Robbie Holland
on behalf of the PINNACLE consortium
Parametric shape models for vessels learned from segmentations via differentiable voxelization
Jul 03, 2025Abstract:Vessels are complex structures in the body that have been studied extensively in multiple representations. While voxelization is the most common of them, meshes and parametric models are critical in various applications due to their desirable properties. However, these representations are typically extracted through segmentations and used disjointly from each other. We propose a framework that joins the three representations under differentiable transformations. By leveraging differentiable voxelization, we automatically extract a parametric shape model of the vessels through shape-to-segmentation fitting, where we learn shape parameters from segmentations without the explicit need for ground-truth shape parameters. The vessel is parametrized as centerlines and radii using cubic B-splines, ensuring smoothness and continuity by construction. Meshes are differentiably extracted from the learned shape parameters, resulting in high-fidelity meshes that can be manipulated post-fit. Our method can accurately capture the geometry of complex vessels, as demonstrated by the volumetric fits in experiments on aortas, aneurysms, and brain vessels.
Specialist vision-language models for clinical ophthalmology
Jul 11, 2024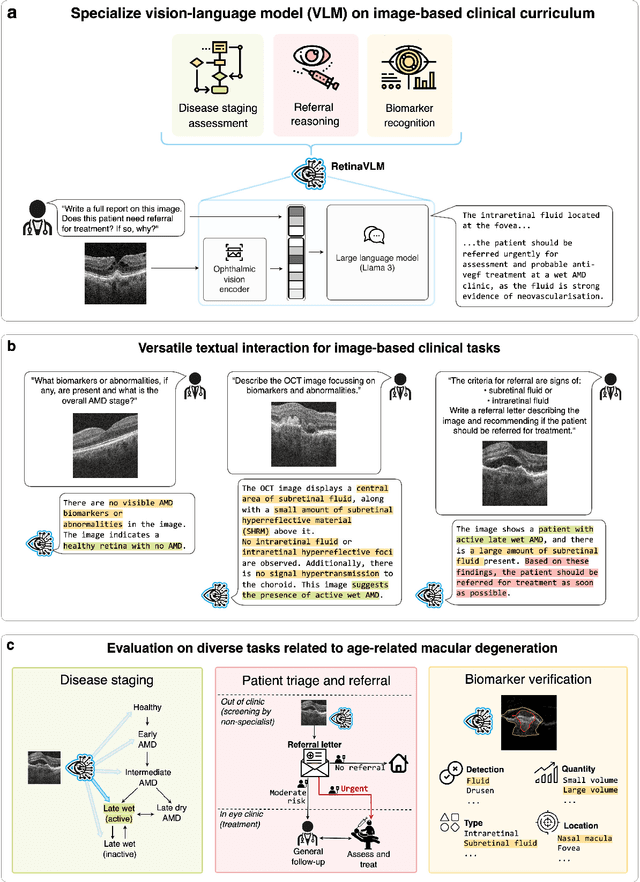

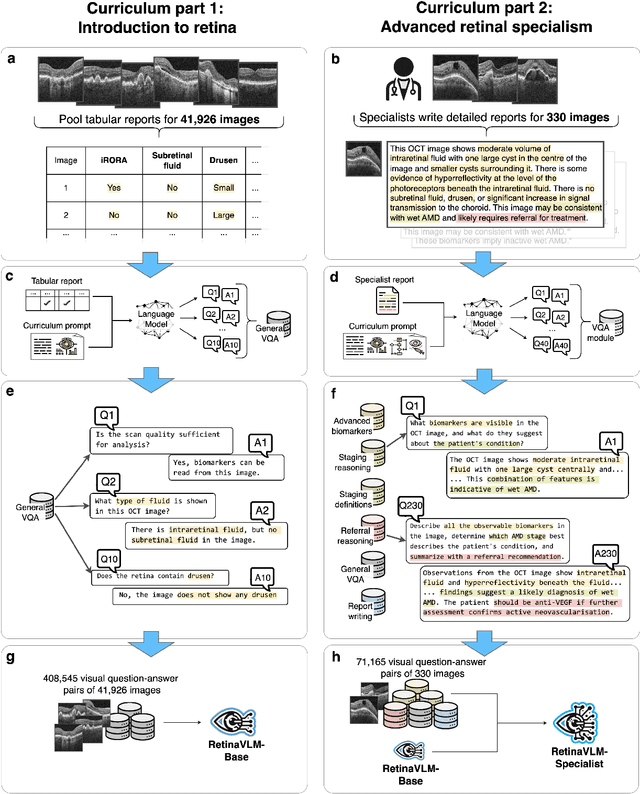

Abstract:Clinicians spend a significant amount of time reviewing medical images and transcribing their findings regarding patient diagnosis, referral and treatment in text form. Vision-language models (VLMs), which automatically interpret images and summarize their findings as text, have enormous potential to alleviate clinical workloads and increase patient access to high-quality medical care. While foundational models have stirred considerable interest in the medical community, it is unclear whether their general capabilities translate to real-world clinical utility. In this work, we show that foundation VLMs markedly underperform compared to practicing ophthalmologists on specialist tasks crucial to the care of patients with age-related macular degeneration (AMD). To address this, we initially identified the essential capabilities required for image-based clinical decision-making, and then developed a curriculum to selectively train VLMs in these skills. The resulting model, RetinaVLM, can be instructed to write reports that significantly outperform those written by leading foundation medical VLMs in disease staging (F1 score of 0.63 vs. 0.11) and patient referral (0.67 vs. 0.39), and approaches the diagnostic performance of junior ophthalmologists (who achieve 0.77 and 0.78 on the respective tasks). Furthermore, in a reader study involving two senior ophthalmologists with up to 32 years of experience, RetinaVLM's reports were found to be similarly correct (78.6% vs. 82.1%) and complete (both 78.6%) as reports written by junior ophthalmologists with up to 10 years of experience. These results demonstrate that our curriculum-based approach provides a blueprint for specializing generalist foundation medical VLMs to handle real-world clinical tasks.
Spatiotemporal Representation Learning for Short and Long Medical Image Time Series
Mar 12, 2024Abstract:Analyzing temporal developments is crucial for the accurate prognosis of many medical conditions. Temporal changes that occur over short time scales are key to assessing the health of physiological functions, such as the cardiac cycle. Moreover, tracking longer term developments that occur over months or years in evolving processes, such as age-related macular degeneration (AMD), is essential for accurate prognosis. Despite the importance of both short and long term analysis to clinical decision making, they remain understudied in medical deep learning. State of the art methods for spatiotemporal representation learning, developed for short natural videos, prioritize the detection of temporal constants rather than temporal developments. Moreover, they do not account for varying time intervals between acquisitions, which are essential for contextualizing observed changes. To address these issues, we propose two approaches. First, we combine clip-level contrastive learning with a novel temporal embedding to adapt to irregular time series. Second, we propose masking and predicting latent frame representations of the temporal sequence. Our two approaches outperform all prior methods on temporally-dependent tasks including cardiac output estimation and three prognostic AMD tasks. Overall, this enables the automated analysis of temporal patterns which are typically overlooked in applications of deep learning to medicine.
A skeletonization algorithm for gradient-based optimization
Sep 05, 2023Abstract:The skeleton of a digital image is a compact representation of its topology, geometry, and scale. It has utility in many computer vision applications, such as image description, segmentation, and registration. However, skeletonization has only seen limited use in contemporary deep learning solutions. Most existing skeletonization algorithms are not differentiable, making it impossible to integrate them with gradient-based optimization. Compatible algorithms based on morphological operations and neural networks have been proposed, but their results often deviate from the geometry and topology of the true medial axis. This work introduces the first three-dimensional skeletonization algorithm that is both compatible with gradient-based optimization and preserves an object's topology. Our method is exclusively based on matrix additions and multiplications, convolutional operations, basic non-linear functions, and sampling from a uniform probability distribution, allowing it to be easily implemented in any major deep learning library. In benchmarking experiments, we prove the advantages of our skeletonization algorithm compared to non-differentiable, morphological, and neural-network-based baselines. Finally, we demonstrate the utility of our algorithm by integrating it with two medical image processing applications that use gradient-based optimization: deep-learning-based blood vessel segmentation, and multimodal registration of the mandible in computed tomography and magnetic resonance images.
Clustering disease trajectories in contrastive feature space for biomarker discovery in age-related macular degeneration
Jan 11, 2023Abstract:Age-related macular degeneration (AMD) is the leading cause of blindness in the elderly. Despite this, the exact dynamics of disease progression are poorly understood. There is a clear need for imaging biomarkers in retinal optical coherence tomography (OCT) that aid the diagnosis, prognosis and management of AMD. However, current grading systems, which coarsely group disease stage into broad categories describing early and intermediate AMD, have very limited prognostic value for the conversion to late AMD. In this paper, we are the first to analyse disease progression as clustered trajectories in a self-supervised feature space. Our method first pretrains an encoder with contrastive learning to project images from longitudinal time series to points in feature space. This enables the creation of disease trajectories, which are then denoised, partitioned and grouped into clusters. These clusters, found in two datasets containing time series of 7,912 patients imaged over eight years, were correlated with known OCT biomarkers. This reinforced efforts by four expert ophthalmologists to investigate clusters, during a clinical comparison and interpretation task, as candidates for time-dependent biomarkers that describe progression of AMD.
Metadata-enhanced contrastive learning from retinal optical coherence tomography images
Aug 04, 2022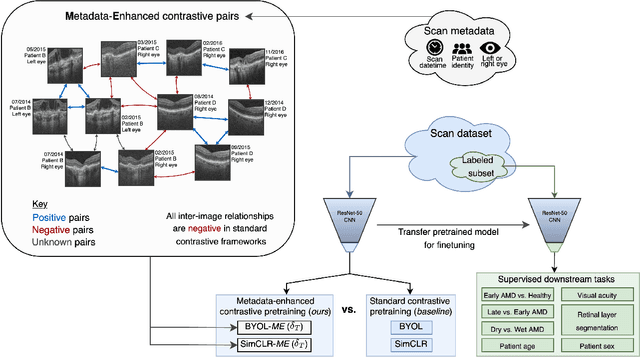
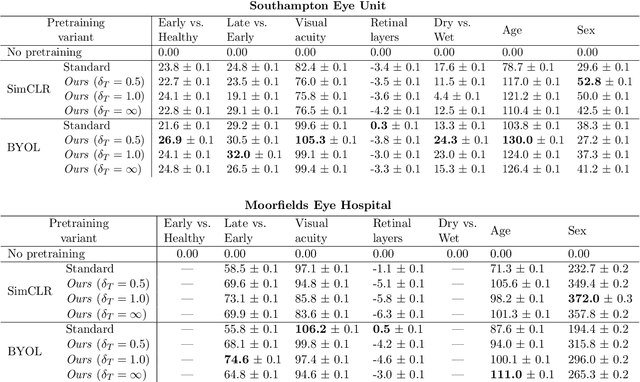
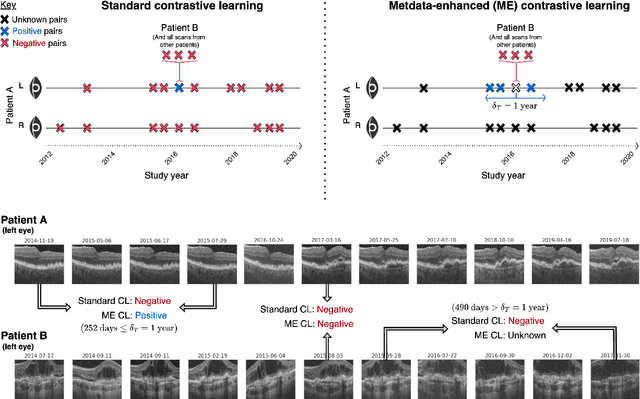
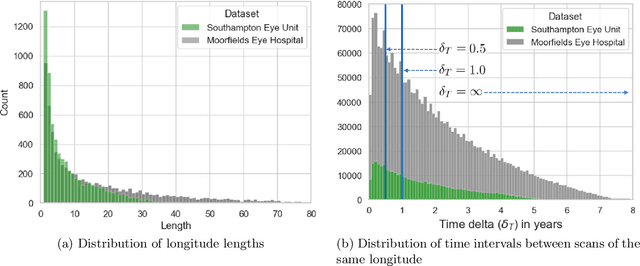
Abstract:Supervised deep learning algorithms hold great potential to automate screening, monitoring and grading of medical images. However, training performant models has typically required vast quantities of labelled data, which is scarcely available in the medical domain. Self-supervised contrastive frameworks relax this dependency by first learning from unlabelled images. In this work we show that pretraining with two contrastive methods, SimCLR and BYOL, improves the utility of deep learning with regard to the clinical assessment of age-related macular degeneration (AMD). In experiments using two large clinical datasets containing 170,427 optical coherence tomography (OCT) images of 7,912 patients, we evaluate benefits attributed to pretraining across seven downstream tasks ranging from AMD stage and type classification to prediction of functional endpoints to segmentation of retinal layers, finding performance significantly increased in six out of seven tasks with fewer labels. However, standard contrastive frameworks have two known weaknesses that are detrimental to pretraining in the medical domain. Several of the image transformations used to create positive contrastive pairs are not applicable to greyscale medical scans. Furthermore, medical images often depict the same anatomical region and disease severity, resulting in numerous misleading negative pairs. To address these issues we develop a novel metadata-enhanced approach that exploits the rich set of inherently available patient information. To this end we employ records for patient identity, eye position (i.e. left or right) and time series data to indicate the typically unknowable set of inter-image contrastive relationships. By leveraging this often neglected information our metadata-enhanced contrastive pretraining leads to further benefits and outperforms conventional contrastive methods in five out of seven downstream tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge