Alexander Unger
Realistic 3D printed imaging tumor phantoms for validation of image processing algorithms
Nov 27, 2022



Abstract:Medical imaging phantoms are widely used for validation and verification of imaging systems and algorithms in surgical guidance and radiation oncology procedures. Especially, for the performance evaluation of new algorithms in the field of medical imaging, manufactured phantoms need to replicate specific properties of the human body, e.g., tissue morphology and radiological properties. Additive manufacturing (AM) technology provides an inexpensive opportunity for accurate anatomical replication with customization capabilities. In this study, we proposed a simple and cheap protocol to manufacture realistic tumor phantoms based on the filament 3D printing technology. Tumor phantoms with both homogenous and heterogenous radiodensity were fabricated. The radiodensity similarity between the printed tumor models and real tumor data from CT images of lung cancer patients was evaluated. Additionally, it was investigated whether a heterogeneity in the 3D printed tumor phantoms as observed in the tumor patient data had an influence on the validation of image registration algorithms. A density range between -217 to 226 HUs was achieved for 3D printed phantoms; this range of radiation attenuation is also observed in the human lung tumor tissue. The resulted HU range could serve as a lookup-table for researchers and phantom manufactures to create realistic CT tumor phantoms with the desired range of radiodensities. The 3D printed tumor phantoms also precisely replicated real lung tumor patient data regarding morphology and could also include life-like heterogeneity of the radiodensity inside the tumor models. An influence of the heterogeneity on accuracy and robustness of the image registration algorithms was not found.
clDice -- a Topology-Preserving Loss Function for Tubular Structure Segmentation
Mar 29, 2020

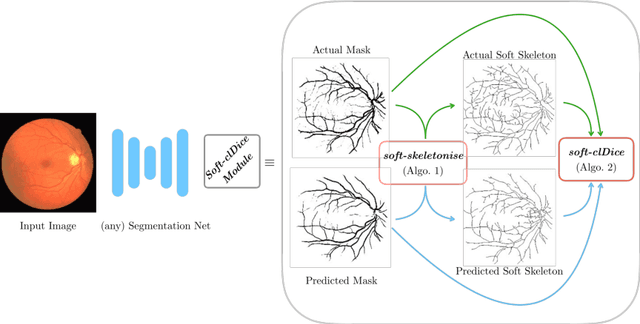
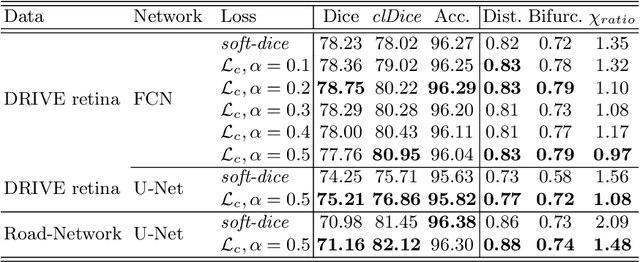
Abstract:Accurate segmentation of tubular, network-like structures, such as vessels, neurons, or roads, is relevant to many fields of research. For such structures, the topology is their most important characteristic, e.g. preserving connectedness: in case of vascular networks, missing a connected vessel entirely alters the blood-flow dynamics. We introduce a novel similarity measure termed clDice, which is calculated on the intersection of the segmentation masks and their (morphological) skeletons. Crucially, we theoretically prove that clDice guarantees topological correctness for binary 2D and 3D segmentation. Extending this, we propose a computationally efficient, differentiable soft-clDice as a loss function for training arbitrary neural segmentation networks. We benchmark the soft-clDice loss for segmentation on four public datasets (2D and 3D). Training on soft-clDice leads to segmentation with more accurate connectivity information, higher graph similarity, and better volumetric scores.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge