Renjiu Hu
Gaussian Primitive Optimized Deformable Retinal Image Registration
Aug 23, 2025



Abstract:Deformable retinal image registration is notoriously difficult due to large homogeneous regions and sparse but critical vascular features, which cause limited gradient signals in standard learning-based frameworks. In this paper, we introduce Gaussian Primitive Optimization (GPO), a novel iterative framework that performs structured message passing to overcome these challenges. After an initial coarse alignment, we extract keypoints at salient anatomical structures (e.g., major vessels) to serve as a minimal set of descriptor-based control nodes (DCN). Each node is modelled as a Gaussian primitive with trainable position, displacement, and radius, thus adapting its spatial influence to local deformation scales. A K-Nearest Neighbors (KNN) Gaussian interpolation then blends and propagates displacement signals from these information-rich nodes to construct a globally coherent displacement field; focusing interpolation on the top (K) neighbors reduces computational overhead while preserving local detail. By strategically anchoring nodes in high-gradient regions, GPO ensures robust gradient flow, mitigating vanishing gradient signal in textureless areas. The framework is optimized end-to-end via a multi-term loss that enforces both keypoint consistency and intensity alignment. Experiments on the FIRE dataset show that GPO reduces the target registration error from 6.2\,px to ~2.4\,px and increases the AUC at 25\,px from 0.770 to 0.938, substantially outperforming existing methods. The source code can be accessed via https://github.com/xintian-99/GPOreg.
Fidelity-Imposed Displacement Editing for the Learn2Reg 2024 SHG-BF Challenge
Oct 28, 2024



Abstract:Co-examination of second-harmonic generation (SHG) and bright-field (BF) microscopy enables the differentiation of tissue components and collagen fibers, aiding the analysis of human breast and pancreatic cancer tissues. However, large discrepancies between SHG and BF images pose challenges for current learning-based registration models in aligning SHG to BF. In this paper, we propose a novel multi-modal registration framework that employs fidelity-imposed displacement editing to address these challenges. The framework integrates batch-wise contrastive learning, feature-based pre-alignment, and instance-level optimization. Experimental results from the Learn2Reg COMULISglobe SHG-BF Challenge validate the effectiveness of our method, securing the 1st place on the online leaderboard.
Large Scale Unsupervised Brain MRI Image Registration Solution for Learn2Reg 2024
Sep 04, 2024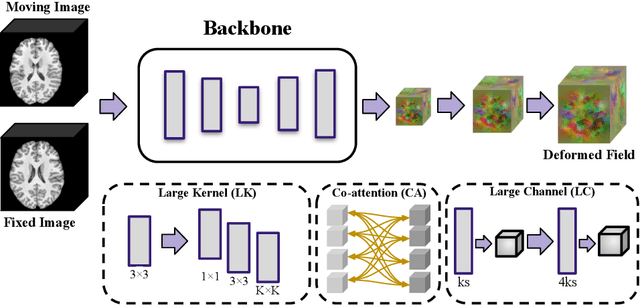
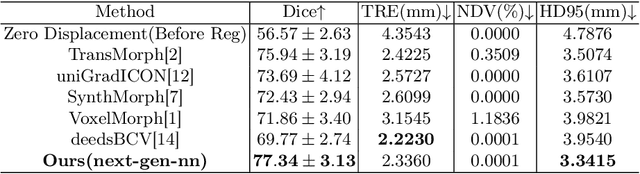
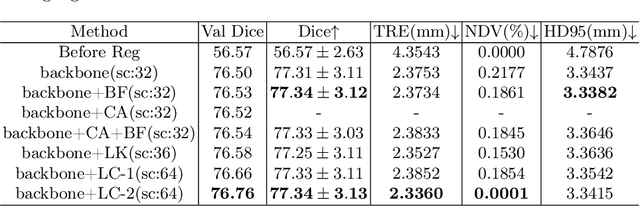
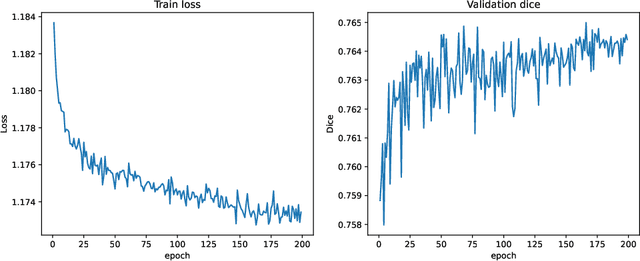
Abstract:In this paper, we summarize the methods and experimental results we proposed for Task 2 in the learn2reg 2024 Challenge. This task focuses on unsupervised registration of anatomical structures in brain MRI images between different patients. The difficulty lies in: (1) without segmentation labels, and (2) a large amount of data. To address these challenges, we built an efficient backbone network and explored several schemes to further enhance registration accuracy. Under the guidance of the NCC loss function and smoothness regularization loss function, we obtained a smooth and reasonable deformation field. According to the leaderboard, our method achieved a Dice coefficient of 77.34%, which is 1.4% higher than the TransMorph. Overall, we won second place on the leaderboard for Task 2.
MemWarp: Discontinuity-Preserving Cardiac Registration with Memorized Anatomical Filters
Jul 10, 2024Abstract:Many existing learning-based deformable image registration methods impose constraints on deformation fields to ensure they are globally smooth and continuous. However, this assumption does not hold in cardiac image registration, where different anatomical regions exhibit asymmetric motions during respiration and movements due to sliding organs within the chest. Consequently, such global constraints fail to accommodate local discontinuities across organ boundaries, potentially resulting in erroneous and unrealistic displacement fields. In this paper, we address this issue with MemWarp, a learning framework that leverages a memory network to store prototypical information tailored to different anatomical regions. MemWarp is different from earlier approaches in two main aspects: firstly, by decoupling feature extraction from similarity matching in moving and fixed images, it facilitates more effective utilization of feature maps; secondly, despite its capability to preserve discontinuities, it eliminates the need for segmentation masks during model inference. In experiments on a publicly available cardiac dataset, our method achieves considerable improvements in registration accuracy and producing realistic deformations, outperforming state-of-the-art methods with a remarkable 7.1\% Dice score improvement over the runner-up semi-supervised method. Source code will be available at https://github.com/tinymilky/Mem-Warp.
Slicer Networks
Jan 18, 2024Abstract:In medical imaging, scans often reveal objects with varied contrasts but consistent internal intensities or textures. This characteristic enables the use of low-frequency approximations for tasks such as segmentation and deformation field estimation. Yet, integrating this concept into neural network architectures for medical image analysis remains underexplored. In this paper, we propose the Slicer Network, a novel architecture designed to leverage these traits. Comprising an encoder utilizing models like vision transformers for feature extraction and a slicer employing a learnable bilateral grid, the Slicer Network strategically refines and upsamples feature maps via a splatting-blurring-slicing process. This introduces an edge-preserving low-frequency approximation for the network outcome, effectively enlarging the effective receptive field. The enhancement not only reduces computational complexity but also boosts overall performance. Experiments across different medical imaging applications, including unsupervised and keypoints-based image registration and lesion segmentation, have verified the Slicer Network's improved accuracy and efficiency.
RimSet: Quantitatively Identifying and Characterizing Chronic Active Multiple Sclerosis Lesion on Quantitative Susceptibility Maps
Dec 28, 2023Abstract:Background: Rim+ lesions in multiple sclerosis (MS), detectable via Quantitative Susceptibility Mapping (QSM), correlate with increased disability. Existing literature lacks quantitative analysis of these lesions. We introduce RimSet for quantitative identification and characterization of rim+ lesions on QSM. Methods: RimSet combines RimSeg, an unsupervised segmentation method using level-set methodology, and radiomic measurements with Local Binary Pattern texture descriptors. We validated RimSet using simulated QSM images and an in vivo dataset of 172 MS subjects with 177 rim+ and 3986 rim-lesions. Results: RimSeg achieved a 78.7% Dice score against the ground truth, with challenges in partial rim lesions. RimSet detected rim+ lesions with a partial ROC AUC of 0.808 and PR AUC of 0.737, surpassing existing methods. QSMRim-Net showed the lowest mean square error (0.85) and high correlation (0.91; 95% CI: 0.88, 0.93) with expert annotations at the subject level.
Spatially Covariant Image Registration with Text Prompts
Nov 27, 2023



Abstract:Medical images are often characterized by their structured anatomical representations and spatially inhomogeneous contrasts. Leveraging anatomical priors in neural networks can greatly enhance their utility in resource-constrained clinical settings. Prior research has harnessed such information for image segmentation, yet progress in deformable image registration has been modest. Our work introduces textSCF, a novel method that integrates spatially covariant filters and textual anatomical prompts encoded by visual-language models, to fill this gap. This approach optimizes an implicit function that correlates text embeddings of anatomical regions to filter weights, relaxing the typical translation-invariance constraint of convolutional operations. TextSCF not only boosts computational efficiency but can also retain or improve registration accuracy. By capturing the contextual interplay between anatomical regions, it offers impressive inter-regional transferability and the ability to preserve structural discontinuities during registration. TextSCF's performance has been rigorously tested on inter-subject brain MRI and abdominal CT registration tasks, outperforming existing state-of-the-art models in the MICCAI Learn2Reg 2021 challenge and leading the leaderboard. In abdominal registrations, textSCF's larger model variant improved the Dice score by 11.3% over the second-best model, while its smaller variant maintained similar accuracy but with an 89.13% reduction in network parameters and a 98.34\% decrease in computational operations.
Multi-delay arterial spin-labeled perfusion estimation with biophysics simulation and deep learning
Nov 17, 2023



Abstract:Purpose: To develop biophysics-based method for estimating perfusion Q from arterial spin labeling (ASL) images using deep learning. Methods: A 3D U-Net (QTMnet) was trained to estimate perfusion from 4D tracer propagation images. The network was trained and tested on simulated 4D tracer concentration data based on artificial vasculature structure generated by constrained constructive optimization (CCO) method. The trained network was further tested in a synthetic brain ASL image based on vasculature network extracted from magnetic resonance (MR) angiography. The estimations from both trained network and a conventional kinetic model were compared in ASL images acquired from eight healthy volunteers. Results: QTMnet accurately reconstructed perfusion Q from concentration data. Relative error of the synthetic brain ASL image was 7.04% for perfusion Q, lower than the error using single-delay ASL model: 25.15% for Q, and multi-delay ASL model: 12.62% for perfusion Q. Conclusion: QTMnet provides accurate estimation on perfusion parameters and is a promising approach as a clinical ASL MRI image processing pipeline.
DAGrid: Directed Accumulator Grid
Jun 05, 2023Abstract:Recent research highlights that the Directed Accumulator (DA), through its parametrization of geometric priors into neural networks, has notably improved the performance of medical image recognition, particularly with small and imbalanced datasets. However, DA's potential in pixel-wise dense predictions is unexplored. To bridge this gap, we present the Directed Accumulator Grid (DAGrid), which allows geometric-preserving filtering in neural networks, thus broadening the scope of DA's applications to include pixel-level dense prediction tasks. DAGrid utilizes homogeneous data types in conjunction with designed sampling grids to construct geometrically transformed representations, retaining intricate geometric information and promoting long-range information propagation within the neural networks. Contrary to its symmetric counterpart, grid sampling, which might lose information in the sampling process, DAGrid aggregates all pixels, ensuring a comprehensive representation in the transformed space. The parallelization of DAGrid on modern GPUs is facilitated using CUDA programming, and also back propagation is enabled for deep neural network training. Empirical results show DAGrid-enhanced neural networks excel in supervised skin lesion segmentation and unsupervised cardiac image registration. Specifically, the network incorporating DAGrid has realized a 70.8% reduction in network parameter size and a 96.8% decrease in FLOPs, while concurrently improving the Dice score for skin lesion segmentation by 1.0% compared to state-of-the-art transformers. Furthermore, it has achieved improvements of 4.4% and 8.2% in the average Dice score and Dice score of the left ventricular mass, respectively, indicating an increase in registration accuracy for cardiac images. The source code is available at https://github.com/tinymilky/DeDA.
DeDA: Deep Directed Accumulator
Mar 15, 2023Abstract:Chronic active multiple sclerosis lesions, also termed as rim+ lesions, can be characterized by a hyperintense rim at the edge of the lesion on quantitative susceptibility maps. These rim+ lesions exhibit a geometrically simple structure, where gradients at the lesion edge are radially oriented and a greater magnitude of gradients is observed in contrast to rim- (non rim+) lesions. However, recent studies have shown that the identification performance of such lesions remains unsatisfied due to the limited amount of data and high class imbalance. In this paper, we propose a simple yet effective image processing operation, deep directed accumulator (DeDA), that provides a new perspective for injecting domain-specific inductive biases (priors) into neural networks for rim+ lesion identification. Given a feature map and a set of sampling grids, DeDA creates and quantizes an accumulator space into finite intervals, and accumulates feature values accordingly. This DeDA operation is a generalized discrete Radon transform and can also be regarded as a symmetric operation to the grid sampling within the forward-backward neural network framework, the process of which is order-agnostic, and can be efficiently implemented with the native CUDA programming. Experimental results on a dataset with 177 rim+ and 3986 rim- lesions show that 10.1% of improvement in a partial (false positive rate<0.1) area under the receiver operating characteristic curve (pROC AUC) and 10.2% of improvement in an area under the precision recall curve (PR AUC) can be achieved respectively comparing to other state-of-the-art methods. The source code is available online at https://github.com/tinymilky/DeDA
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge