Eleni Chiou
Tissue Aware Nuclei Detection and Classification Model for Histopathology Images
Nov 17, 2025Abstract:Accurate nuclei detection and classification are fundamental to computational pathology, yet existing approaches are hindered by reliance on detailed expert annotations and insufficient use of tissue context. We present Tissue-Aware Nuclei Detection (TAND), a novel framework achieving joint nuclei detection and classification using point-level supervision enhanced by tissue mask conditioning. TAND couples a ConvNeXt-based encoder-decoder with a frozen Virchow-2 tissue segmentation branch, where semantic tissue probabilities selectively modulate the classification stream through a novel multi-scale Spatial Feature-wise Linear Modulation (Spatial-FiLM). On the PUMA benchmark, TAND achieves state-of-the-art performance, surpassing both tissue-agnostic baselines and mask-supervised methods. Notably, our approach demonstrates remarkable improvements in tissue-dependent cell types such as epithelium, endothelium, and stroma. To the best of our knowledge, this is the first method to condition per-cell classification on learned tissue masks, offering a practical pathway to reduce annotation burden.
Beyond Deterministic Translation for Unsupervised Domain Adaptation
Mar 11, 2022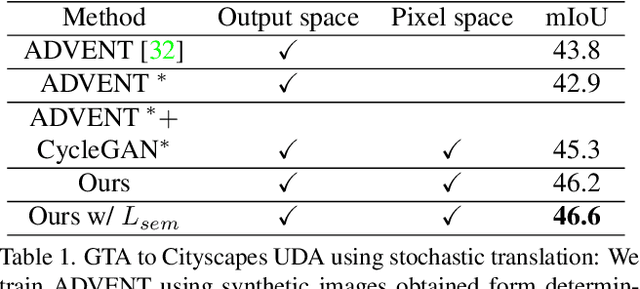
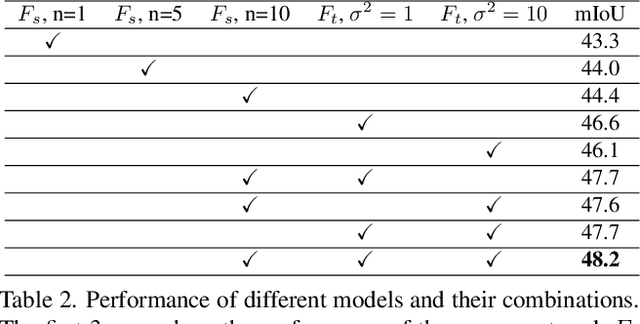



Abstract:In this work we challenge the common approach of using a one-to-one mapping ('translation') between the source and target domains in unsupervised domain adaptation (UDA). Instead, we rely on stochastic translation to capture inherent translation ambiguities. This allows us to (i) train more accurate target networks by generating multiple outputs conditioned on the same source image, leveraging both accurate translation and data augmentation for appearance variability, (ii) impute robust pseudo-labels for the target data by averaging the predictions of a source network on multiple translated versions of a single target image and (iii) train and ensemble diverse networks in the target domain by modulating the degree of stochasticity in the translations. We report improvements over strong recent baselines, leading to state-of-the-art UDA results on two challenging semantic segmentation benchmarks.
Unsupervised Domain Adaptation with Semantic Consistency across Heterogeneous Modalities for MRI Prostate Lesion Segmentation
Sep 19, 2021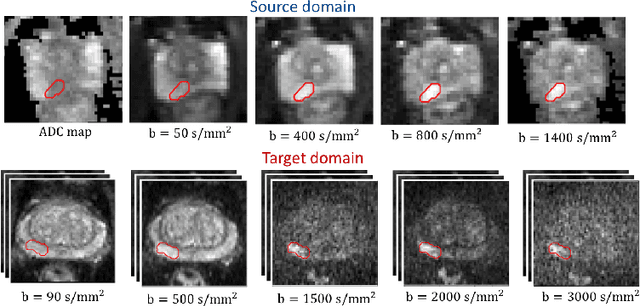
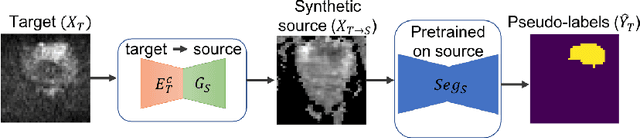

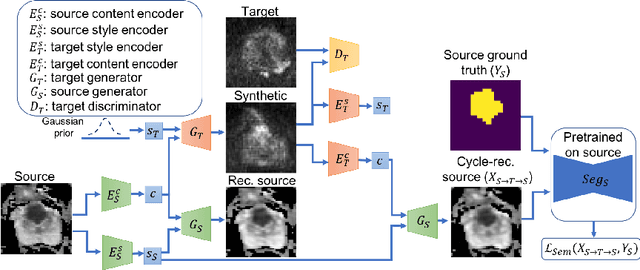

Abstract:Any novel medical imaging modality that differs from previous protocols e.g. in the number of imaging channels, introduces a new domain that is heterogeneous from previous ones. This common medical imaging scenario is rarely considered in the domain adaptation literature, which handles shifts across domains of the same dimensionality. In our work we rely on stochastic generative modeling to translate across two heterogeneous domains at pixel space and introduce two new loss functions that promote semantic consistency. Firstly, we introduce a semantic cycle-consistency loss in the source domain to ensure that the translation preserves the semantics. Secondly, we introduce a pseudo-labelling loss, where we translate target data to source, label them by a source-domain network, and use the generated pseudo-labels to supervise the target-domain network. Our results show that this allows us to extract systematically better representations for the target domain. In particular, we address the challenge of enhancing performance on VERDICT-MRI, an advanced diffusion-weighted imaging technique, by exploiting labeled mp-MRI data. When compared to several unsupervised domain adaptation approaches, our approach yields substantial improvements, that consistently carry over to the semi-supervised and supervised learning settings.
Harnessing Uncertainty in Domain Adaptation for MRI Prostate Lesion Segmentation
Oct 14, 2020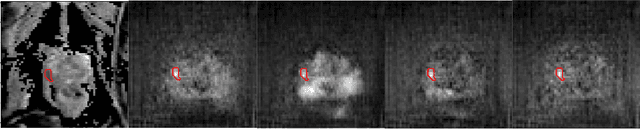
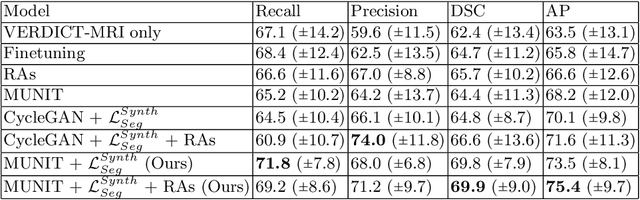
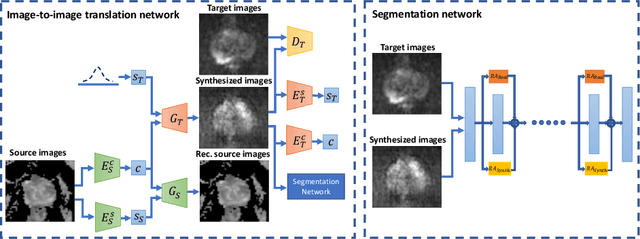
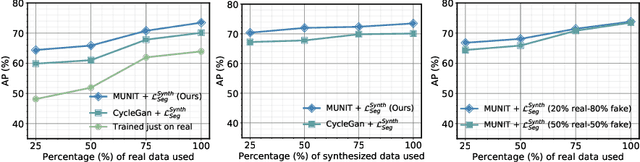
Abstract:The need for training data can impede the adoption of novel imaging modalities for learning-based medical image analysis. Domain adaptation methods partially mitigate this problem by translating training data from a related source domain to a novel target domain, but typically assume that a one-to-one translation is possible. Our work addresses the challenge of adapting to a more informative target domain where multiple target samples can emerge from a single source sample. In particular we consider translating from mp-MRI to VERDICT, a richer MRI modality involving an optimized acquisition protocol for cancer characterization. We explicitly account for the inherent uncertainty of this mapping and exploit it to generate multiple outputs conditioned on a single input. Our results show that this allows us to extract systematically better image representations for the target domain, when used in tandem with both simple, CycleGAN-based baselines, as well as more powerful approaches that integrate discriminative segmentation losses and/or residual adapters. When compared to its deterministic counterparts, our approach yields substantial improvements across a broad range of dataset sizes, increasingly strong baselines, and evaluation measures.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge