William Grisaitis
The Animation Transformer: Visual Correspondence via Segment Matching
Sep 08, 2021

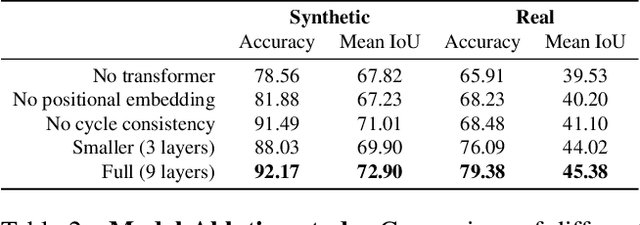
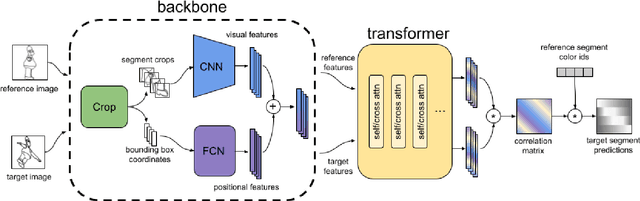
Abstract:Visual correspondence is a fundamental building block on the way to building assistive tools for hand-drawn animation. However, while a large body of work has focused on learning visual correspondences at the pixel-level, few approaches have emerged to learn correspondence at the level of line enclosures (segments) that naturally occur in hand-drawn animation. Exploiting this structure in animation has numerous benefits: it avoids the intractable memory complexity of attending to individual pixels in high resolution images and enables the use of real-world animation datasets that contain correspondence information at the level of per-segment colors. To that end, we propose the Animation Transformer (AnT) which uses a transformer-based architecture to learn the spatial and visual relationships between segments across a sequence of images. AnT enables practical ML-assisted colorization for professional animation workflows and is publicly accessible as a creative tool in Cadmium.
Anisotropic EM Segmentation by 3D Affinity Learning and Agglomeration
Aug 03, 2018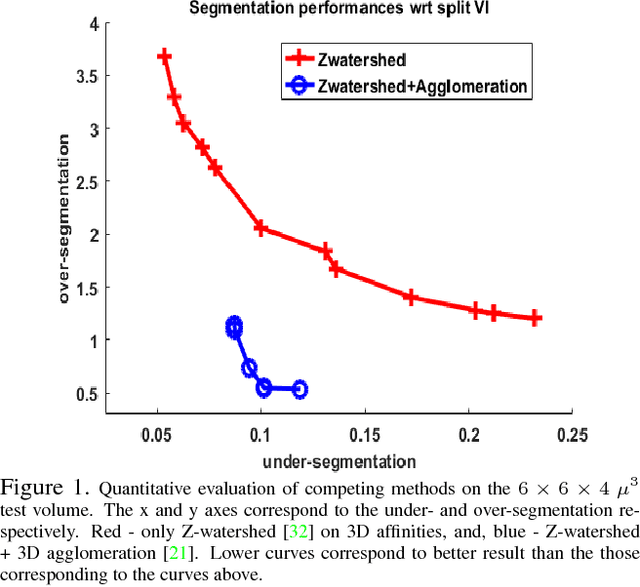
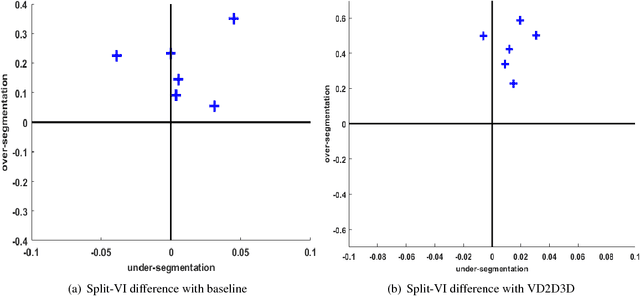
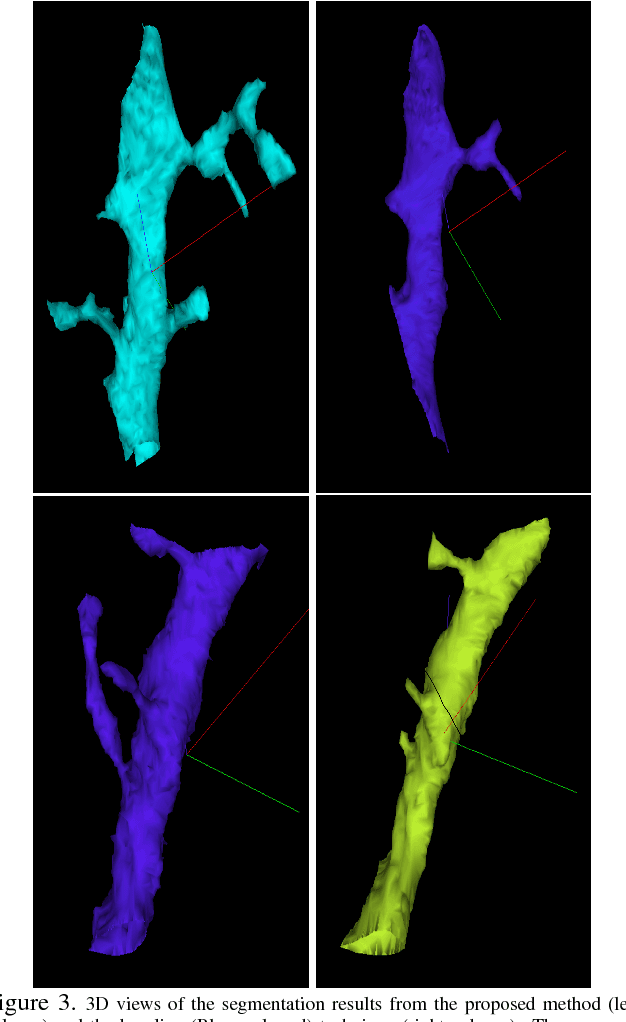
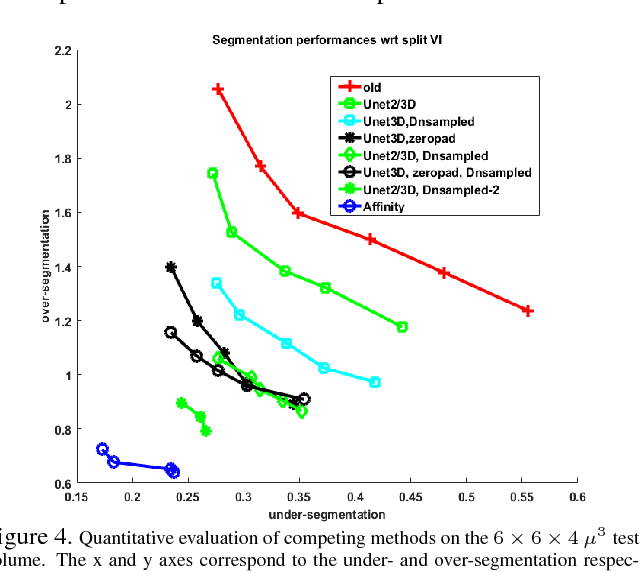
Abstract:The field of connectomics has recently produced neuron wiring diagrams from relatively large brain regions from multiple animals. Most of these neural reconstructions were computed from isotropic (e.g., FIBSEM) or near isotropic (e.g., SBEM) data. In spite of the remarkable progress on algorithms in recent years, automatic dense reconstruction from anisotropic data remains a challenge for the connectomics community. One significant hurdle in the segmentation of anisotropic data is the difficulty in generating a suitable initial over-segmentation. In this study, we present a segmentation method for anisotropic EM data that agglomerates a 3D over-segmentation computed from the 3D affinity prediction. A 3D U-net is trained to predict 3D affinities by the MALIS approach. Experiments on multiple datasets demonstrates the strength and robustness of the proposed method for anisotropic EM segmentation.
A Deep Structured Learning Approach Towards Automating Connectome Reconstruction from 3D Electron Micrographs
Sep 24, 2017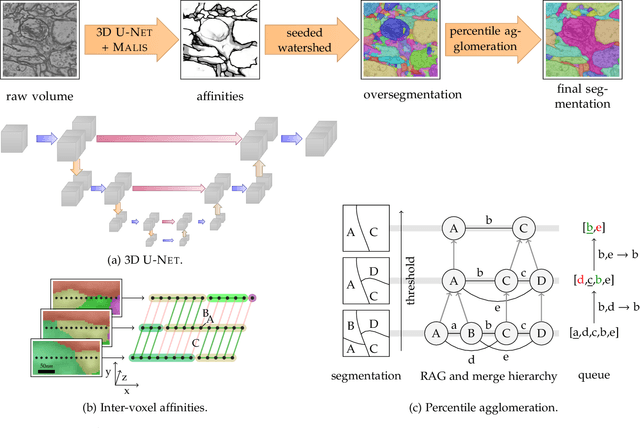

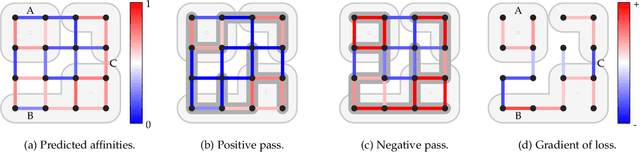
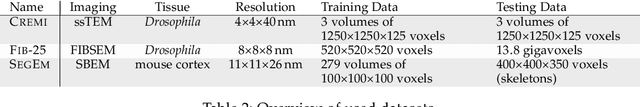
Abstract:We present a deep structured learning method for neuron segmentation from 3D electron microscopy (EM) which improves significantly upon the state of the art in terms of accuracy and scalability. Our method consists of a 3D U-Net classifier predicting affinity graphs on voxels, followed by iterative region agglomeration. We train the U-Net using a new structured loss based on MALIS that encourages topological correctness. Our extension consists of two parts: First, an $O(n\log(n))$ method to compute the loss gradient, improving over the originally proposed $O(n^2)$ algorithm. Second, we compute the gradient in two separate passes to avoid spurious contributions in early training stages. Our affinity predictions are accurate enough that simple agglomeration outperforms more involved methods used earlier on inferior predictions. We present results on three datasets (CREMI, FIB, and SegEM) of different imaging techniques and animals and achieve improvements over previous results of 27%, 15%, and 250%. Our findings suggest that a single 3D segmentation strategy can be applied to both isotropic and anisotropic EM data. The runtime of our method scales with $O(n)$ in the size of the volume and achieves a throughput of about 2.6 seconds per megavoxel, allowing processing of very large datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge