intersection between low-quality images, deemed by both the task-specific and task-agnostic IQA, may help to differentiate the underpinning factors that caused the poor target task performance. For example, common imaging artefacts may not adversely affect the target task, which would lead to a low task-agnostic quality and a high task-specific quality, whilst individual cases considered clinically challenging, which can not be improved by better imaging equipment or protocols, is likely to result in a high task-agnostic quality but a low task-specific quality. We first describe a flexible reward shaping strategy which allows for the adjustment of weighting between task-agnostic and task-specific quality scoring. Furthermore, we evaluate the proposed algorithm using a clinically challenging target task of prostate tumour segmentation on multiparametric magnetic resonance (mpMR) images, from 850 patients. The proposed reward shaping strategy, with appropriately weighted task-specific and task-agnostic qualities, successfully identified samples that need re-acquisition due to defected imaging process.
Unsupervised Domain Adaptation with Semantic Consistency across Heterogeneous Modalities for MRI Prostate Lesion Segmentation
Sep 19, 2021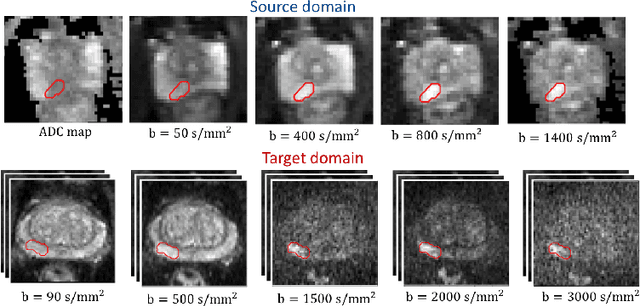
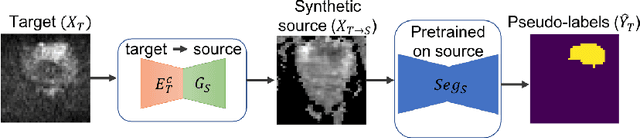

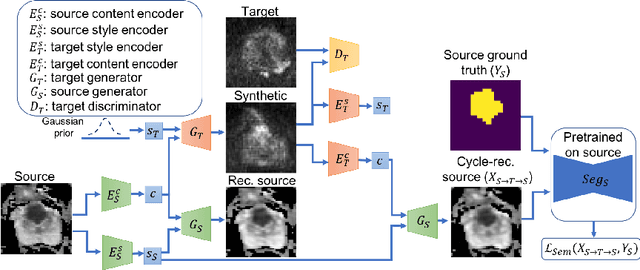

Abstract:Any novel medical imaging modality that differs from previous protocols e.g. in the number of imaging channels, introduces a new domain that is heterogeneous from previous ones. This common medical imaging scenario is rarely considered in the domain adaptation literature, which handles shifts across domains of the same dimensionality. In our work we rely on stochastic generative modeling to translate across two heterogeneous domains at pixel space and introduce two new loss functions that promote semantic consistency. Firstly, we introduce a semantic cycle-consistency loss in the source domain to ensure that the translation preserves the semantics. Secondly, we introduce a pseudo-labelling loss, where we translate target data to source, label them by a source-domain network, and use the generated pseudo-labels to supervise the target-domain network. Our results show that this allows us to extract systematically better representations for the target domain. In particular, we address the challenge of enhancing performance on VERDICT-MRI, an advanced diffusion-weighted imaging technique, by exploiting labeled mp-MRI data. When compared to several unsupervised domain adaptation approaches, our approach yields substantial improvements, that consistently carry over to the semi-supervised and supervised learning settings.
Morphological Change Forecasting for Prostate Glands using Feature-based Registration and Kernel Density Extrapolation
Jan 16, 2021
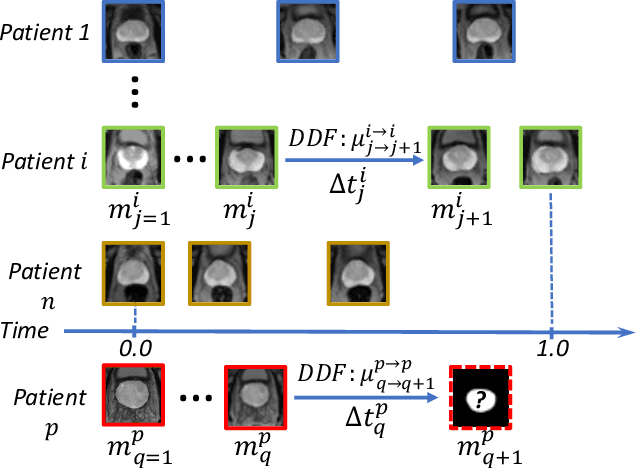


Abstract:Organ morphology is a key indicator for prostate disease diagnosis and prognosis. For instance, In longitudinal study of prostate cancer patients under active surveillance, the volume, boundary smoothness and their changes are closely monitored on time-series MR image data. In this paper, we describe a new framework for forecasting prostate morphological changes, as the ability to detect such changes earlier than what is currently possible may enable timely treatment or avoiding unnecessary confirmatory biopsies. In this work, an efficient feature-based MR image registration is first developed to align delineated prostate gland capsules to quantify the morphological changes using the inferred dense displacement fields (DDFs). We then propose to use kernel density estimation (KDE) of the probability density of the DDF-represented \textit{future morphology changes}, between current and future time points, before the future data become available. The KDE utilises a novel distance function that takes into account morphology, stage-of-progression and duration-of-change, which are considered factors in such subject-specific forecasting. We validate the proposed approach on image masks unseen to registration network training, without using any data acquired at the future target time points. The experiment results are presented on a longitudinal data set with 331 images from 73 patients, yielding an average Dice score of 0.865 on a holdout set, between the ground-truth and the image masks warped by the KDE-predicted-DDFs.
Harnessing Uncertainty in Domain Adaptation for MRI Prostate Lesion Segmentation
Oct 14, 2020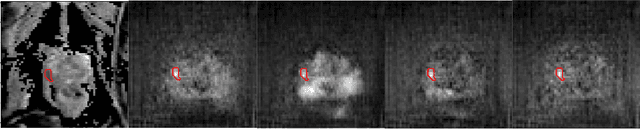
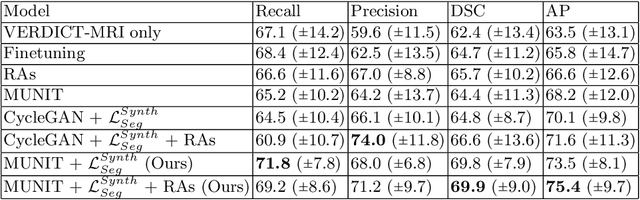
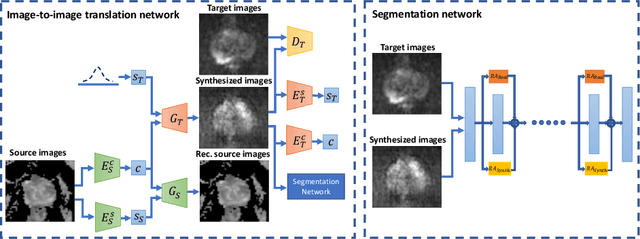
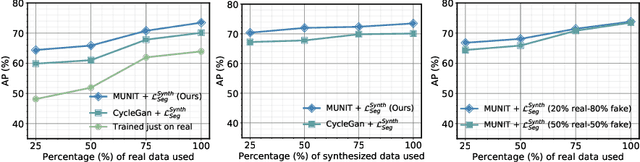
Abstract:The need for training data can impede the adoption of novel imaging modalities for learning-based medical image analysis. Domain adaptation methods partially mitigate this problem by translating training data from a related source domain to a novel target domain, but typically assume that a one-to-one translation is possible. Our work addresses the challenge of adapting to a more informative target domain where multiple target samples can emerge from a single source sample. In particular we consider translating from mp-MRI to VERDICT, a richer MRI modality involving an optimized acquisition protocol for cancer characterization. We explicitly account for the inherent uncertainty of this mapping and exploit it to generate multiple outputs conditioned on a single input. Our results show that this allows us to extract systematically better image representations for the target domain, when used in tandem with both simple, CycleGAN-based baselines, as well as more powerful approaches that integrate discriminative segmentation losses and/or residual adapters. When compared to its deterministic counterparts, our approach yields substantial improvements across a broad range of dataset sizes, increasingly strong baselines, and evaluation measures.
Longitudinal Image Registration with Temporal-order and Subject-specificity Discrimination
Aug 29, 2020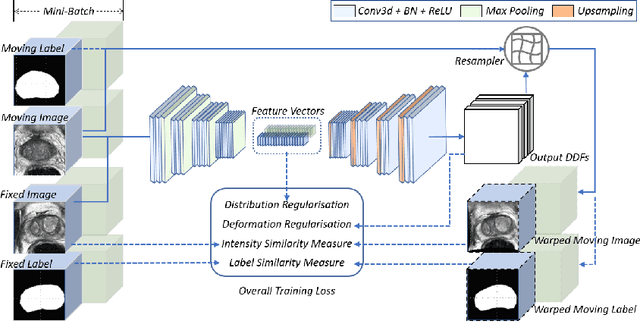

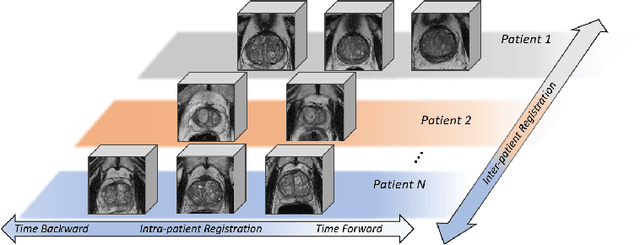

Abstract:Morphological analysis of longitudinal MR images plays a key role in monitoring disease progression for prostate cancer patients, who are placed under an active surveillance program. In this paper, we describe a learning-based image registration algorithm to quantify changes on regions of interest between a pair of images from the same patient, acquired at two different time points. Combining intensity-based similarity and gland segmentation as weak supervision, the population-data-trained registration networks significantly lowered the target registration errors (TREs) on holdout patient data, compared with those before registration and those from an iterative registration algorithm. Furthermore, this work provides a quantitative analysis on several longitudinal-data-sampling strategies and, in turn, we propose a novel regularisation method based on maximum mean discrepancy, between differently-sampled training image pairs. Based on 216 3D MR images from 86 patients, we report a mean TRE of 5.6 mm and show statistically significant differences between the different training data sampling strategies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge