Andrew Lotery
Stochastic Siamese MAE Pretraining for Longitudinal Medical Images
Dec 29, 2025Abstract:Temporally aware image representations are crucial for capturing disease progression in 3D volumes of longitudinal medical datasets. However, recent state-of-the-art self-supervised learning approaches like Masked Autoencoding (MAE), despite their strong representation learning capabilities, lack temporal awareness. In this paper, we propose STAMP (Stochastic Temporal Autoencoder with Masked Pretraining), a Siamese MAE framework that encodes temporal information through a stochastic process by conditioning on the time difference between the 2 input volumes. Unlike deterministic Siamese approaches, which compare scans from different time points but fail to account for the inherent uncertainty in disease evolution, STAMP learns temporal dynamics stochastically by reframing the MAE reconstruction loss as a conditional variational inference objective. We evaluated STAMP on two OCT and one MRI datasets with multiple visits per patient. STAMP pretrained ViT models outperformed both existing temporal MAE methods and foundation models on different late stage Age-Related Macular Degeneration and Alzheimer's Disease progression prediction which require models to learn the underlying non-deterministic temporal dynamics of the diseases.
Forecasting Disease Progression with Parallel Hyperplanes in Longitudinal Retinal OCT
Sep 30, 2024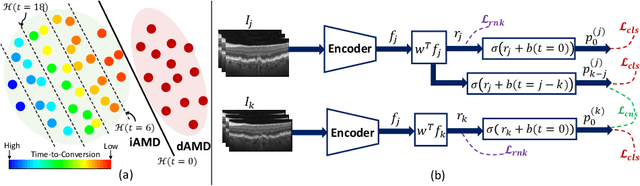
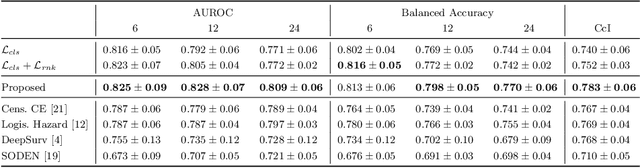
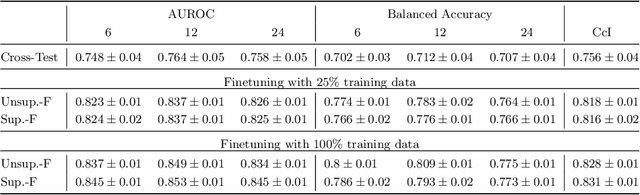
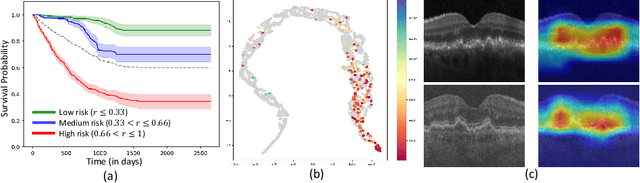
Abstract:Predicting future disease progression risk from medical images is challenging due to patient heterogeneity, and subtle or unknown imaging biomarkers. Moreover, deep learning (DL) methods for survival analysis are susceptible to image domain shifts across scanners. We tackle these issues in the task of predicting late dry Age-related Macular Degeneration (dAMD) onset from retinal OCT scans. We propose a novel DL method for survival prediction to jointly predict from the current scan a risk score, inversely related to time-to-conversion, and the probability of conversion within a time interval $t$. It uses a family of parallel hyperplanes generated by parameterizing the bias term as a function of $t$. In addition, we develop unsupervised losses based on intra-subject image pairs to ensure that risk scores increase over time and that future conversion predictions are consistent with AMD stage prediction using actual scans of future visits. Such losses enable data-efficient fine-tuning of the trained model on new unlabeled datasets acquired with a different scanner. Extensive evaluation on two large datasets acquired with different scanners resulted in a mean AUROCs of 0.82 for Dataset-1 and 0.83 for Dataset-2, across prediction intervals of 6,12 and 24 months.
Spatiotemporal Representation Learning for Short and Long Medical Image Time Series
Mar 12, 2024Abstract:Analyzing temporal developments is crucial for the accurate prognosis of many medical conditions. Temporal changes that occur over short time scales are key to assessing the health of physiological functions, such as the cardiac cycle. Moreover, tracking longer term developments that occur over months or years in evolving processes, such as age-related macular degeneration (AMD), is essential for accurate prognosis. Despite the importance of both short and long term analysis to clinical decision making, they remain understudied in medical deep learning. State of the art methods for spatiotemporal representation learning, developed for short natural videos, prioritize the detection of temporal constants rather than temporal developments. Moreover, they do not account for varying time intervals between acquisitions, which are essential for contextualizing observed changes. To address these issues, we propose two approaches. First, we combine clip-level contrastive learning with a novel temporal embedding to adapt to irregular time series. Second, we propose masking and predicting latent frame representations of the temporal sequence. Our two approaches outperform all prior methods on temporally-dependent tasks including cardiac output estimation and three prognostic AMD tasks. Overall, this enables the automated analysis of temporal patterns which are typically overlooked in applications of deep learning to medicine.
3DTINC: Time-Equivariant Non-Contrastive Learning for Predicting Disease Progression from Longitudinal OCTs
Dec 28, 2023



Abstract:Self-supervised learning (SSL) has emerged as a powerful technique for improving the efficiency and effectiveness of deep learning models. Contrastive methods are a prominent family of SSL that extract similar representations of two augmented views of an image while pushing away others in the representation space as negatives. However, the state-of-the-art contrastive methods require large batch sizes and augmentations designed for natural images that are impractical for 3D medical images. To address these limitations, we propose a new longitudinal SSL method, 3DTINC, based on non-contrastive learning. It is designed to learn perturbation-invariant features for 3D optical coherence tomography (OCT) volumes, using augmentations specifically designed for OCT. We introduce a new non-contrastive similarity loss term that learns temporal information implicitly from intra-patient scans acquired at different times. Our experiments show that this temporal information is crucial for predicting progression of retinal diseases, such as age-related macular degeneration (AMD). After pretraining with 3DTINC, we evaluated the learned representations and the prognostic models on two large-scale longitudinal datasets of retinal OCTs where we predict the conversion to wet-AMD within a six months interval. Our results demonstrate that each component of our contributions is crucial for learning meaningful representations useful in predicting disease progression from longitudinal volumetric scans.
Pretrained Deep 2.5D Models for Efficient Predictive Modeling from Retinal OCT
Jul 25, 2023Abstract:In the field of medical imaging, 3D deep learning models play a crucial role in building powerful predictive models of disease progression. However, the size of these models presents significant challenges, both in terms of computational resources and data requirements. Moreover, achieving high-quality pretraining of 3D models proves to be even more challenging. To address these issues, hybrid 2.5D approaches provide an effective solution for utilizing 3D volumetric data efficiently using 2D models. Combining 2D and 3D techniques offers a promising avenue for optimizing performance while minimizing memory requirements. In this paper, we explore 2.5D architectures based on a combination of convolutional neural networks (CNNs), long short-term memory (LSTM), and Transformers. In addition, leveraging the benefits of recent non-contrastive pretraining approaches in 2D, we enhanced the performance and data efficiency of 2.5D techniques even further. We demonstrate the effectiveness of architectures and associated pretraining on a task of predicting progression to wet age-related macular degeneration (AMD) within a six-month period on two large longitudinal OCT datasets.
Morph-SSL: Self-Supervision with Longitudinal Morphing to Predict AMD Progression from OCT
Apr 17, 2023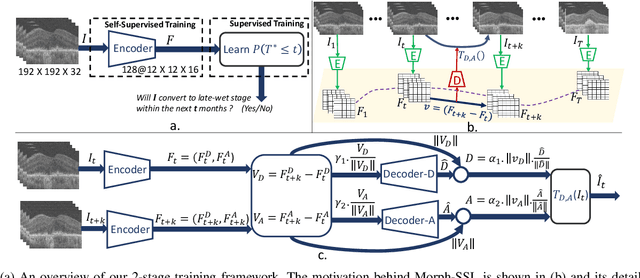
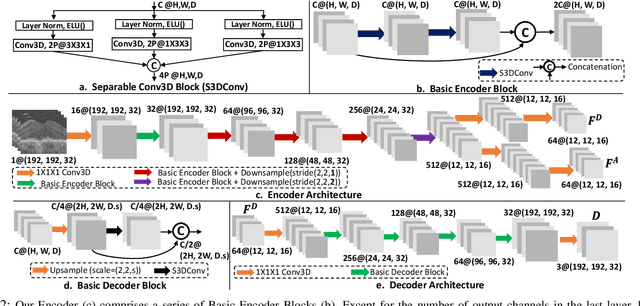

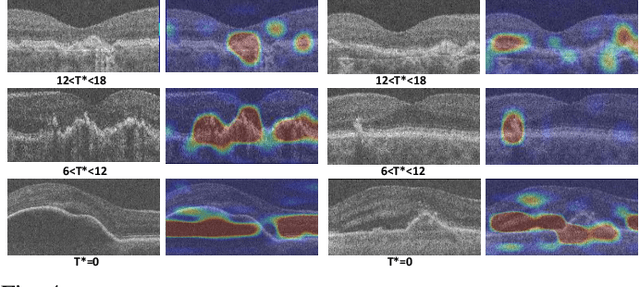
Abstract:The lack of reliable biomarkers makes predicting the conversion from intermediate to neovascular age-related macular degeneration (iAMD, nAMD) a challenging task. We develop a Deep Learning (DL) model to predict the future risk of conversion of an eye from iAMD to nAMD from its current OCT scan. Although eye clinics generate vast amounts of longitudinal OCT scans to monitor AMD progression, only a small subset can be manually labeled for supervised DL. To address this issue, we propose Morph-SSL, a novel Self-supervised Learning (SSL) method for longitudinal data. It uses pairs of unlabelled OCT scans from different visits and involves morphing the scan from the previous visit to the next. The Decoder predicts the transformation for morphing and ensures a smooth feature manifold that can generate intermediate scans between visits through linear interpolation. Next, the Morph-SSL trained features are input to a Classifier which is trained in a supervised manner to model the cumulative probability distribution of the time to conversion with a sigmoidal function. Morph-SSL was trained on unlabelled scans of 399 eyes (3570 visits). The Classifier was evaluated with a five-fold cross-validation on 2418 scans from 343 eyes with clinical labels of the conversion date. The Morph-SSL features achieved an AUC of 0.766 in predicting the conversion to nAMD within the next 6 months, outperforming the same network when trained end-to-end from scratch or pre-trained with popular SSL methods. Automated prediction of the future risk of nAMD onset can enable timely treatment and individualized AMD management.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge