Stephan Saalfeld
HHMI Janelia Research Campus
DeepPD: Joint Phase and Object Estimation from Phase Diversity with Neural Calibration of a Deformable Mirror
Apr 19, 2025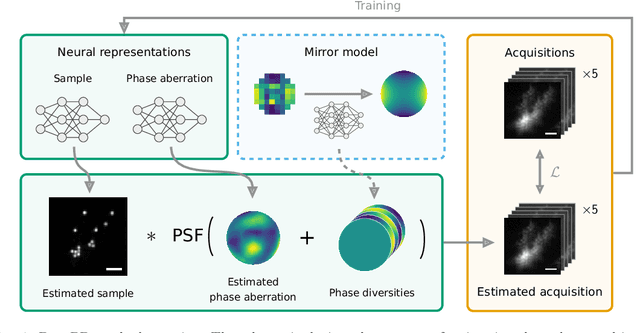


Abstract:Sample-induced aberrations and optical imperfections limit the resolution of fluorescence microscopy. Phase diversity is a powerful technique that leverages complementary phase information in sequentially acquired images with deliberately introduced aberrations--the phase diversities--to enable phase and object reconstruction and restore diffraction-limited resolution. These phase diversities are typically introduced into the optical path via a deformable mirror. Existing phase-diversity-based methods are limited to Zernike modes, require large numbers of diversity images, or depend on accurate mirror calibration--which are all suboptimal. We present DeepPD, a deep learning-based framework that combines neural representations of the object and wavefront with a learned model of the deformable mirror to jointly estimate both object and phase from only five images. DeepPD improves robustness and reconstruction quality over previous approaches, even under severe aberrations. We demonstrate its performance on calibration targets and biological samples, including immunolabeled myosin in fixed PtK2 cells.
Decomposing heterogeneous dynamical systems with graph neural networks
Jul 27, 2024Abstract:Natural physical, chemical, and biological dynamical systems are often complex, with heterogeneous components interacting in diverse ways. We show that graph neural networks can be designed to jointly learn the interaction rules and the structure of the heterogeneity from data alone. The learned latent structure and dynamics can be used to virtually decompose the complex system which is necessary to parameterize and infer the underlying governing equations. We tested the approach with simulation experiments of moving particles and vector fields that interact with each other. While our current aim is to better understand and validate the approach with simulated data, we anticipate it to become a generally applicable tool to uncover the governing rules underlying complex dynamics observed in nature.
Synaptic Cleft Segmentation in Non-Isotropic Volume Electron Microscopy of the Complete Drosophila Brain
May 07, 2018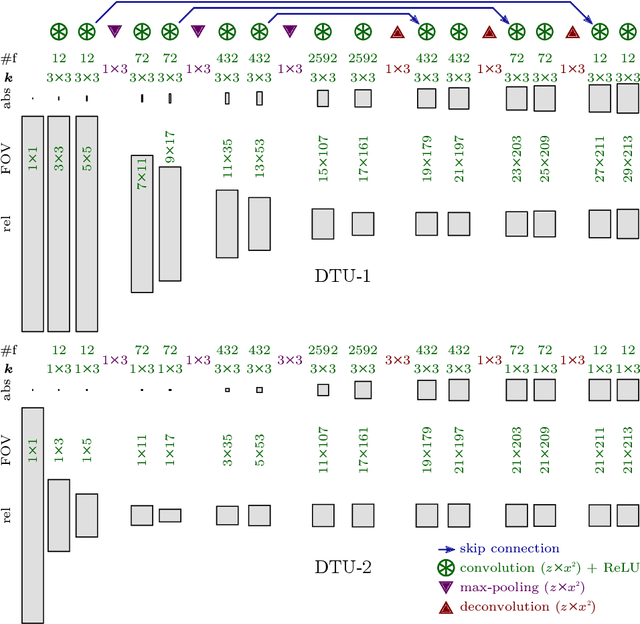
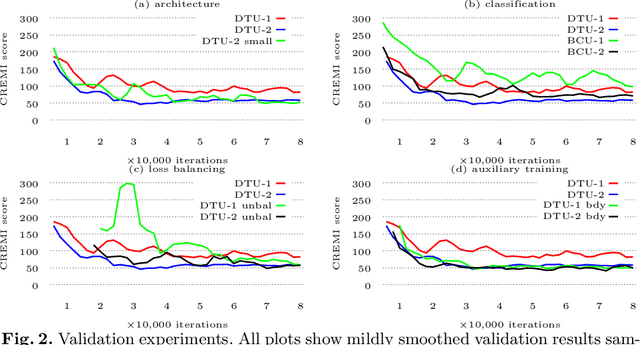
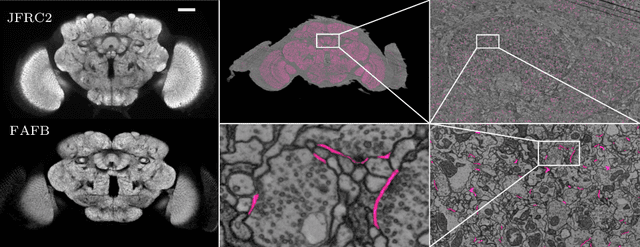
Abstract:Neural circuit reconstruction at single synapse resolution is increasingly recognized as crucially important to decipher the function of biological nervous systems. Volume electron microscopy in serial transmission or scanning mode has been demonstrated to provide the necessary resolution to segment or trace all neurites and to annotate all synaptic connections. Automatic annotation of synaptic connections has been done successfully in near isotropic electron microscopy of vertebrate model organisms. Results on non-isotropic data in insect models, however, are not yet on par with human annotation. We designed a new 3D-U-Net architecture to optimally represent isotropic fields of view in non-isotropic data. We used regression on a signed distance transform of manually annotated synaptic clefts of the CREMI challenge dataset to train this model and observed significant improvement over the state of the art. We developed open source software for optimized parallel prediction on very large volumetric datasets and applied our model to predict synaptic clefts in a 50 tera-voxels dataset of the complete Drosophila brain. Our model generalizes well to areas far away from where training data was available.
Joint Deformable Registration of Large EM Image Volumes: A Matrix Solver Approach
Apr 26, 2018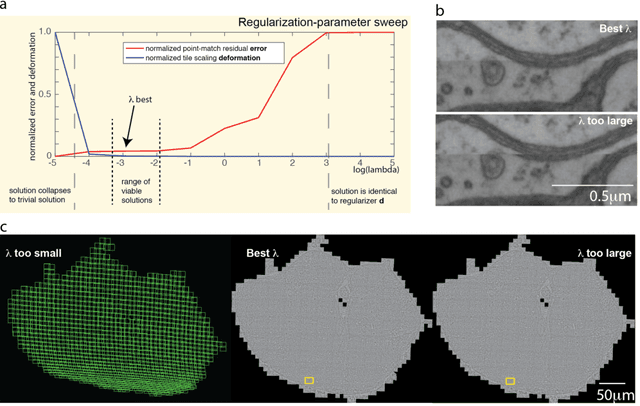
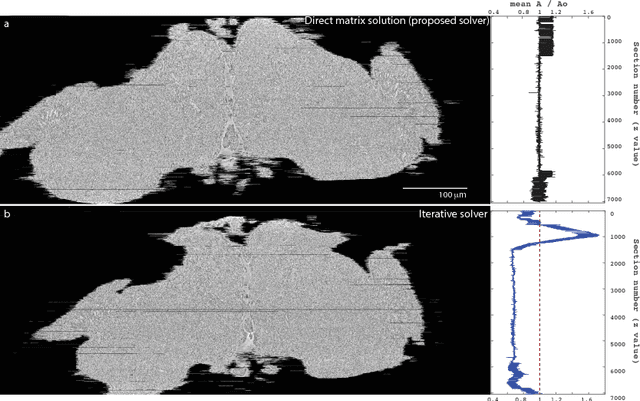
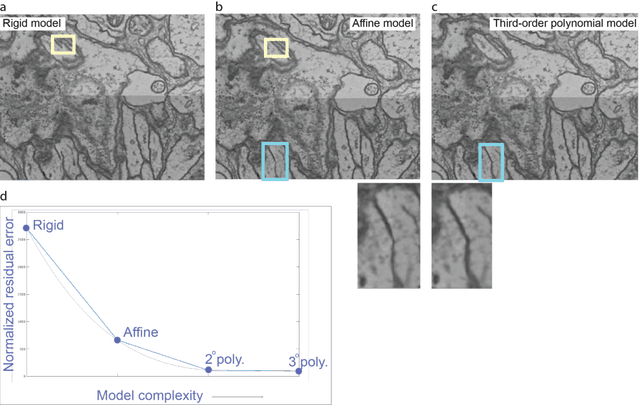
Abstract:Large electron microscopy image datasets for connectomics are typically composed of thousands to millions of partially overlapping two-dimensional images (tiles), which must be registered into a coherent volume prior to further analysis. A common registration strategy is to find matching features between neighboring and overlapping image pairs, followed by a numerical estimation of optimal image deformation using a so-called solver program. Existing solvers are inadequate for large data volumes, and inefficient for small-scale image registration. In this work, an efficient and accurate matrix-based solver method is presented. A linear system is constructed that combines minimization of feature-pair square distances with explicit constraints in a regularization term. In absence of reliable priors for regularization, we show how to construct a rigid-model approximation to use as prior. The linear system is solved using available computer programs, whose performance on typical registration tasks we briefly compare, and to which future scale-up is delegated. Our method is applied to the joint alignment of 2.67 million images, with more than 200 million point-pairs and has been used for successfully aligning the first full adult fruit fly brain.
A Deep Structured Learning Approach Towards Automating Connectome Reconstruction from 3D Electron Micrographs
Sep 24, 2017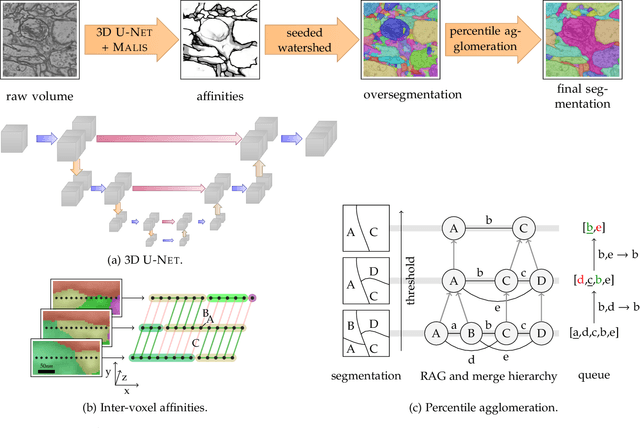

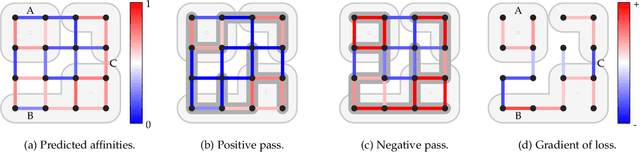
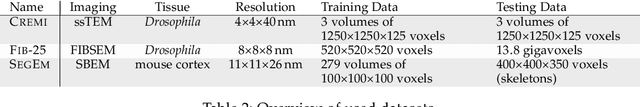
Abstract:We present a deep structured learning method for neuron segmentation from 3D electron microscopy (EM) which improves significantly upon the state of the art in terms of accuracy and scalability. Our method consists of a 3D U-Net classifier predicting affinity graphs on voxels, followed by iterative region agglomeration. We train the U-Net using a new structured loss based on MALIS that encourages topological correctness. Our extension consists of two parts: First, an $O(n\log(n))$ method to compute the loss gradient, improving over the originally proposed $O(n^2)$ algorithm. Second, we compute the gradient in two separate passes to avoid spurious contributions in early training stages. Our affinity predictions are accurate enough that simple agglomeration outperforms more involved methods used earlier on inferior predictions. We present results on three datasets (CREMI, FIB, and SegEM) of different imaging techniques and animals and achieve improvements over previous results of 27%, 15%, and 250%. Our findings suggest that a single 3D segmentation strategy can be applied to both isotropic and anisotropic EM data. The runtime of our method scales with $O(n)$ in the size of the volume and achieves a throughput of about 2.6 seconds per megavoxel, allowing processing of very large datasets.
The Candidate Multi-Cut for Cell Segmentation
Jul 04, 2017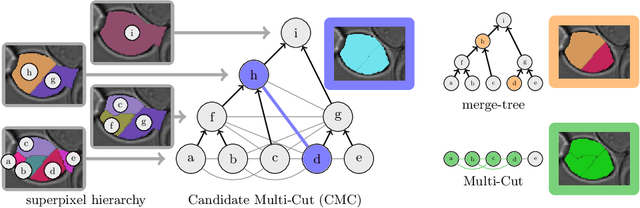
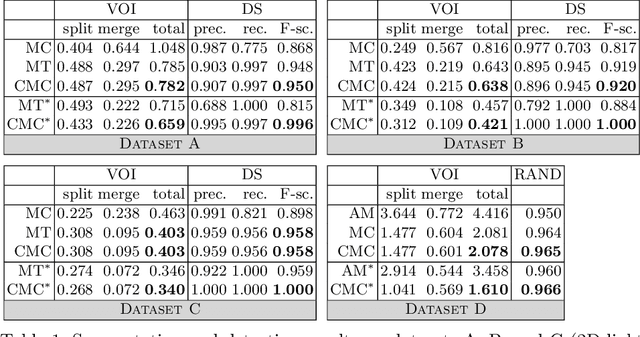

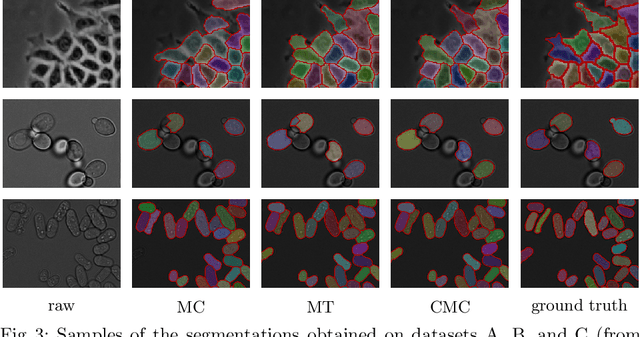
Abstract:Two successful approaches for the segmentation of biomedical images are (1) the selection of segment candidates from a merge-tree, and (2) the clustering of small superpixels by solving a Multi-Cut problem. In this paper, we introduce a model that unifies both approaches. Our model, the Candidate Multi-Cut (CMC), allows joint selection and clustering of segment candidates from a merge-tree. This way, we overcome the respective limitations of the individual methods: (1) the space of possible segmentations is not constrained to candidates of a merge-tree, and (2) the decision for clustering can be made on candidates larger than superpixels, using features over larger contexts. We solve the optimization problem of selecting and clustering of candidates using an integer linear program. On datasets of 2D light microscopy of cell populations and 3D electron microscopy of neurons, we show that our method generalizes well and generates more accurate segmentations than merge-tree or Multi-Cut methods alone.
Deep Learning for Isotropic Super-Resolution from Non-Isotropic 3D Electron Microscopy
Jun 09, 2017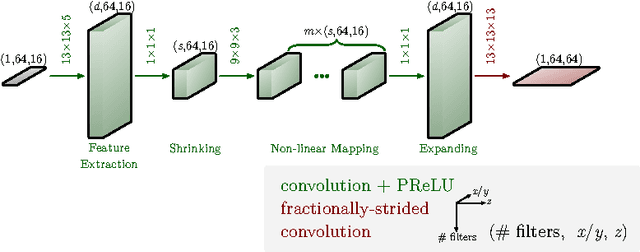
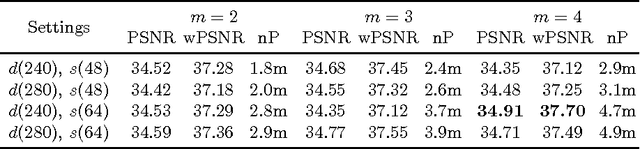
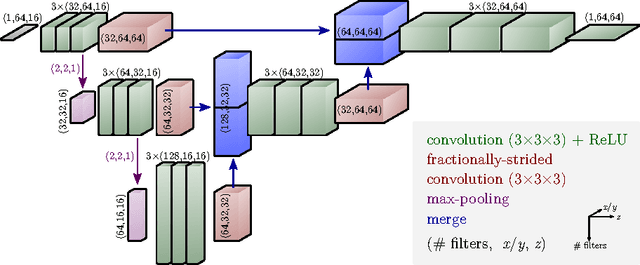
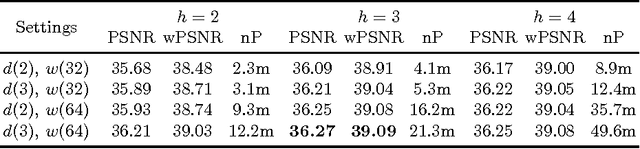
Abstract:The most sophisticated existing methods to generate 3D isotropic super-resolution (SR) from non-isotropic electron microscopy (EM) are based on learned dictionaries. Unfortunately, none of the existing methods generate practically satisfying results. For 2D natural images, recently developed super-resolution methods that use deep learning have been shown to significantly outperform the previous state of the art. We have adapted one of the most successful architectures (FSRCNN) for 3D super-resolution, and compared its performance to a 3D U-Net architecture that has not been used previously to generate super-resolution. We trained both architectures on artificially downscaled isotropic ground truth from focused ion beam milling scanning EM (FIB-SEM) and tested the performance for various hyperparameter settings. Our results indicate that both architectures can successfully generate 3D isotropic super-resolution from non-isotropic EM, with the U-Net performing consistently better. We propose several promising directions for practical application.
Image-Based Correction of Continuous and Discontinuous Non-Planar Axial Distortion in Serial Section Microscopy
Jun 17, 2016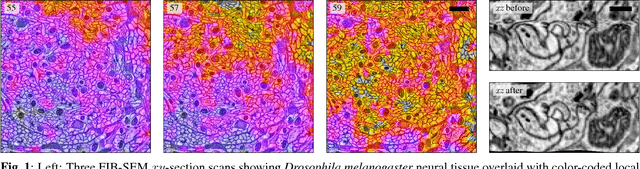
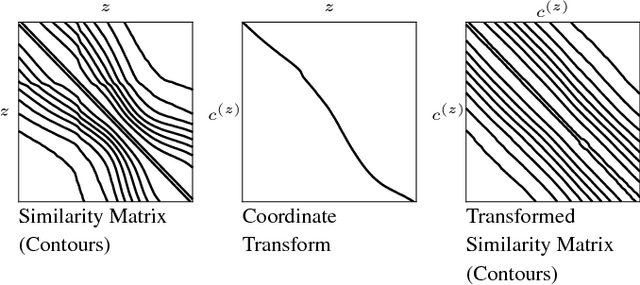
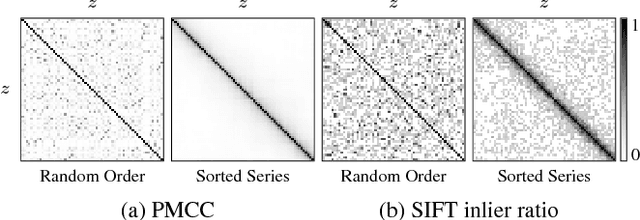
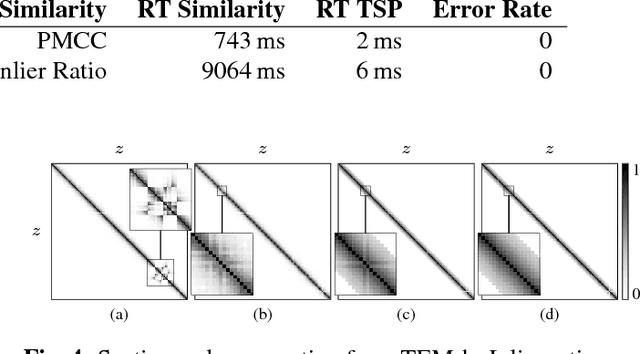
Abstract:Motivation: Serial section microscopy is an established method for detailed anatomy reconstruction of biological specimen. During the last decade, high resolution electron microscopy (EM) of serial sections has become the de-facto standard for reconstruction of neural connectivity at ever increasing scales (EM connectomics). In serial section microscopy, the axial dimension of the volume is sampled by physically removing thin sections from the embedded specimen and subsequently imaging either the block-face or the section series. This process has limited precision leading to inhomogeneous non-planar sampling of the axial dimension of the volume which, in turn, results in distorted image volumes. This includes that section series may be collected and imaged in unknown order. Results: We developed methods to identify and correct these distortions through image-based signal analysis without any additional physical apparatus or measurements. We demonstrate the efficacy of our methods in proof of principle experiments and application to real world problems. Availability and Implementation: We made our work available as libraries for the ImageJ distribution Fiji and for deployment in a high performance parallel computing environment. Our sources are open and available at http://github.com/saalfeldla/section-sort, http://github.com/saalfeldlab/em-thickness-estimation, and http://github.com/saalfeldlab/z-spacing-spark. Contact: saalfelds@janelia.hhmi.org
Robust Registration of Calcium Images by Learned Contrast Synthesis
Nov 03, 2015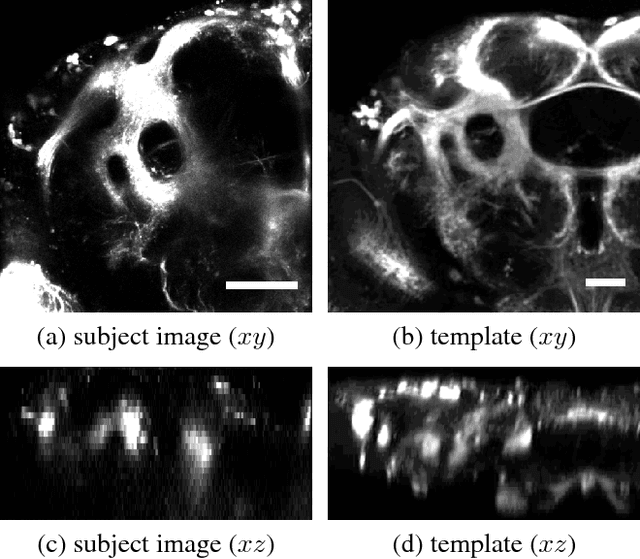
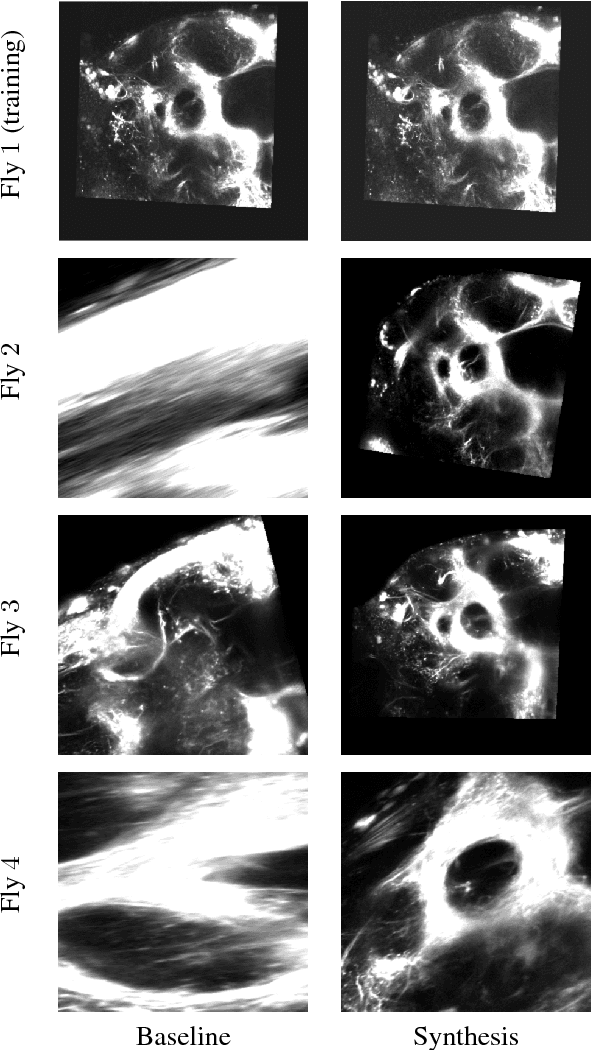
Abstract:Multi-modal image registration is a challenging task that is vital to fuse complementary signals for subsequent analyses. Despite much research into cost functions addressing this challenge, there exist cases in which these are ineffective. In this work, we show that (1) this is true for the registration of in-vivo Drosophila brain volumes visualizing genetically encoded calcium indicators to an nc82 atlas and (2) that machine learning based contrast synthesis can yield improvements. More specifically, the number of subjects for which the registration outright failed was greatly reduced (from 40% to 15%) by using a synthesized image.
Post-acquisition image based compensation for thickness variation in microscopy section series
May 27, 2015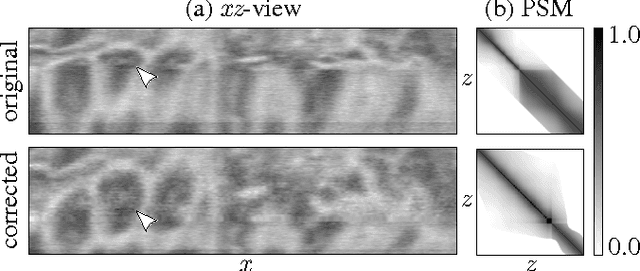
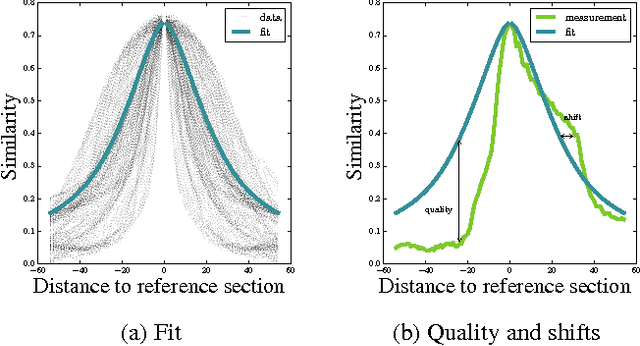

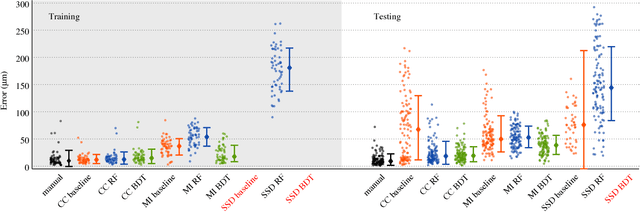
Abstract:Serial section Microscopy is an established method for volumetric anatomy reconstruction. Section series imaged with Electron Microscopy are currently vital for the reconstruction of the synaptic connectivity of entire animal brains such as that of Drosophila melanogaster. The process of removing ultrathin layers from a solid block containing the specimen, however, is a fragile procedure and has limited precision with respect to section thickness. We have developed a method to estimate the relative z-position of each individual section as a function of signal change across the section series. First experiments show promising results on both serial section Transmission Electron Microscopy (ssTEM) data and Focused Ion Beam Scanning Electron Microscopy (FIB-SEM) series. We made our solution available as Open Source plugins for the TrakEM2 software and the ImageJ distribution Fiji.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge