Simon K. Warfield
FetDTIAlign: A Deep Learning Framework for Affine and Deformable Registration of Fetal Brain dMRI
Feb 03, 2025Abstract:Diffusion MRI (dMRI) provides unique insights into fetal brain microstructure in utero. Longitudinal and cross-sectional fetal dMRI studies can reveal crucial neurodevelopmental changes but require precise spatial alignment across scans and subjects. This is challenging due to low data quality, rapid brain development, and limited anatomical landmarks. Existing registration methods, designed for high-quality adult data, struggle with these complexities. To address this, we introduce FetDTIAlign, a deep learning approach for fetal brain dMRI registration, enabling accurate affine and deformable alignment. FetDTIAlign features a dual-encoder architecture and iterative feature-based inference, reducing the impact of noise and low resolution. It optimizes network configurations and domain-specific features at each registration stage, enhancing both robustness and accuracy. We validated FetDTIAlign on data from 23 to 36 weeks gestation, covering 60 white matter tracts. It consistently outperformed two classical optimization-based methods and a deep learning pipeline, achieving superior anatomical correspondence. Further validation on external data from the Developing Human Connectome Project confirmed its generalizability across acquisition protocols. Our results demonstrate the feasibility of deep learning for fetal brain dMRI registration, providing a more accurate and reliable alternative to classical techniques. By enabling precise cross-subject and tract-specific analyses, FetDTIAlign supports new discoveries in early brain development.
Streamline tractography of the fetal brain in utero with machine learning
Aug 26, 2024



Abstract:Diffusion-weighted magnetic resonance imaging (dMRI) is the only non-invasive tool for studying white matter tracts and structural connectivity of the brain. These assessments rely heavily on tractography techniques, which reconstruct virtual streamlines representing white matter fibers. Much effort has been devoted to improving tractography methodology for adult brains, while tractography of the fetal brain has been largely neglected. Fetal tractography faces unique difficulties due to low dMRI signal quality, immature and rapidly developing brain structures, and paucity of reference data. This work presents the first machine learning model for fetal tractography. The model input consists of five sources of information: (1) Fiber orientation, inferred from a diffusion tensor fit to the dMRI signal; (2) Directions of recent propagation steps; (3) Global spatial information, encoded as distances to keypoints in the brain cortex; (4) Tissue segmentation information; and (5) Prior information about the expected local fiber orientations supplied with an atlas. In order to mitigate the local tensor estimation error, a large spatial context around the current point in the diffusion tensor image is encoded using convolutional and attention neural network modules. Moreover, the diffusion tensor information at a hypothetical next point is included in the model input. Filtering rules based on anatomically constrained tractography are applied to prune implausible streamlines. We trained the model on manually-refined whole-brain fetal tractograms and validated the trained model on an independent set of 11 test scans with gestational ages between 23 and 36 weeks. Results show that our proposed method achieves superior performance across all evaluated tracts. The new method can significantly advance the capabilities of dMRI for studying normal and abnormal brain development in utero.
A self-attention model for robust rigid slice-to-volume registration of functional MRI
Apr 06, 2024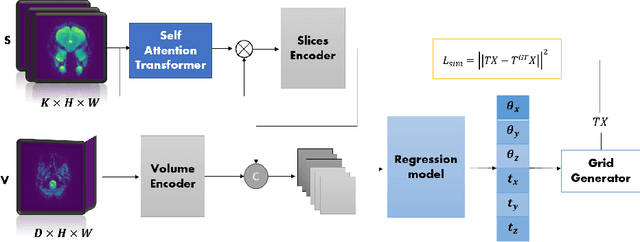
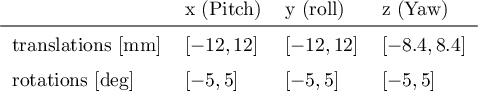
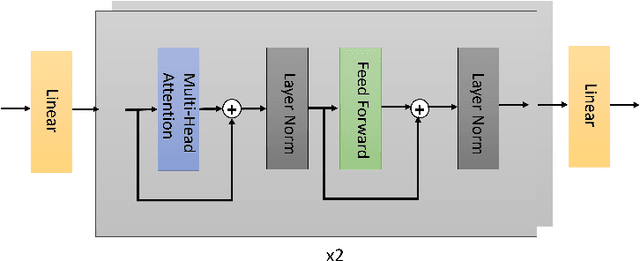
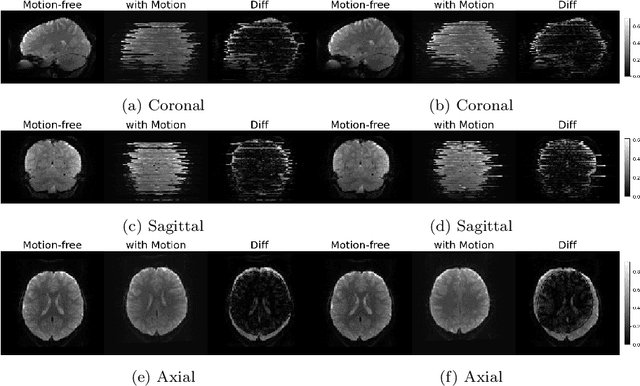
Abstract:Functional Magnetic Resonance Imaging (fMRI) is vital in neuroscience, enabling investigations into brain disorders, treatment monitoring, and brain function mapping. However, head motion during fMRI scans, occurring between shots of slice acquisition, can result in distortion, biased analyses, and increased costs due to the need for scan repetitions. Therefore, retrospective slice-level motion correction through slice-to-volume registration (SVR) is crucial. Previous studies have utilized deep learning (DL) based models to address the SVR task; however, they overlooked the uncertainty stemming from the input stack of slices and did not assign weighting or scoring to each slice. In this work, we introduce an end-to-end SVR model for aligning 2D fMRI slices with a 3D reference volume, incorporating a self-attention mechanism to enhance robustness against input data variations and uncertainties. It utilizes independent slice and volume encoders and a self-attention module to assign pixel-wise scores for each slice. We conducted evaluation experiments on 200 images involving synthetic rigid motion generated from 27 subjects belonging to the test set, from the publicly available Healthy Brain Network (HBN) dataset. Our experimental results demonstrate that our model achieves competitive performance in terms of alignment accuracy compared to state-of-the-art deep learning-based methods (Euclidean distance of $0.93$ [mm] vs. $1.86$ [mm]). Furthermore, our approach exhibits significantly faster registration speed compared to conventional iterative methods ($0.096$ sec. vs. $1.17$ sec.). Our end-to-end SVR model facilitates real-time head motion tracking during fMRI acquisition, ensuring reliability and robustness against uncertainties in inputs. source code, which includes the training and evaluations, will be available soon.
Anatomically Constrained Tractography of the Fetal Brain
Mar 04, 2024Abstract:Diffusion-weighted Magnetic Resonance Imaging (dMRI) is increasingly used to study the fetal brain in utero. An important computation enabled by dMRI is streamline tractography, which has unique applications such as tract-specific analysis of the brain white matter and structural connectivity assessment. However, due to the low fetal dMRI data quality and the challenging nature of tractography, existing methods tend to produce highly inaccurate results. They generate many false streamlines while failing to reconstruct streamlines that constitute the major white matter tracts. In this paper, we advocate for anatomically constrained tractography based on an accurate segmentation of the fetal brain tissue directly in the dMRI space. We develop a deep learning method to compute the segmentation automatically. Experiments on independent test data show that this method can accurately segment the fetal brain tissue and drastically improve tractography results. It enables the reconstruction of highly curved tracts such as optic radiations. Importantly, our method infers the tissue segmentation and streamline propagation direction from a diffusion tensor fit to the dMRI data, making it applicable to routine fetal dMRI scans. The proposed method can lead to significant improvements in the accuracy and reproducibility of quantitative assessment of the fetal brain with dMRI.
IVIM-Morph: Motion-compensated quantitative Intra-voxel Incoherent Motion (IVIM) analysis for functional fetal lung maturity assessment from diffusion-weighted MRI data
Jan 17, 2024
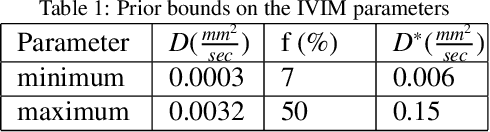

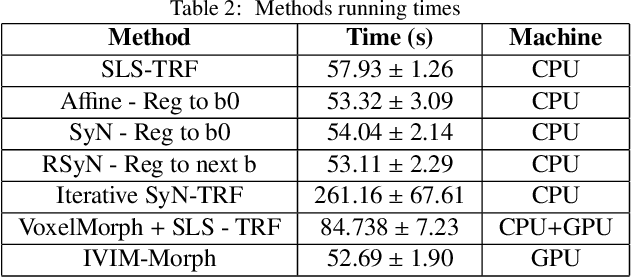
Abstract:Quantitative analysis of pseudo-diffusion in diffusion-weighted magnetic resonance imaging (DWI) data shows potential for assessing fetal lung maturation and generating valuable imaging biomarkers. Yet, the clinical utility of DWI data is hindered by unavoidable fetal motion during acquisition. We present IVIM-morph, a self-supervised deep neural network model for motion-corrected quantitative analysis of DWI data using the Intra-voxel Incoherent Motion (IVIM) model. IVIM-morph combines two sub-networks, a registration sub-network, and an IVIM model fitting sub-network, enabling simultaneous estimation of IVIM model parameters and motion. To promote physically plausible image registration, we introduce a biophysically informed loss function that effectively balances registration and model-fitting quality. We validated the efficacy of IVIM-morph by establishing a correlation between the predicted IVIM model parameters of the lung and gestational age (GA) using fetal DWI data of 39 subjects. IVIM-morph exhibited a notably improved correlation with gestational age (GA) when performing in-vivo quantitative analysis of fetal lung DWI data during the canalicular phase. IVIM-morph shows potential in developing valuable biomarkers for non-invasive assessment of fetal lung maturity with DWI data. Moreover, its adaptability opens the door to potential applications in other clinical contexts where motion compensation is essential for quantitative DWI analysis. The IVIM-morph code is readily available at: https://github.com/TechnionComputationalMRILab/qDWI-Morph.
3D Brainformer: 3D Fusion Transformer for Brain Tumor Segmentation
Apr 28, 2023



Abstract:Magnetic resonance imaging (MRI) is critically important for brain mapping in both scientific research and clinical studies. Precise segmentation of brain tumors facilitates clinical diagnosis, evaluations, and surgical planning. Deep learning has recently emerged to improve brain tumor segmentation and achieved impressive results. Convolutional architectures are widely used to implement those neural networks. By the nature of limited receptive fields, however, those architectures are subject to representing long-range spatial dependencies of the voxel intensities in MRI images. Transformers have been leveraged recently to address the above limitations of convolutional networks. Unfortunately, the majority of current Transformers-based methods in segmentation are performed with 2D MRI slices, instead of 3D volumes. Moreover, it is difficult to incorporate the structures between layers because each head is calculated independently in the Multi-Head Self-Attention mechanism (MHSA). In this work, we proposed a 3D Transformer-based segmentation approach. We developed a Fusion-Head Self-Attention mechanism (FHSA) to combine each attention head through attention logic and weight mapping, for the exploration of the long-range spatial dependencies in 3D MRI images. We implemented a plug-and-play self-attention module, named the Infinite Deformable Fusion Transformer Module (IDFTM), to extract features on any deformable feature maps. We applied our approach to the task of brain tumor segmentation, and assessed it on the public BRATS datasets. The experimental results demonstrated that our proposed approach achieved superior performance, in comparison to several state-of-the-art segmentation methods.
CORPS: Cost-free Rigorous Pseudo-labeling based on Similarity-ranking for Brain MRI Segmentation
May 19, 2022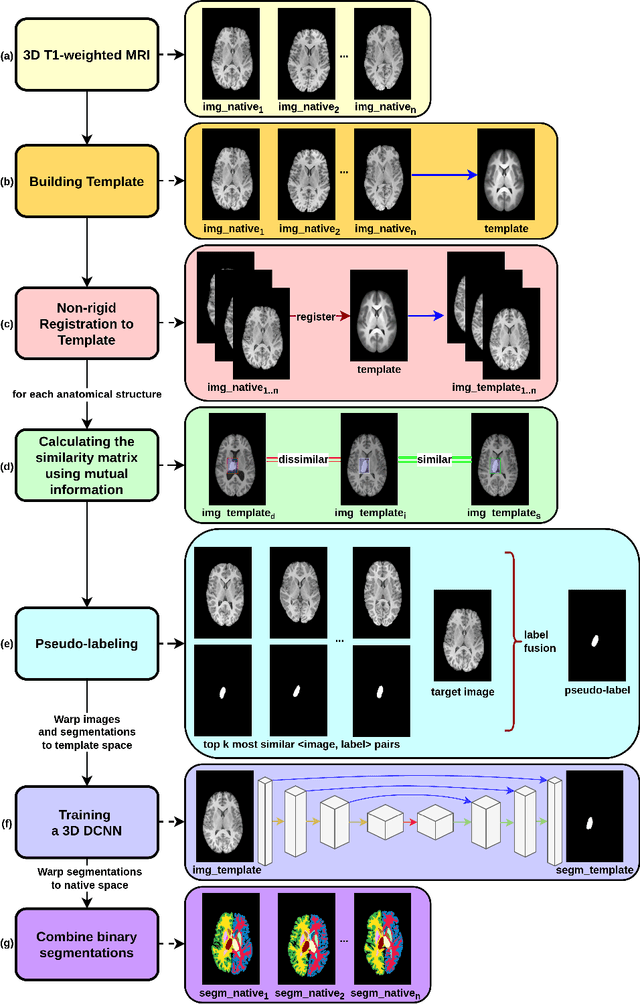
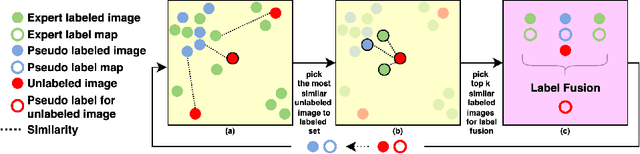
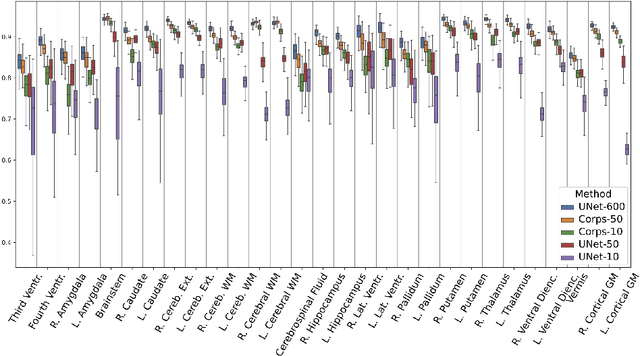
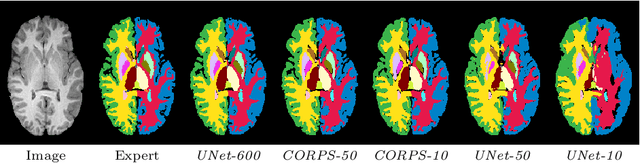
Abstract:Segmentation of brain magnetic resonance images (MRI) is crucial for the analysis of the human brain and diagnosis of various brain disorders. The drawbacks of time-consuming and error-prone manual delineation procedures are aimed to be alleviated by atlas-based and supervised machine learning methods where the former methods are computationally intense and the latter methods lack a sufficiently large number of labeled data. With this motivation, we propose CORPS, a semi-supervised segmentation framework built upon a novel atlas-based pseudo-labeling method and a 3D deep convolutional neural network (DCNN) for 3D brain MRI segmentation. In this work, we propose to generate expert-level pseudo-labels for unlabeled set of images in an order based on a local intensity-based similarity score to existing labeled set of images and using a novel atlas-based label fusion method. Then, we propose to train a 3D DCNN on the combination of expert and pseudo labeled images for binary segmentation of each anatomical structure. The binary segmentation approach is proposed to avoid the poor performance of multi-class segmentation methods on limited and imbalanced data. This also allows to employ a lightweight and efficient 3D DCNN in terms of the number of filters and reserve memory resources for training the binary networks on full-scale and full-resolution 3D MRI volumes instead of 2D/3D patches or 2D slices. Thus, the proposed framework can encapsulate the spatial contiguity in each dimension and enhance context-awareness. The experimental results demonstrate the superiority of the proposed framework over the baseline method both qualitatively and quantitatively without additional labeling cost for manual labeling.
Calibrated Diffusion Tensor Estimation
Nov 21, 2021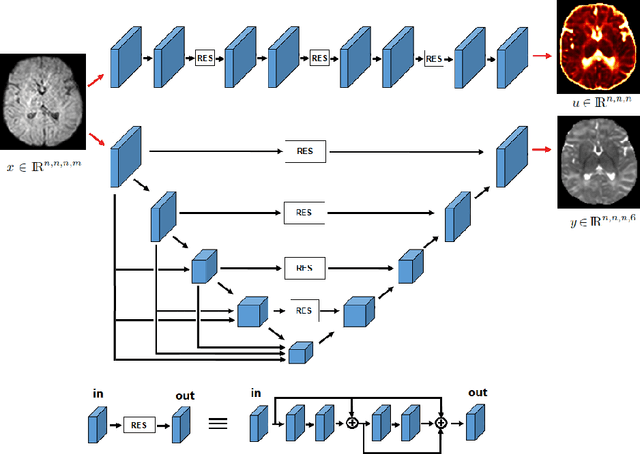
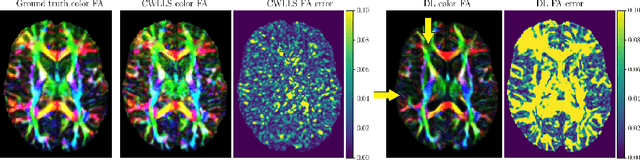
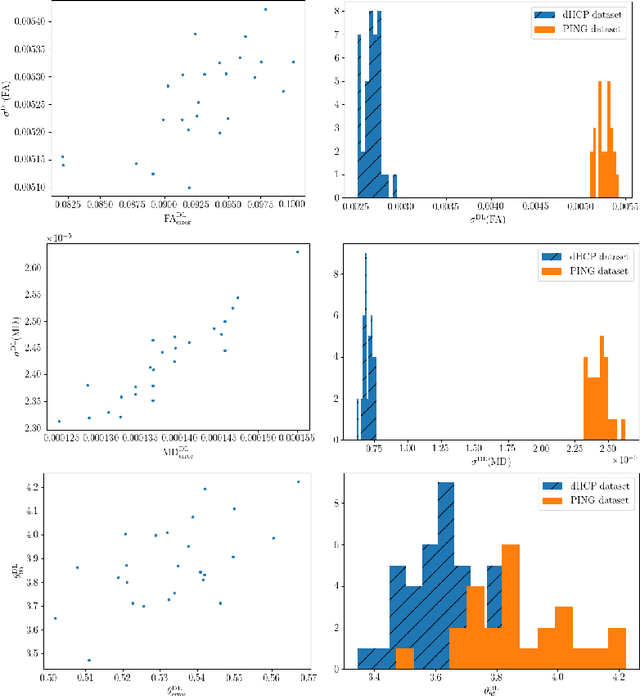
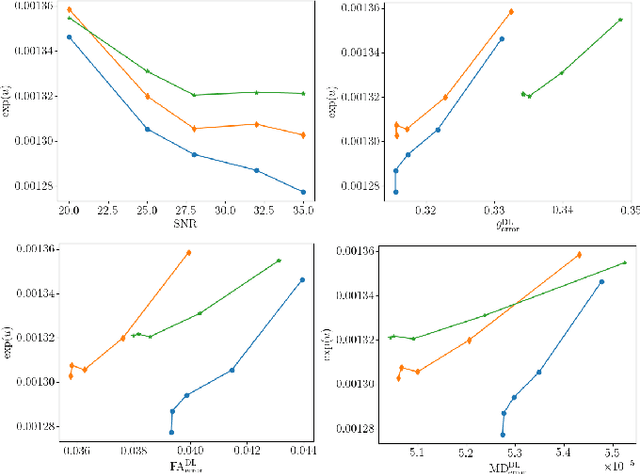
Abstract:It is highly desirable to know how uncertain a model's predictions are, especially for models that are complex and hard to understand as in deep learning. Although there has been a growing interest in using deep learning methods in diffusion-weighted MRI, prior works have not addressed the issue of model uncertainty. Here, we propose a deep learning method to estimate the diffusion tensor and compute the estimation uncertainty. Data-dependent uncertainty is computed directly by the network and learned via loss attenuation. Model uncertainty is computed using Monte Carlo dropout. We also propose a new method for evaluating the quality of predicted uncertainties. We compare the new method with the standard least-squares tensor estimation and bootstrap-based uncertainty computation techniques. Our experiments show that when the number of measurements is small the deep learning method is more accurate and its uncertainty predictions are better calibrated than the standard methods. We show that the estimation uncertainties computed by the new method can highlight the model's biases, detect domain shift, and reflect the strength of noise in the measurements. Our study shows the importance and practical value of modeling prediction uncertainties in deep learning-based diffusion MRI analysis.
A machine learning-based method for estimating the number and orientations of major fascicles in diffusion-weighted magnetic resonance imaging
Jun 19, 2020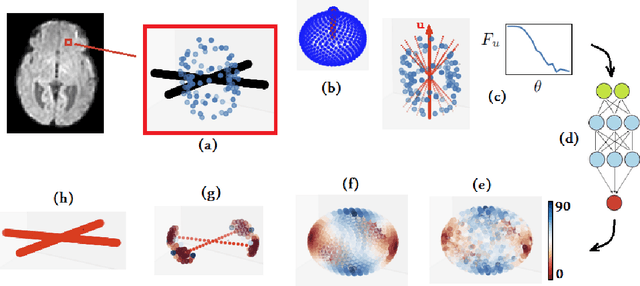
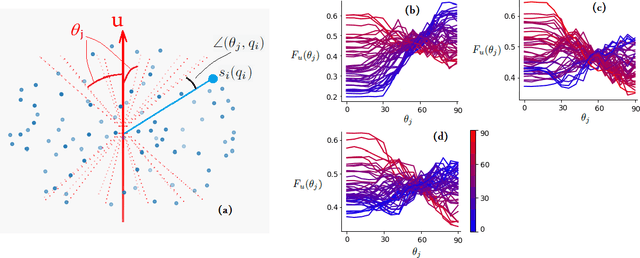

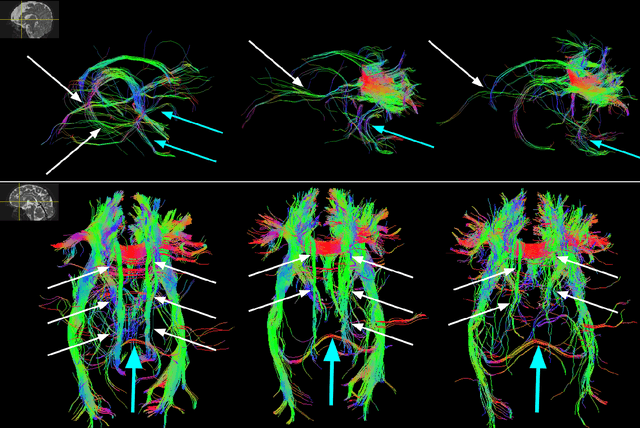
Abstract:Multi-compartment modeling of diffusion-weighted magnetic resonance imaging measurements is necessary for accurate brain connectivity analysis. Existing methods for estimating the number and orientations of fascicles in an imaging voxel either depend on non-convex optimization techniques that are sensitive to initialization and measurement noise, or are prone to predicting spurious fascicles. In this paper, we propose a machine learning-based technique that can accurately estimate the number and orientations of fascicles in a voxel. Our method can be trained with either simulated or real diffusion-weighted imaging data. Our method estimates the angle to the closest fascicle for each direction in a set of discrete directions uniformly spread on the unit sphere. This information is then processed to extract the number and orientations of fascicles in a voxel. On realistic simulated phantom data with known ground truth, our method predicts the number and orientations of crossing fascicles more accurately than several existing methods. It also leads to more accurate tractography. On real data, our method is better than or compares favorably with standard methods in terms of robustness to measurement down-sampling and also in terms of expert quality assessment of tractography results.
Critical Assessment of Transfer Learning for Medical Image Segmentation with Fully Convolutional Neural Networks
May 30, 2020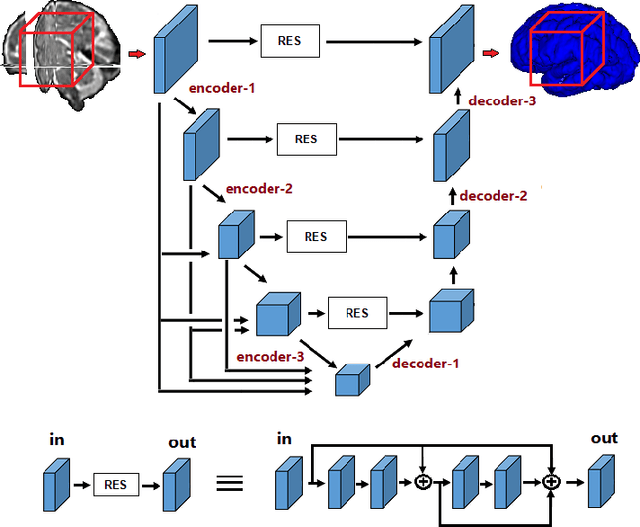
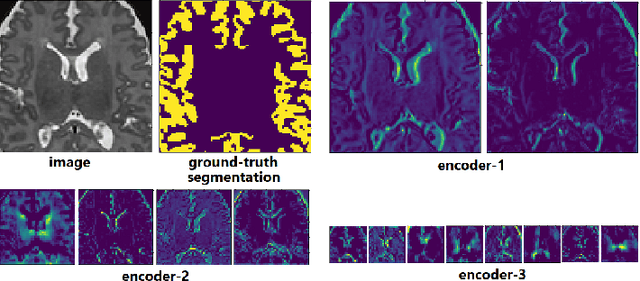
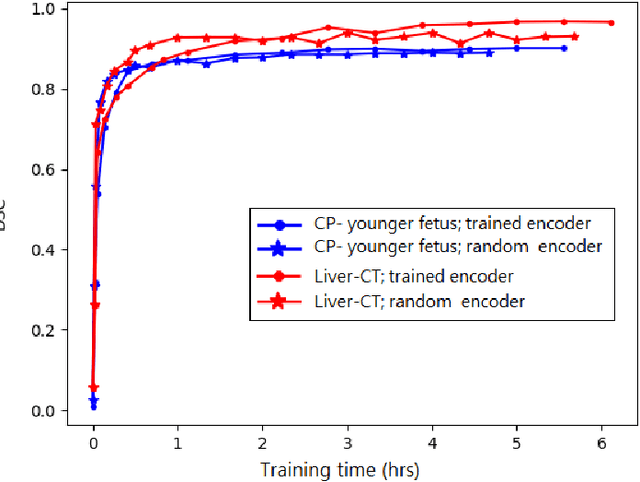
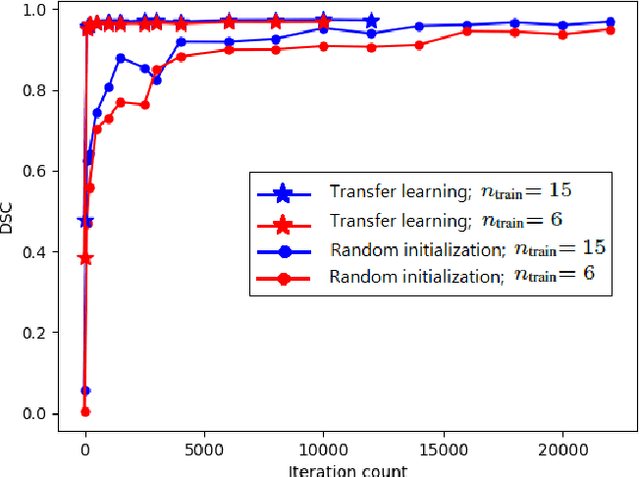
Abstract:Transfer learning is widely used for training machine learning models. Here, we study the role of transfer learning for training fully convolutional networks (FCNs) for medical image segmentation. Our experiments show that although transfer learning reduces the training time on the target task, the improvement in segmentation accuracy is highly task/data-dependent. Larger improvements in accuracy are observed when the segmentation task is more challenging and the target training data is smaller. We observe that convolutional filters of an FCN change little during training for medical image segmentation, and still look random at convergence. We further show that quite accurate FCNs can be built by freezing the encoder section of the network at random values and only training the decoder section. At least for medical image segmentation, this finding challenges the common belief that the encoder section needs to learn data/task-specific representations. We examine the evolution of FCN representations to gain a better insight into the effects of transfer learning on the training dynamics. Our analysis shows that although FCNs trained via transfer learning learn different representations than FCNs trained with random initialization, the variability among FCNs trained via transfer learning can be as high as that among FCNs trained with random initialization. Moreover, feature reuse is not restricted to the early encoder layers; rather, it can be more significant in deeper layers. These findings offer new insights and suggest alternative ways of training FCNs for medical image segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge