Eva Schnider
Plots Unlock Time-Series Understanding in Multimodal Models
Oct 03, 2024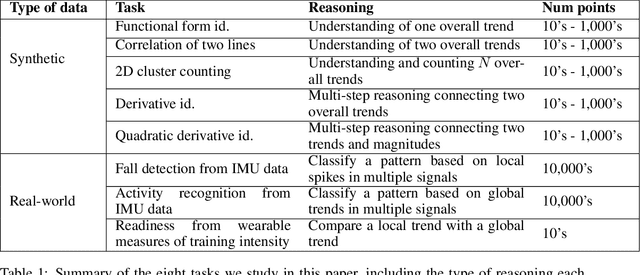

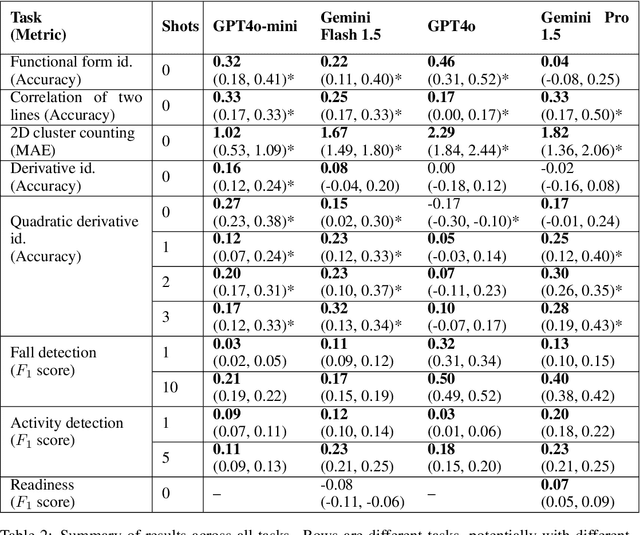
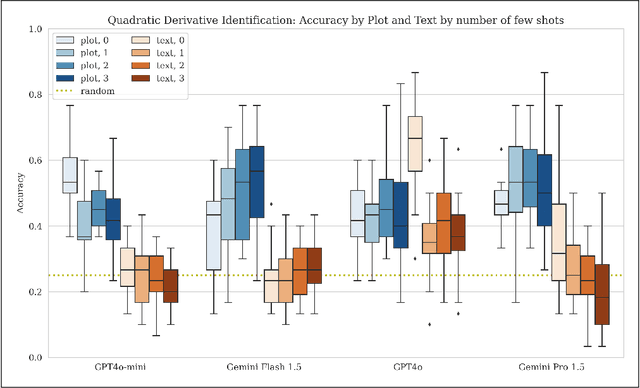
Abstract:While multimodal foundation models can now natively work with data beyond text, they remain underutilized in analyzing the considerable amounts of multi-dimensional time-series data in fields like healthcare, finance, and social sciences, representing a missed opportunity for richer, data-driven insights. This paper proposes a simple but effective method that leverages the existing vision encoders of these models to "see" time-series data via plots, avoiding the need for additional, potentially costly, model training. Our empirical evaluations show that this approach outperforms providing the raw time-series data as text, with the additional benefit that visual time-series representations demonstrate up to a 90% reduction in model API costs. We validate our hypothesis through synthetic data tasks of increasing complexity, progressing from simple functional form identification on clean data, to extracting trends from noisy scatter plots. To demonstrate generalizability from synthetic tasks with clear reasoning steps to more complex, real-world scenarios, we apply our approach to consumer health tasks - specifically fall detection, activity recognition, and readiness assessment - which involve heterogeneous, noisy data and multi-step reasoning. The overall success in plot performance over text performance (up to an 120% performance increase on zero-shot synthetic tasks, and up to 150% performance increase on real-world tasks), across both GPT and Gemini model families, highlights our approach's potential for making the best use of the native capabilities of foundation models.
MINT: A wrapper to make multi-modal and multi-image AI models interactive
Jan 22, 2024Abstract:During the diagnostic process, doctors incorporate multimodal information including imaging and the medical history - and similarly medical AI development has increasingly become multimodal. In this paper we tackle a more subtle challenge: doctors take a targeted medical history to obtain only the most pertinent pieces of information; how do we enable AI to do the same? We develop a wrapper method named MINT (Make your model INTeractive) that automatically determines what pieces of information are most valuable at each step, and ask for only the most useful information. We demonstrate the efficacy of MINT wrapping a skin disease prediction model, where multiple images and a set of optional answers to $25$ standard metadata questions (i.e., structured medical history) are used by a multi-modal deep network to provide a differential diagnosis. We show that MINT can identify whether metadata inputs are needed and if so, which question to ask next. We also demonstrate that when collecting multiple images, MINT can identify if an additional image would be beneficial, and if so, which type of image to capture. We showed that MINT reduces the number of metadata and image inputs needed by 82% and 36.2% respectively, while maintaining predictive performance. Using real-world AI dermatology system data, we show that needing fewer inputs can retain users that may otherwise fail to complete the system submission and drop off without a diagnosis. Qualitative examples show MINT can closely mimic the step-by-step decision making process of a clinical workflow and how this is different for straight forward cases versus more difficult, ambiguous cases. Finally we demonstrate how MINT is robust to different underlying multi-model classifiers and can be easily adapted to user requirements without significant model re-training.
Improved distinct bone segmentation in upper-body CT through multi-resolution networks
Jan 31, 2023Abstract:Purpose: Automated distinct bone segmentation from CT scans is widely used in planning and navigation workflows. U-Net variants are known to provide excellent results in supervised semantic segmentation. However, in distinct bone segmentation from upper body CTs a large field of view and a computationally taxing 3D architecture are required. This leads to low-resolution results lacking detail or localisation errors due to missing spatial context when using high-resolution inputs. Methods: We propose to solve this problem by using end-to-end trainable segmentation networks that combine several 3D U-Nets working at different resolutions. Our approach, which extends and generalizes HookNet and MRN, captures spatial information at a lower resolution and skips the encoded information to the target network, which operates on smaller high-resolution inputs. We evaluated our proposed architecture against single resolution networks and performed an ablation study on information concatenation and the number of context networks. Results: Our proposed best network achieves a median DSC of 0.86 taken over all 125 segmented bone classes and reduces the confusion among similar-looking bones in different locations. These results outperform our previously published 3D U-Net baseline results on the task and distinct-bone segmentation results reported by other groups. Conclusion: The presented multi-resolution 3D U-Nets address current shortcomings in bone segmentation from upper-body CT scans by allowing for capturing a larger field of view while avoiding the cubic growth of the input pixels and intermediate computations that quickly outgrow the computational capacities in 3D. The approach thus improves the accuracy and efficiency of distinct bone segmentation from upper-body CT.
Ensemble uncertainty as a criterion for dataset expansion in distinct bone segmentation from upper-body CT images
Aug 19, 2022
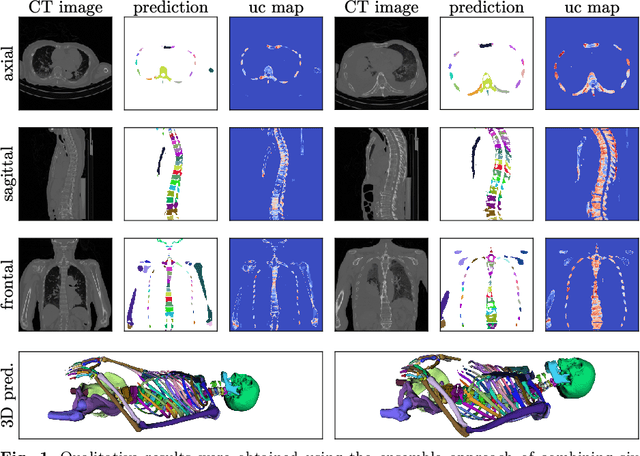
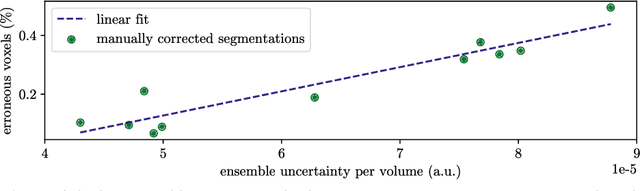
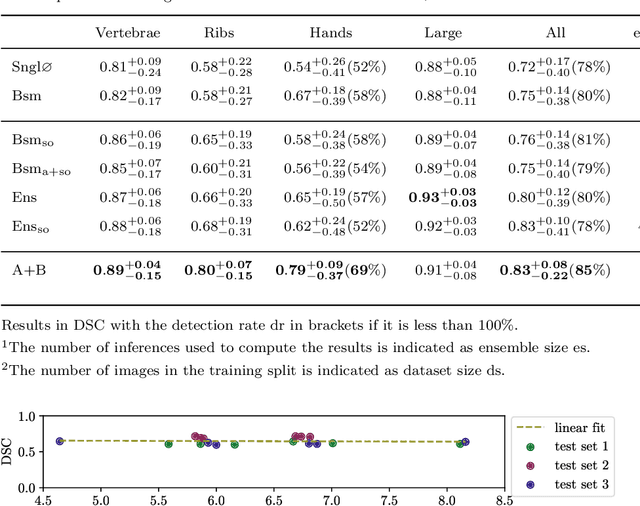
Abstract:Purpose: The localisation and segmentation of individual bones is an important preprocessing step in many planning and navigation applications. It is, however, a time-consuming and repetitive task if done manually. This is true not only for clinical practice but also for the acquisition of training data. We therefore not only present an end-to-end learnt algorithm that is capable of segmenting 125 distinct bones in an upper-body CT, but also provide an ensemble-based uncertainty measure that helps to single out scans to enlarge the training dataset with. Methods We create fully automated end-to-end learnt segmentations using a neural network architecture inspired by the 3D-Unet and fully supervised training. The results are improved using ensembles and inference-time augmentation. We examine the relationship of ensemble-uncertainty to an unlabelled scan's prospective usefulness as part of the training dataset. Results: Our methods are evaluated on an in-house dataset of 16 upper-body CT scans with a resolution of \SI{2}{\milli\meter} per dimension. Taking into account all 125 bones in our label set, our most successful ensemble achieves a median dice score coefficient of 0.83. We find a lack of correlation between a scan's ensemble uncertainty and its prospective influence on the accuracies achieved within an enlarged training set. At the same time, we show that the ensemble uncertainty correlates to the number of voxels that need manual correction after an initial automated segmentation, thus minimising the time required to finalise a new ground truth segmentation. Conclusion: In combination, scans with low ensemble uncertainty need less annotator time while yielding similar future DSC improvements. They are thus ideal candidates to enlarge a training set for upper-body distinct bone segmentation from CT scans. }
Maintaining fairness across distribution shift: do we have viable solutions for real-world applications?
Feb 02, 2022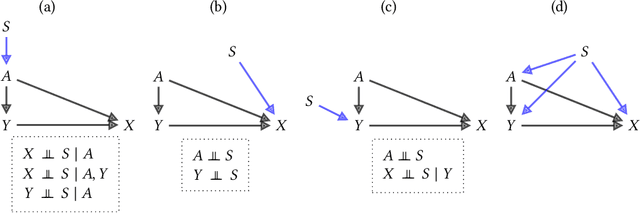
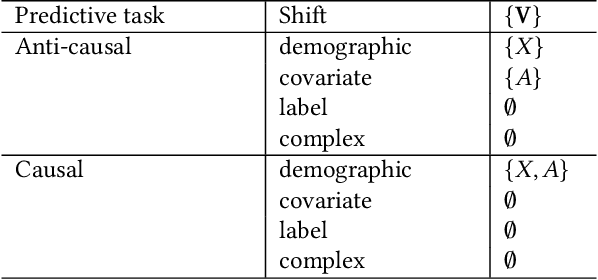
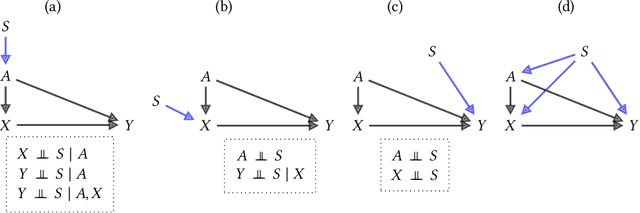
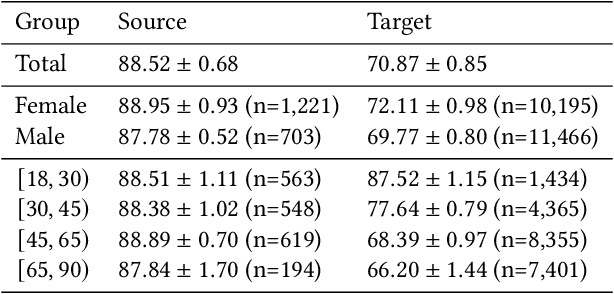
Abstract:Fairness and robustness are often considered as orthogonal dimensions when evaluating machine learning models. However, recent work has revealed interactions between fairness and robustness, showing that fairness properties are not necessarily maintained under distribution shift. In healthcare settings, this can result in e.g. a model that performs fairly according to a selected metric in "hospital A" showing unfairness when deployed in "hospital B". While a nascent field has emerged to develop provable fair and robust models, it typically relies on strong assumptions about the shift, limiting its impact for real-world applications. In this work, we explore the settings in which recently proposed mitigation strategies are applicable by referring to a causal framing. Using examples of predictive models in dermatology and electronic health records, we show that real-world applications are complex and often invalidate the assumptions of such methods. Our work hence highlights technical, practical, and engineering gaps that prevent the development of robustly fair machine learning models for real-world applications. Finally, we discuss potential remedies at each step of the machine learning pipeline.
3D Segmentation Networks for Excessive Numbers of Classes: Distinct Bone Segmentation in Upper Bodies
Oct 14, 2020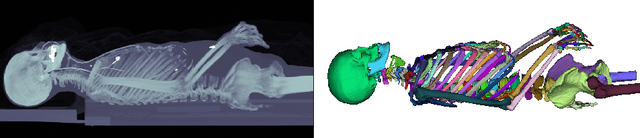
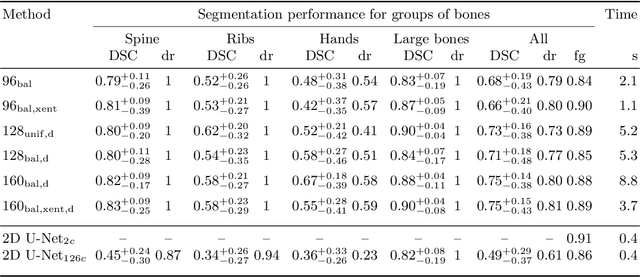
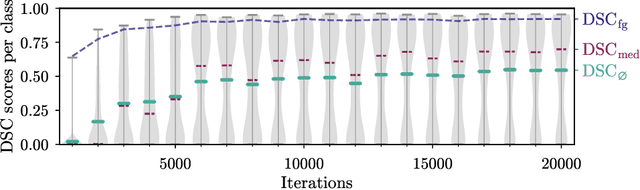
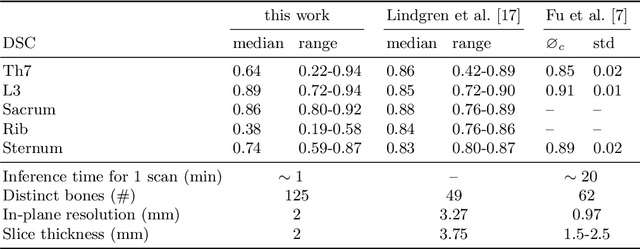
Abstract:Segmentation of distinct bones plays a crucial role in diagnosis, planning, navigation, and the assessment of bone metastasis. It supplies semantic knowledge to visualisation tools for the planning of surgical interventions and the education of health professionals. Fully supervised segmentation of 3D data using Deep Learning methods has been extensively studied for many tasks but is usually restricted to distinguishing only a handful of classes. With 125 distinct bones, our case includes many more labels than typical 3D segmentation tasks. For this reason, the direct adaptation of most established methods is not possible. This paper discusses the intricacies of training a 3D segmentation network in a many-label setting and shows necessary modifications in network architecture, loss function, and data augmentation. As a result, we demonstrate the robustness of our method by automatically segmenting over one hundred distinct bones simultaneously in an end-to-end learnt fashion from a CT-scan.
* 10 pages, 3 figures, 2 tables, accepted into MICCAI 2020 International Workshop on Machine Learning in Medical Imaging
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge