Larissa Heinrich
Graph neural networks uncover structure and functions underlying the activity of simulated neural assemblies
Feb 11, 2026Abstract:Graph neural networks trained to predict observable dynamics can be used to decompose the temporal activity of complex heterogeneous systems into simple, interpretable representations. Here we apply this framework to simulated neural assemblies with thousands of neurons and demonstrate that it can jointly reveal the connectivity matrix, the neuron types, the signaling functions, and in some cases hidden external stimuli. In contrast to existing machine learning approaches such as recurrent neural networks and transformers, which emphasize predictive accuracy but offer limited interpretability, our method provides both reliable forecasts of neural activity and interpretable decomposition of the mechanisms governing large neural assemblies.
DeepPD: Joint Phase and Object Estimation from Phase Diversity with Neural Calibration of a Deformable Mirror
Apr 19, 2025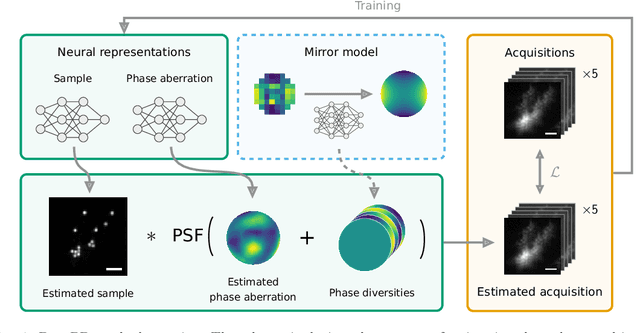
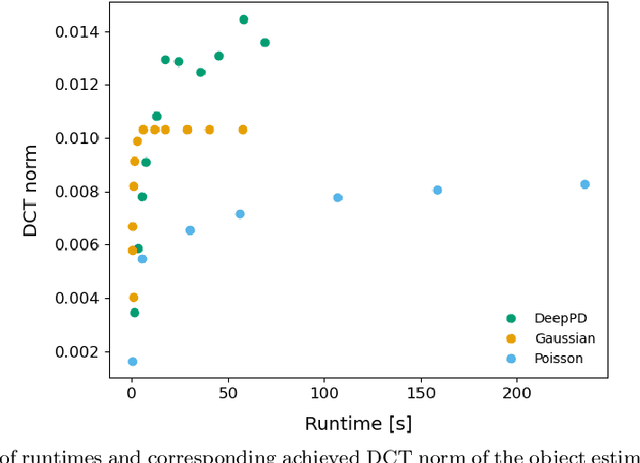


Abstract:Sample-induced aberrations and optical imperfections limit the resolution of fluorescence microscopy. Phase diversity is a powerful technique that leverages complementary phase information in sequentially acquired images with deliberately introduced aberrations--the phase diversities--to enable phase and object reconstruction and restore diffraction-limited resolution. These phase diversities are typically introduced into the optical path via a deformable mirror. Existing phase-diversity-based methods are limited to Zernike modes, require large numbers of diversity images, or depend on accurate mirror calibration--which are all suboptimal. We present DeepPD, a deep learning-based framework that combines neural representations of the object and wavefront with a learned model of the deformable mirror to jointly estimate both object and phase from only five images. DeepPD improves robustness and reconstruction quality over previous approaches, even under severe aberrations. We demonstrate its performance on calibration targets and biological samples, including immunolabeled myosin in fixed PtK2 cells.
DaCapo: a modular deep learning framework for scalable 3D image segmentation
Aug 05, 2024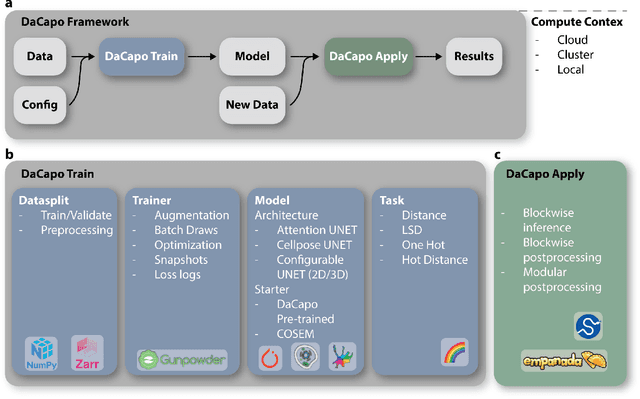
Abstract:DaCapo is a specialized deep learning library tailored to expedite the training and application of existing machine learning approaches on large, near-isotropic image data. In this correspondence, we introduce DaCapo's unique features optimized for this specific domain, highlighting its modular structure, efficient experiment management tools, and scalable deployment capabilities. We discuss its potential to improve access to large-scale, isotropic image segmentation and invite the community to explore and contribute to this open-source initiative.
Decomposing heterogeneous dynamical systems with graph neural networks
Jul 27, 2024



Abstract:Natural physical, chemical, and biological dynamical systems are often complex, with heterogeneous components interacting in diverse ways. We show that graph neural networks can be designed to jointly learn the interaction rules and the structure of the heterogeneity from data alone. The learned latent structure and dynamics can be used to virtually decompose the complex system which is necessary to parameterize and infer the underlying governing equations. We tested the approach with simulation experiments of moving particles and vector fields that interact with each other. While our current aim is to better understand and validate the approach with simulated data, we anticipate it to become a generally applicable tool to uncover the governing rules underlying complex dynamics observed in nature.
Synaptic Cleft Segmentation in Non-Isotropic Volume Electron Microscopy of the Complete Drosophila Brain
May 07, 2018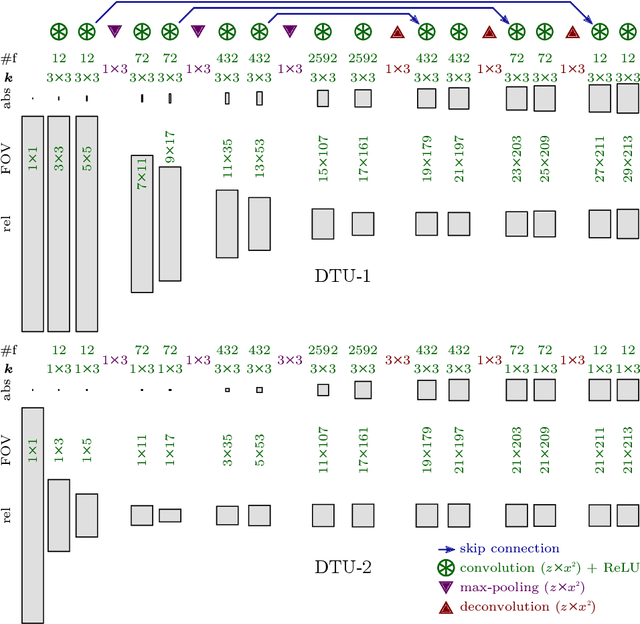
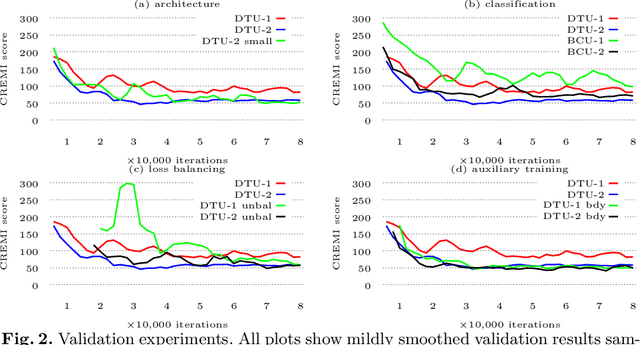
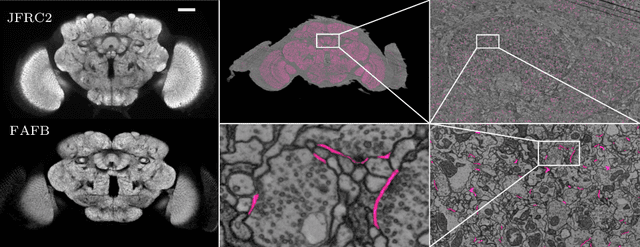
Abstract:Neural circuit reconstruction at single synapse resolution is increasingly recognized as crucially important to decipher the function of biological nervous systems. Volume electron microscopy in serial transmission or scanning mode has been demonstrated to provide the necessary resolution to segment or trace all neurites and to annotate all synaptic connections. Automatic annotation of synaptic connections has been done successfully in near isotropic electron microscopy of vertebrate model organisms. Results on non-isotropic data in insect models, however, are not yet on par with human annotation. We designed a new 3D-U-Net architecture to optimally represent isotropic fields of view in non-isotropic data. We used regression on a signed distance transform of manually annotated synaptic clefts of the CREMI challenge dataset to train this model and observed significant improvement over the state of the art. We developed open source software for optimized parallel prediction on very large volumetric datasets and applied our model to predict synaptic clefts in a 50 tera-voxels dataset of the complete Drosophila brain. Our model generalizes well to areas far away from where training data was available.
Deep Learning for Isotropic Super-Resolution from Non-Isotropic 3D Electron Microscopy
Jun 09, 2017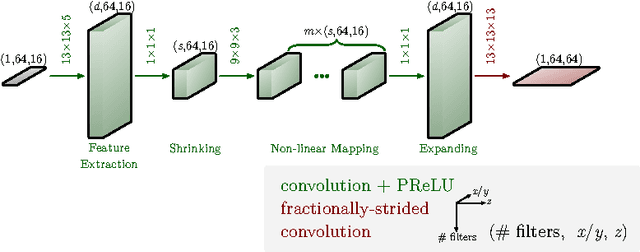
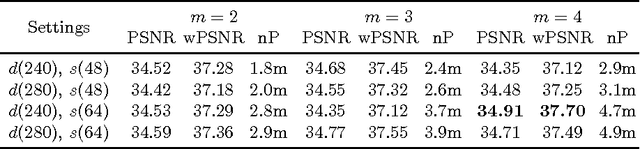
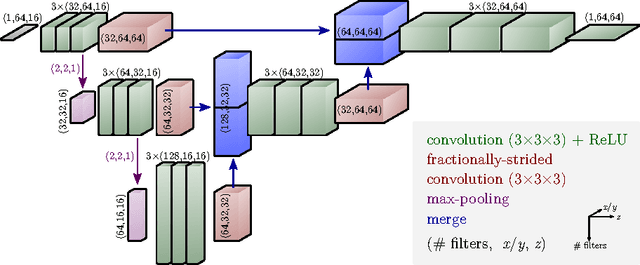
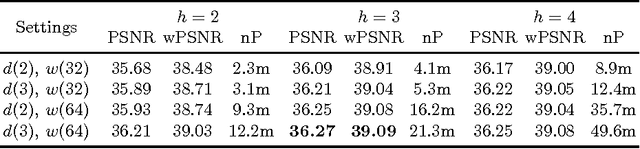
Abstract:The most sophisticated existing methods to generate 3D isotropic super-resolution (SR) from non-isotropic electron microscopy (EM) are based on learned dictionaries. Unfortunately, none of the existing methods generate practically satisfying results. For 2D natural images, recently developed super-resolution methods that use deep learning have been shown to significantly outperform the previous state of the art. We have adapted one of the most successful architectures (FSRCNN) for 3D super-resolution, and compared its performance to a 3D U-Net architecture that has not been used previously to generate super-resolution. We trained both architectures on artificially downscaled isotropic ground truth from focused ion beam milling scanning EM (FIB-SEM) and tested the performance for various hyperparameter settings. Our results indicate that both architectures can successfully generate 3D isotropic super-resolution from non-isotropic EM, with the U-Net performing consistently better. We propose several promising directions for practical application.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge