Vishal Dey
AutoSDT: Scaling Data-Driven Discovery Tasks Toward Open Co-Scientists
Jun 09, 2025



Abstract:Despite long-standing efforts in accelerating scientific discovery with AI, building AI co-scientists remains challenging due to limited high-quality data for training and evaluation. To tackle this data scarcity issue, we present AutoSDT, an automatic pipeline that collects high-quality coding tasks in real-world data-driven discovery workflows. AutoSDT leverages the coding capabilities and parametric knowledge of LLMs to search for diverse sources, select ecologically valid tasks, and synthesize accurate task instructions and code solutions. Using our pipeline, we construct AutoSDT-5K, a dataset of 5,404 coding tasks for data-driven discovery that covers four scientific disciplines and 756 unique Python packages. To the best of our knowledge, AutoSDT-5K is the only automatically collected and the largest open dataset for data-driven scientific discovery. Expert feedback on a subset of 256 tasks shows the effectiveness of AutoSDT: 93% of the collected tasks are ecologically valid, and 92.2% of the synthesized programs are functionally correct. Trained on AutoSDT-5K, the Qwen2.5-Coder-Instruct LLM series, dubbed AutoSDT-Coder, show substantial improvement on two challenging data-driven discovery benchmarks, ScienceAgentBench and DiscoveryBench. Most notably, AutoSDT-Coder-32B reaches the same level of performance as GPT-4o on ScienceAgentBench with a success rate of 7.8%, doubling the performance of its base model. On DiscoveryBench, it lifts the hypothesis matching score to 8.1, bringing a 17.4% relative improvement and closing the gap between open-weight models and GPT-4o.
Large Language Models for Controllable Multi-property Multi-objective Molecule Optimization
May 29, 2025Abstract:In real-world drug design, molecule optimization requires selectively improving multiple molecular properties up to pharmaceutically relevant levels, while maintaining others that already meet such criteria. However, existing computational approaches and instruction-tuned LLMs fail to capture such nuanced property-specific objectives, limiting their practical applicability. To address this, we introduce C-MuMOInstruct, the first instruction-tuning dataset focused on multi-property optimization with explicit, property-specific objectives. Leveraging C-MuMOInstruct, we develop GeLLMO-Cs, a series of instruction-tuned LLMs that can perform targeted property-specific optimization. Our experiments across 5 in-distribution and 5 out-of-distribution tasks show that GeLLMO-Cs consistently outperform strong baselines, achieving up to 126% higher success rate. Notably, GeLLMO-Cs exhibit impressive 0-shot generalization to novel optimization tasks and unseen instructions. This offers a step toward a foundational LLM to support realistic, diverse optimizations with property-specific objectives. C-MuMOInstruct and code are accessible through https://github.com/ninglab/GeLLMO-C.
$\mathtt{GeLLM^3O}$: Generalizing Large Language Models for Multi-property Molecule Optimization
Feb 19, 2025



Abstract:Despite recent advancements, most computational methods for molecule optimization are constrained to single- or double-property optimization tasks and suffer from poor scalability and generalizability to novel optimization tasks. Meanwhile, Large Language Models (LLMs) demonstrate remarkable out-of-domain generalizability to novel tasks. To demonstrate LLMs' potential for molecule optimization, we introduce $\mathtt{MoMUInstruct}$, the first high-quality instruction-tuning dataset specifically focused on complex multi-property molecule optimization tasks. Leveraging $\mathtt{MoMUInstruct}$, we develop $\mathtt{GeLLM^3O}$s, a series of instruction-tuned LLMs for molecule optimization. Extensive evaluations across 5 in-domain and 5 out-of-domain tasks demonstrate that $\mathtt{GeLLM^3O}$s consistently outperform state-of-the-art baselines. $\mathtt{GeLLM^3O}$s also exhibit outstanding zero-shot generalization to unseen tasks, significantly outperforming powerful closed-source LLMs. Such strong generalizability demonstrates the tremendous potential of $\mathtt{GeLLM^3O}$s as foundational models for molecule optimization, thereby tackling novel optimization tasks without resource-intensive retraining. $\mathtt{MoMUInstruct}$, models, and code are accessible through https://github.com/ninglab/GeLLMO.
ScienceAgentBench: Toward Rigorous Assessment of Language Agents for Data-Driven Scientific Discovery
Oct 07, 2024



Abstract:The advancements of language language models (LLMs) have piqued growing interest in developing LLM-based language agents to automate scientific discovery end-to-end, which has sparked both excitement and skepticism about the true capabilities of such agents. In this work, we argue that for an agent to fully automate scientific discovery, it must be able to complete all essential tasks in the workflow. Thus, we call for rigorous assessment of agents on individual tasks in a scientific workflow before making bold claims on end-to-end automation. To this end, we present ScienceAgentBench, a new benchmark for evaluating language agents for data-driven scientific discovery. To ensure the scientific authenticity and real-world relevance of our benchmark, we extract 102 tasks from 44 peer-reviewed publications in four disciplines and engage nine subject matter experts to validate them. We unify the target output for every task to a self-contained Python program file and employ an array of evaluation metrics to examine the generated programs, execution results, and costs. Each task goes through multiple rounds of manual validation by annotators and subject matter experts to ensure its annotation quality and scientific plausibility. We also propose two effective strategies to mitigate data contamination concerns. Using our benchmark, we evaluate five open-weight and proprietary LLMs, each with three frameworks: direct prompting, OpenHands, and self-debug. Given three attempts for each task, the best-performing agent can only solve 32.4% of the tasks independently and 34.3% with expert-provided knowledge. These results underscore the limited capacities of current language agents in generating code for data-driven discovery, let alone end-to-end automation for scientific research.
Enhancing Molecular Property Prediction with Auxiliary Learning and Task-Specific Adaptation
Jan 29, 2024Abstract:Pretrained Graph Neural Networks have been widely adopted for various molecular property prediction tasks. Despite their ability to encode structural and relational features of molecules, traditional fine-tuning of such pretrained GNNs on the target task can lead to poor generalization. To address this, we explore the adaptation of pretrained GNNs to the target task by jointly training them with multiple auxiliary tasks. This could enable the GNNs to learn both general and task-specific features, which may benefit the target task. However, a major challenge is to determine the relatedness of auxiliary tasks with the target task. To address this, we investigate multiple strategies to measure the relevance of auxiliary tasks and integrate such tasks by adaptively combining task gradients or by learning task weights via bi-level optimization. Additionally, we propose a novel gradient surgery-based approach, Rotation of Conflicting Gradients ($\mathtt{RCGrad}$), that learns to align conflicting auxiliary task gradients through rotation. Our experiments with state-of-the-art pretrained GNNs demonstrate the efficacy of our proposed methods, with improvements of up to 7.7% over fine-tuning. This suggests that incorporating auxiliary tasks along with target task fine-tuning can be an effective way to improve the generalizability of pretrained GNNs for molecular property prediction.
Precision Anti-Cancer Drug Selection via Neural Ranking
Jun 30, 2023



Abstract:Personalized cancer treatment requires a thorough understanding of complex interactions between drugs and cancer cell lines in varying genetic and molecular contexts. To address this, high-throughput screening has been used to generate large-scale drug response data, facilitating data-driven computational models. Such models can capture complex drug-cell line interactions across various contexts in a fully data-driven manner. However, accurately prioritizing the most sensitive drugs for each cell line still remains a significant challenge. To address this, we developed neural ranking approaches that leverage large-scale drug response data across multiple cell lines from diverse cancer types. Unlike existing approaches that primarily utilize regression and classification techniques for drug response prediction, we formulated the objective of drug selection and prioritization as a drug ranking problem. In this work, we proposed two neural listwise ranking methods that learn latent representations of drugs and cell lines, and then use those representations to score drugs in each cell line via a learnable scoring function. Specifically, we developed a neural listwise ranking method, List-One, on top of the existing method ListNet. Additionally, we proposed a novel listwise ranking method, List-All, that focuses on all the sensitive drugs instead of the top sensitive drug, unlike List-One. Our results demonstrate that List-All outperforms the best baseline with significant improvements of as much as 8.6% in hit@20 across 50% test cell lines. Furthermore, our analyses suggest that the learned latent spaces from our proposed methods demonstrate informative clustering structures and capture relevant underlying biological features. Moreover, our comprehensive empirical evaluation provides a thorough and objective comparison of the performance of different methods (including our proposed ones).
Improving Compound Activity Classification via Deep Transfer and Representation Learning
Nov 14, 2021


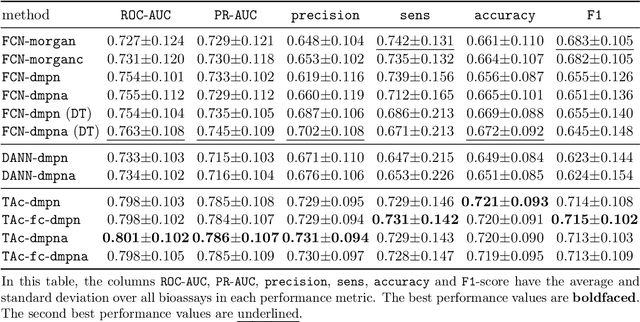
Abstract:Recent advances in molecular machine learning, especially deep neural networks such as Graph Neural Networks (GNNs) for predicting structure activity relationships (SAR) have shown tremendous potential in computer-aided drug discovery. However, the applicability of such deep neural networks are limited by the requirement of large amounts of training data. In order to cope with limited training data for a target task, transfer learning for SAR modeling has been recently adopted to leverage information from data of related tasks. In this work, in contrast to the popular parameter-based transfer learning such as pretraining, we develop novel deep transfer learning methods TAc and TAc-fc to leverage source domain data and transfer useful information to the target domain. TAc learns to generate effective molecular features that can generalize well from one domain to another, and increase the classification performance in the target domain. Additionally, TAc-fc extends TAc by incorporating novel components to selectively learn feature-wise and compound-wise transferability. We used the bioassay screening data from PubChem, and identified 120 pairs of bioassays such that the active compounds in each pair are more similar to each other compared to its inactive compounds. Overall, TAc achieves the best performance with average ROC-AUC of 0.801; it significantly improves ROC-AUC of 83% target tasks with average task-wise performance improvement of 7.102%, compared to the best baseline FCN-dmpna (DT). Our experiments clearly demonstrate that TAc achieves significant improvement over all baselines across a large number of target tasks. Furthermore, although TAc-fc achieves slightly worse ROC-AUC on average compared to TAc (0.798 vs 0.801), TAc-fc still achieves the best performance on more tasks in terms of PR-AUC and F1 compared to other methods.
Adversarial Attacks and Defences: A Survey
Sep 28, 2018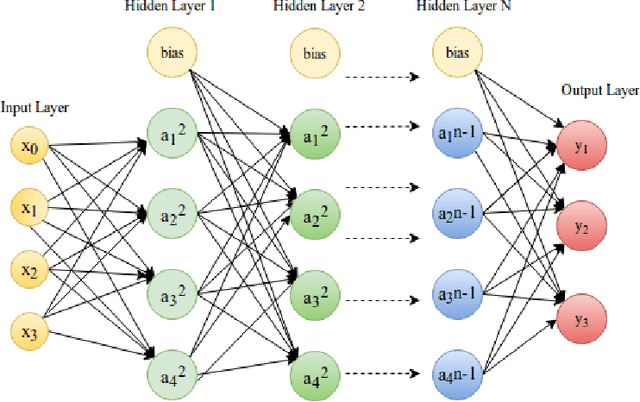
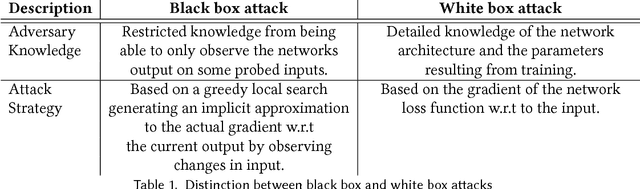


Abstract:Deep learning has emerged as a strong and efficient framework that can be applied to a broad spectrum of complex learning problems which were difficult to solve using the traditional machine learning techniques in the past. In the last few years, deep learning has advanced radically in such a way that it can surpass human-level performance on a number of tasks. As a consequence, deep learning is being extensively used in most of the recent day-to-day applications. However, security of deep learning systems are vulnerable to crafted adversarial examples, which may be imperceptible to the human eye, but can lead the model to misclassify the output. In recent times, different types of adversaries based on their threat model leverage these vulnerabilities to compromise a deep learning system where adversaries have high incentives. Hence, it is extremely important to provide robustness to deep learning algorithms against these adversaries. However, there are only a few strong countermeasures which can be used in all types of attack scenarios to design a robust deep learning system. In this paper, we attempt to provide a detailed discussion on different types of adversarial attacks with various threat models and also elaborate the efficiency and challenges of recent countermeasures against them.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge