Sepideh Hatamikia
Multimodal Connectome Fusion via Cross-Attention for Autism Spectrum Disorder Classification Using Graph Learning
Mar 16, 2026Abstract:Autism spectrum disorder (ASD) is a complex neurodevelopmental condition characterized by atypical functional brain connectivity and subtle structural alterations. rs-fMRI has been widely used to identify disruptions in large-scale brain networks, while structural MRI provides complementary information about morphological organization. Despite their complementary nature, effectively integrating these heterogeneous imaging modalities within a unified framework remains challenging. This study proposes a multimodal graph learning framework that preserves the dominant role of functional connectivity while integrating structural imaging and phenotypic information for ASD classification. The proposed framework is evaluated on ABIDE-I dataset. Each subject is represented as a node within a population graph. Functional and structural features are extracted as modality-specific node attributes, while inter-subject relationships are modeled using a pairwise association encoder (PAE) based on phenotypic information. Two Edge Variational GCNs are trained to learn subject-level embeddings. To enable effective multimodal integration, we introduce a novel asymmetric transformer-based cross-attention mechanism that allows functional embeddings to selectively incorporate complementary structural information while preserving functional dominance. The fused embeddings are then passed to a MLP for ASD classification. Using stratified 10-fold cross-validation, the framework achieved an AUC of 87.3% and an accuracy of 84.4%. Under leave-one-site-out cross-validation (LOSO-CV), the model achieved an average cross-site accuracy of 82.0%, outperforming existing methods by approximately 3% under 10-fold cross-validation and 7% under LOSO-CV. The proposed framework effectively integrates heterogeneous multimodal data from the multi-site ABIDE-I dataset, improving automated ASD classification across imaging sites.
NucFuseRank: Dataset Fusion and Performance Ranking for Nuclei Instance Segmentation
Jan 27, 2026Abstract:Nuclei instance segmentation in hematoxylin and eosin (H&E)-stained images plays an important role in automated histological image analysis, with various applications in downstream tasks. While several machine learning and deep learning approaches have been proposed for nuclei instance segmentation, most research in this field focuses on developing new segmentation algorithms and benchmarking them on a limited number of arbitrarily selected public datasets. In this work, rather than focusing on model development, we focused on the datasets used for this task. Based on an extensive literature review, we identified manually annotated, publicly available datasets of H&E-stained images for nuclei instance segmentation and standardized them into a unified input and annotation format. Using two state-of-the-art segmentation models, one based on convolutional neural networks (CNNs) and one based on a hybrid CNN and vision transformer architecture, we systematically evaluated and ranked these datasets based on their nuclei instance segmentation performance. Furthermore, we proposed a unified test set (NucFuse-test) for fair cross-dataset evaluation and a unified training set (NucFuse-train) for improved segmentation performance by merging images from multiple datasets. By evaluating and ranking the datasets, performing comprehensive analyses, generating fused datasets, conducting external validation, and making our implementation publicly available, we provided a new benchmark for training, testing, and evaluating nuclei instance segmentation models on H&E-stained histological images.
Developing Predictive and Robust Radiomics Models for Chemotherapy Response in High-Grade Serous Ovarian Carcinoma
Jan 13, 2026Abstract:Objectives: High-grade serous ovarian carcinoma (HGSOC) is typically diagnosed at an advanced stage with extensive peritoneal metastases, making treatment challenging. Neoadjuvant chemotherapy (NACT) is often used to reduce tumor burden before surgery, but about 40% of patients show limited response. Radiomics, combined with machine learning (ML), offers a promising non-invasive method for predicting NACT response by analyzing computed tomography (CT) imaging data. This study aimed to improve response prediction in HGSOC patients undergoing NACT by integration different feature selection methods. Materials and methods: A framework for selecting robust radiomics features was introduced by employing an automated randomisation algorithm to mimic inter-observer variability, ensuring a balance between feature robustness and prediction accuracy. Four response metrics were used: chemotherapy response score (CRS), RECIST, volume reduction (VolR), and diameter reduction (DiaR). Lesions in different anatomical sites were studied. Pre- and post-NACT CT scans were used for feature extraction and model training on one cohort, and an independent cohort was used for external testing. Results: The best prediction performance was achieved using all lesions combined for VolR prediction, with an AUC of 0.83. Omental lesions provided the best results for CRS prediction (AUC 0.77), while pelvic lesions performed best for DiaR (AUC 0.76). Conclusion: The integration of robustness into the feature selection processes ensures the development of reliable models and thus facilitates the implementation of the radiomics models in clinical applications for HGSOC patients. Future work should explore further applications of radiomics in ovarian cancer, particularly in real-time clinical settings.
From ACR O-RADS 2022 to Explainable Deep Learning: Comparative Performance of Expert Radiologists, Convolutional Neural Networks, Vision Transformers, and Fusion Models in Ovarian Masses
Nov 09, 2025Abstract:Background: The 2022 update of the Ovarian-Adnexal Reporting and Data System (O-RADS) ultrasound classification refines risk stratification for adnexal lesions, yet human interpretation remains subject to variability and conservative thresholds. Concurrently, deep learning (DL) models have demonstrated promise in image-based ovarian lesion characterization. This study evaluates radiologist performance applying O-RADS v2022, compares it to leading convolutional neural network (CNN) and Vision Transformer (ViT) models, and investigates the diagnostic gains achieved by hybrid human-AI frameworks. Methods: In this single-center, retrospective cohort study, a total of 512 adnexal mass images from 227 patients (110 with at least one malignant cyst) were included. Sixteen DL models, including DenseNets, EfficientNets, ResNets, VGGs, Xception, and ViTs, were trained and validated. A hybrid model integrating radiologist O-RADS scores with DL-predicted probabilities was also built for each scheme. Results: Radiologist-only O-RADS assessment achieved an AUC of 0.683 and an overall accuracy of 68.0%. CNN models yielded AUCs of 0.620 to 0.908 and accuracies of 59.2% to 86.4%, while ViT16-384 reached the best performance, with an AUC of 0.941 and an accuracy of 87.4%. Hybrid human-AI frameworks further significantly enhanced the performance of CNN models; however, the improvement for ViT models was not statistically significant (P-value >0.05). Conclusions: DL models markedly outperform radiologist-only O-RADS v2022 assessment, and the integration of expert scores with AI yields the highest diagnostic accuracy and discrimination. Hybrid human-AI paradigms hold substantial potential to standardize pelvic ultrasound interpretation, reduce false positives, and improve detection of high-risk lesions.
A Review on the Integration of Artificial Intelligence and Medical Imaging in IVF Ovarian Stimulation
Dec 27, 2024



Abstract:Artificial intelligence (AI) has emerged as a powerful tool to enhance decision-making and optimize treatment protocols in in vitro fertilization (IVF). In particular, AI shows significant promise in supporting decision-making during the ovarian stimulation phase of the IVF process. This review evaluates studies focused on the applications of AI combined with medical imaging in ovarian stimulation, examining methodologies, outcomes, and current limitations. Our analysis of 13 studies on this topic reveals that, reveal that while AI algorithms demonstrated notable potential in predicting optimal hormonal dosages, trigger timing, and oocyte retrieval outcomes, the medical imaging data utilized predominantly came from two-dimensional (2D) ultrasound which mainly involved basic quantifications, such as follicle size and number, with limited use of direct feature extraction or advanced image analysis techniques. This points to an underexplored opportunity where advanced image analysis approaches, such as deep learning, and more diverse imaging modalities, like three-dimensional (3D) ultrasound, could unlock deeper insights. Additionally, the lack of explainable AI (XAI) in most studies raises concerns about the transparency and traceability of AI-driven decisions - key factors for clinical adoption and trust. Furthermore, many studies relied on single-center designs and small datasets, which limit the generalizability of their findings. This review highlights the need for integrating advanced imaging analysis techniques with explainable AI methodologies, as well as the importance of leveraging multicenter collaborations and larger datasets. Addressing these gaps has the potential to enhance ovarian stimulation management, paving the way for efficient, personalized, and data-driven treatment pathways that improve IVF outcomes.
An Integrated Optimization and Deep Learning Pipeline for Predicting Live Birth Success in IVF Using Feature Optimization and Transformer-Based Models
Dec 27, 2024Abstract:In vitro fertilization (IVF) is a widely utilized assisted reproductive technology, yet predicting its success remains challenging due to the multifaceted interplay of clinical, demographic, and procedural factors. This study develops a robust artificial intelligence (AI) pipeline aimed at predicting live birth outcomes in IVF treatments. The pipeline uses anonymized data from 2010 to 2018, obtained from the Human Fertilization and Embryology Authority (HFEA). We evaluated the prediction performance of live birth success as a binary outcome (success/failure) by integrating different feature selection methods, such as principal component analysis (PCA) and particle swarm optimization (PSO), with different traditional machine learning-based classifiers including random forest (RF) and decision tree, as well as deep learning-based classifiers including custom transformer-based model and a tab transformer model with an attention mechanism. Our research demonstrated that the best performance was achieved by combining PSO for feature selection with the TabTransformer-based deep learning model, yielding an accuracy of 99.50% and an AUC of 99.96%, highlighting its significant performance to predict live births. This study establishes a highly accurate AI pipeline for predicting live birth outcomes in IVF, demonstrating its potential to enhance personalized fertility treatments.
Advanced Hybrid Deep Learning Model for Enhanced Classification of Osteosarcoma Histopathology Images
Oct 29, 2024



Abstract:Recent advances in machine learning are transforming medical image analysis, particularly in cancer detection and classification. Techniques such as deep learning, especially convolutional neural networks (CNNs) and vision transformers (ViTs), are now enabling the precise analysis of complex histopathological images, automating detection, and enhancing classification accuracy across various cancer types. This study focuses on osteosarcoma (OS), the most common bone cancer in children and adolescents, which affects the long bones of the arms and legs. Early and accurate detection of OS is essential for improving patient outcomes and reducing mortality. However, the increasing prevalence of cancer and the demand for personalized treatments create challenges in achieving precise diagnoses and customized therapies. We propose a novel hybrid model that combines convolutional neural networks (CNN) and vision transformers (ViT) to improve diagnostic accuracy for OS using hematoxylin and eosin (H&E) stained histopathological images. The CNN model extracts local features, while the ViT captures global patterns from histopathological images. These features are combined and classified using a Multi-Layer Perceptron (MLP) into four categories: non-tumor (NT), non-viable tumor (NVT), viable tumor (VT), and none-viable ratio (NVR). Using the Cancer Imaging Archive (TCIA) dataset, the model achieved an accuracy of 99.08%, precision of 99.10%, recall of 99.28%, and an F1-score of 99.23%. This is the first successful four-class classification using this dataset, setting a new benchmark in OS research and offering promising potential for future diagnostic advancements.
Evaluating Pre-trained Convolutional Neural Networks and Foundation Models as Feature Extractors for Content-based Medical Image Retrieval
Sep 14, 2024
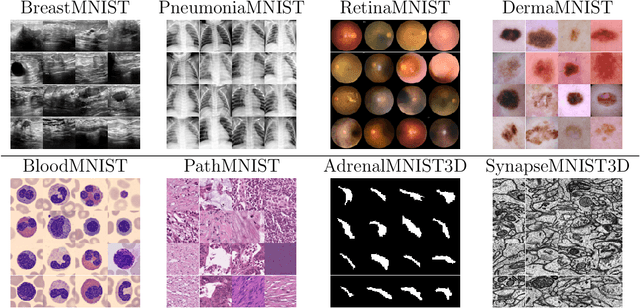


Abstract:Medical image retrieval refers to the task of finding similar images for given query images in a database, with applications such as diagnosis support, treatment planning, and educational tools for inexperienced medical practitioners. While traditional medical image retrieval was performed using clinical metadata, content-based medical image retrieval (CBMIR) relies on the characteristic features of the images, such as color, texture, shape, and spatial features. Many approaches have been proposed for CBMIR, and among them, using pre-trained convolutional neural networks (CNNs) is a widely utilized approach. However, considering the recent advances in the development of foundation models for various computer vision tasks, their application for CBMIR can be also investigated for its potentially superior performance. In this study, we used several pre-trained feature extractors from well-known pre-trained CNNs (VGG19, ResNet-50, DenseNet121, and EfficientNetV2M) and pre-trained foundation models (MedCLIP, BioMedCLIP, OpenCLIP, CONCH and UNI) and investigated the CBMIR performance on a subset of the MedMNIST V2 dataset, including eight types of 2D and 3D medical images. Furthermore, we also investigated the effect of image size on the CBMIR performance. Our results show that, overall, for the 2D datasets, foundation models deliver superior performance by a large margin compared to CNNs, with UNI providing the best overall performance across all datasets and image sizes. For 3D datasets, CNNs and foundation models deliver more competitive performance, with CONCH achieving the best overall performance. Moreover, our findings confirm that while using larger image sizes (especially for 2D datasets) yields slightly better performance, competitive CBMIR performance can still be achieved even with smaller image sizes. Our codes to generate and reproduce the results are available on GitHub.
A study of why we need to reassess full reference image quality assessment with medical images
May 29, 2024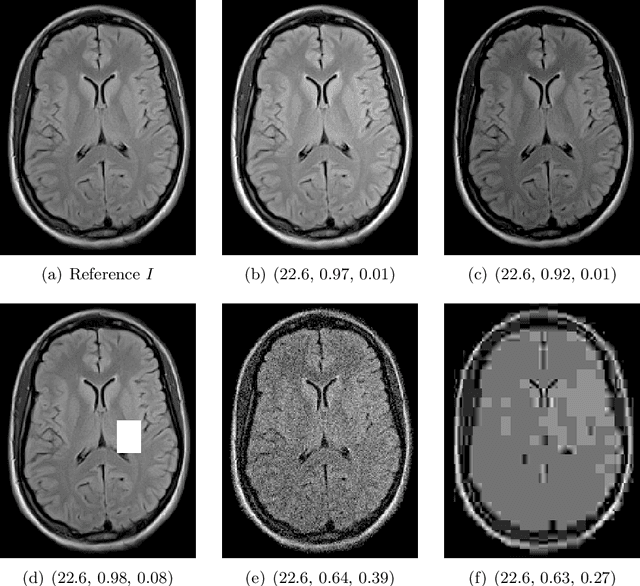
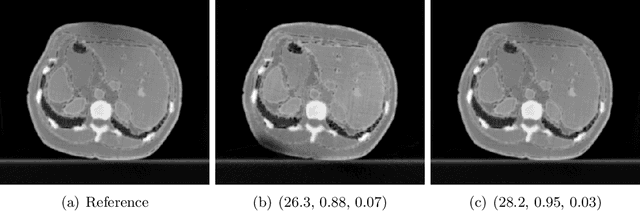
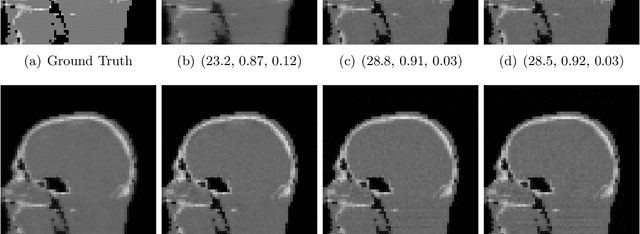
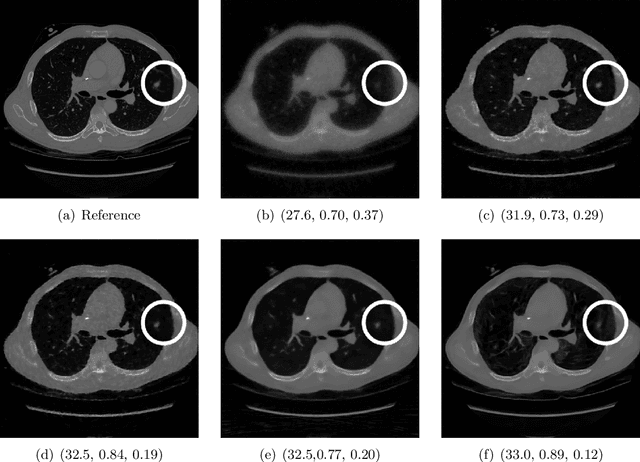

Abstract:Image quality assessment (IQA) is not just indispensable in clinical practice to ensure high standards, but also in the development stage of novel algorithms that operate on medical images with reference data. This paper provides a structured and comprehensive collection of examples where the two most common full reference (FR) image quality measures prove to be unsuitable for the assessment of novel algorithms using different kinds of medical images, including real-world MRI, CT, OCT, X-Ray, digital pathology and photoacoustic imaging data. In particular, the FR-IQA measures PSNR and SSIM are known and tested for working successfully in many natural imaging tasks, but discrepancies in medical scenarios have been noted in the literature. Inconsistencies arising in medical images are not surprising, as they have very different properties than natural images which have not been targeted nor tested in the development of the mentioned measures, and therefore might imply wrong judgement of novel methods for medical images. Therefore, improvement is urgently needed in particular in this era of AI to increase explainability, reproducibility and generalizability in machine learning for medical imaging and beyond. On top of the pitfalls we will provide ideas for future research as well as suggesting guidelines for the usage of FR-IQA measures applied to medical images.
Breast Histopathology Image Retrieval by Attention-based Adversarially Regularized Variational Graph Autoencoder with Contrastive Learning-Based Feature Extraction
May 07, 2024



Abstract:Breast cancer is a significant global health concern, particularly for women. Early detection and appropriate treatment are crucial in mitigating its impact, with histopathology examinations playing a vital role in swift diagnosis. However, these examinations often require a substantial workforce and experienced medical experts for proper recognition and cancer grading. Automated image retrieval systems have the potential to assist pathologists in identifying cancerous tissues, thereby accelerating the diagnostic process. Nevertheless, due to considerable variability among the tissue and cell patterns in histological images, proposing an accurate image retrieval model is very challenging. This work introduces a novel attention-based adversarially regularized variational graph autoencoder model for breast histological image retrieval. Additionally, we incorporated cluster-guided contrastive learning as the graph feature extractor to boost the retrieval performance. We evaluated the proposed model's performance on two publicly available datasets of breast cancer histological images and achieved superior or very competitive retrieval performance, with average mAP scores of 96.5% for the BreakHis dataset and 94.7% for the BACH dataset, and mVP scores of 91.9% and 91.3%, respectively. Our proposed retrieval model has the potential to be used in clinical settings to enhance diagnostic performance and ultimately benefit patients.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge