Omid Rohanian
Improving Recursive Transformers with Mixture of LoRAs
Dec 17, 2025



Abstract:Parameter sharing in recursive transformers reduces model size but collapses layer-wise expressivity. We propose Mixture of LoRAs (MoL), a lightweight conditional-computation mechanism that inserts Low-Rank Adaptation (LoRA) experts inside a shared feed-forward network (FFN). MoL enables token-conditional weight-space modulation of the shared FFN without untying backbone parameters, unlike prior approaches that add fixed or externally attached adapters. We pretrain a modernised recursive architecture, ModernALBERT, integrating rotary embeddings, GeGLU, FlashAttention, and a distillation-based initialisation. Across GLUE, SQuAD-v2, and BEIR, ModernALBERT (50M--120M) achieves state-of-the-art performance among compact models and surpasses larger fully parameterised baselines. We also propose an expert-merging procedure that compresses MoL into a single adapter at inference while preserving accuracy, enabling efficient deployment. Our results show that conditional weight-space modulation effectively restores the expressivity lost under aggressive parameter sharing in recursive transformers.
Rapid Biomedical Research Classification: The Pandemic PACT Advanced Categorisation Engine
Jul 14, 2024



Abstract:This paper introduces the Pandemic PACT Advanced Categorisation Engine (PPACE) along with its associated dataset. PPACE is a fine-tuned model developed to automatically classify research abstracts from funded biomedical projects according to WHO-aligned research priorities. This task is crucial for monitoring research trends and identifying gaps in global health preparedness and response. Our approach builds on human-annotated projects, which are allocated one or more categories from a predefined list. A large language model is then used to generate `rationales' explaining the reasoning behind these annotations. This augmented data, comprising expert annotations and rationales, is subsequently used to fine-tune a smaller, more efficient model. Developed as part of the Pandemic PACT project, which aims to track and analyse research funding and clinical evidence for a wide range of diseases with outbreak potential, PPACE supports informed decision-making by research funders, policymakers, and independent researchers. We introduce and release both the trained model and the instruction-based dataset used for its training. Our evaluation shows that PPACE significantly outperforms its baselines. The release of PPACE and its associated dataset offers valuable resources for researchers in multilabel biomedical document classification and supports advancements in aligning biomedical research with key global health priorities.
Large Language Models in Healthcare: A Comprehensive Benchmark
Apr 25, 2024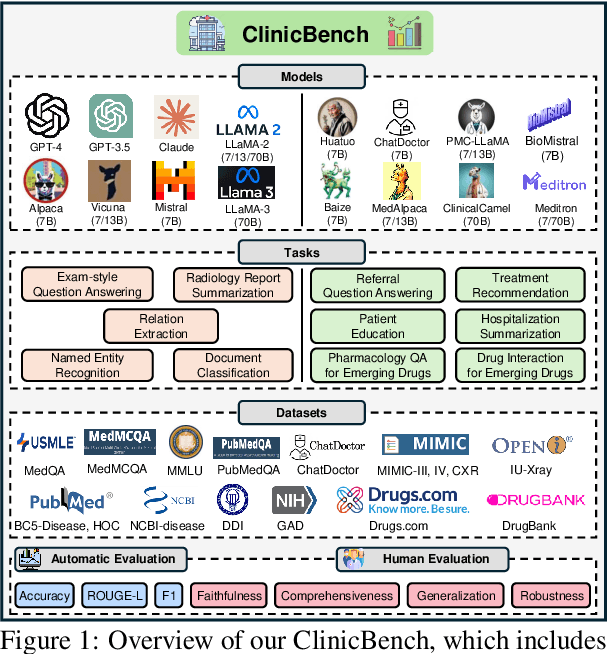



Abstract:The adoption of large language models (LLMs) to assist clinicians has attracted remarkable attention. Existing works mainly adopt the close-ended question-answering task with answer options for evaluation. However, in real clinical settings, many clinical decisions, such as treatment recommendations, involve answering open-ended questions without pre-set options. Meanwhile, existing studies mainly use accuracy to assess model performance. In this paper, we comprehensively benchmark diverse LLMs in healthcare, to clearly understand their strengths and weaknesses. Our benchmark contains seven tasks and thirteen datasets across medical language generation, understanding, and reasoning. We conduct a detailed evaluation of the existing sixteen LLMs in healthcare under both zero-shot and few-shot (i.e., 1,3,5-shot) learning settings. We report the results on five metrics (i.e. matching, faithfulness, comprehensiveness, generalizability, and robustness) that are critical in achieving trust from clinical users. We further invite medical experts to conduct human evaluation.
Efficiency at Scale: Investigating the Performance of Diminutive Language Models in Clinical Tasks
Feb 16, 2024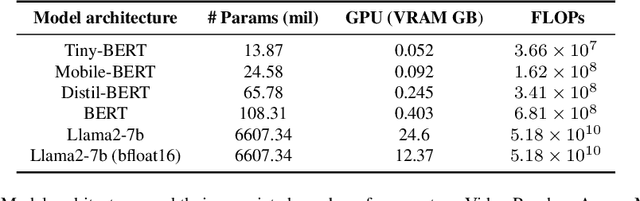
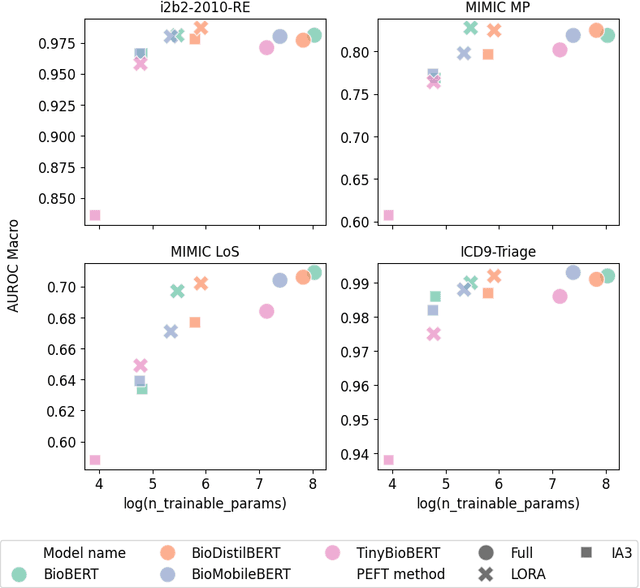
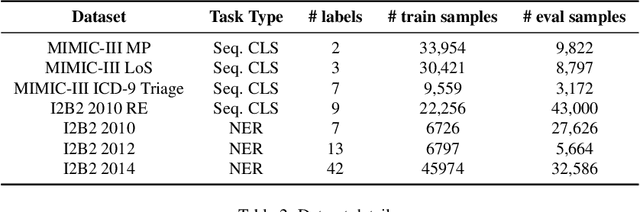
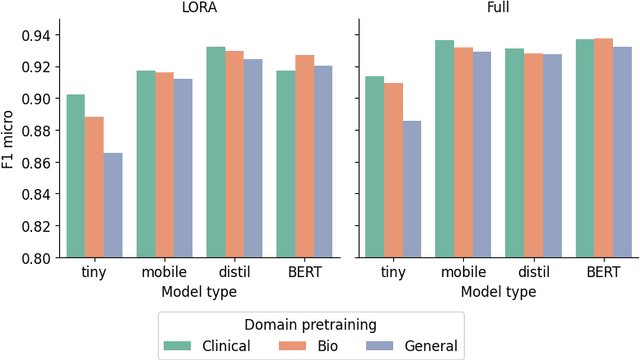
Abstract:The entry of large language models (LLMs) into research and commercial spaces has led to a trend of ever-larger models, with initial promises of generalisability, followed by a widespread desire to downsize and create specialised models without the need for complete fine-tuning, using Parameter Efficient Fine-tuning (PEFT) methods. We present an investigation into the suitability of different PEFT methods to clinical decision-making tasks, across a range of model sizes, including extremely small models with as few as $25$ million parameters. Our analysis shows that the performance of most PEFT approaches varies significantly from one task to another, with the exception of LoRA, which maintains relatively high performance across all model sizes and tasks, typically approaching or matching full fine-tuned performance. The effectiveness of PEFT methods in the clinical domain is evident, particularly for specialised models which can operate on low-cost, in-house computing infrastructure. The advantages of these models, in terms of speed and reduced training costs, dramatically outweighs any performance gain from large foundation LLMs. Furthermore, we highlight how domain-specific pre-training interacts with PEFT methods and model size, and discuss how these factors interplay to provide the best efficiency-performance trade-off. Full code available at: tbd.
Exploring the Effectiveness of Instruction Tuning in Biomedical Language Processing
Dec 31, 2023Abstract:Large Language Models (LLMs), particularly those similar to ChatGPT, have significantly influenced the field of Natural Language Processing (NLP). While these models excel in general language tasks, their performance in domain-specific downstream tasks such as biomedical and clinical Named Entity Recognition (NER), Relation Extraction (RE), and Medical Natural Language Inference (NLI) is still evolving. In this context, our study investigates the potential of instruction tuning for biomedical language processing, applying this technique to two general LLMs of substantial scale. We present a comprehensive, instruction-based model trained on a dataset that consists of approximately $200,000$ instruction-focused samples. This dataset represents a carefully curated compilation of existing data, meticulously adapted and reformatted to align with the specific requirements of our instruction-based tasks. This initiative represents an important step in utilising such models to achieve results on par with specialised encoder-only models like BioBERT and BioClinicalBERT for various classical biomedical NLP tasks. Our work includes an analysis of the dataset's composition and its impact on model performance, providing insights into the intricacies of instruction tuning. By sharing our codes, models, and the distinctively assembled instruction-based dataset, we seek to encourage ongoing research and development in this area.
Lightweight Transformers for Clinical Natural Language Processing
Feb 09, 2023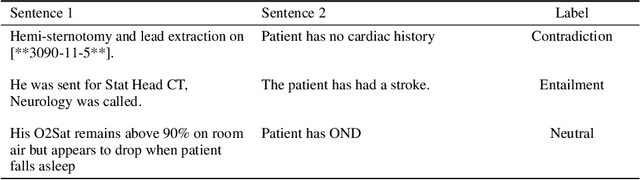
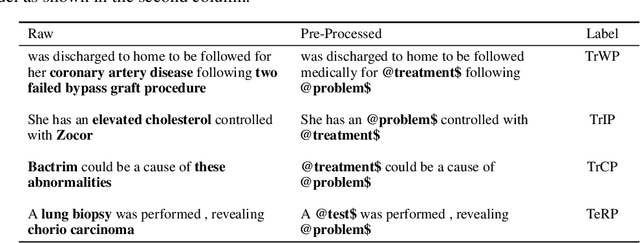
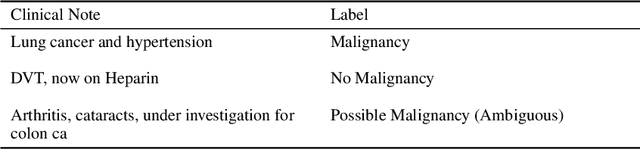
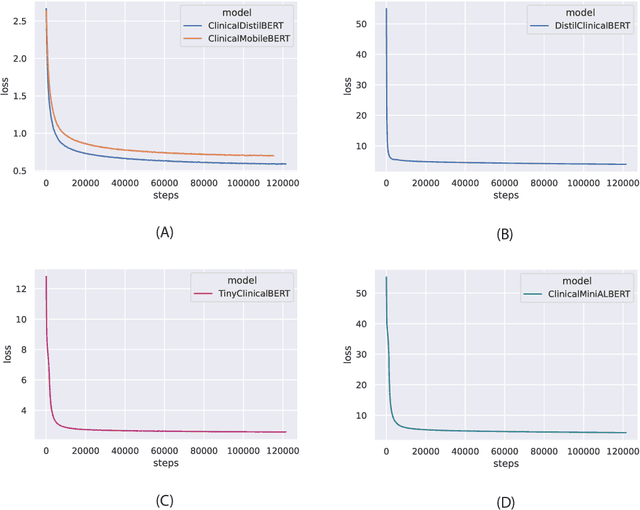
Abstract:Specialised pre-trained language models are becoming more frequent in NLP since they can potentially outperform models trained on generic texts. BioBERT and BioClinicalBERT are two examples of such models that have shown promise in medical NLP tasks. Many of these models are overparametrised and resource-intensive, but thanks to techniques like Knowledge Distillation (KD), it is possible to create smaller versions that perform almost as well as their larger counterparts. In this work, we specifically focus on development of compact language models for processing clinical texts (i.e. progress notes, discharge summaries etc). We developed a number of efficient lightweight clinical transformers using knowledge distillation and continual learning, with the number of parameters ranging from 15 million to 65 million. These models performed comparably to larger models such as BioBERT and ClinicalBioBERT and significantly outperformed other compact models trained on general or biomedical data. Our extensive evaluation was done across several standard datasets and covered a wide range of clinical text-mining tasks, including Natural Language Inference, Relation Extraction, Named Entity Recognition, and Sequence Classification. To our knowledge, this is the first comprehensive study specifically focused on creating efficient and compact transformers for clinical NLP tasks. The models and code used in this study can be found on our Huggingface profile at https://huggingface.co/nlpie and Github page at https://github.com/nlpie-research/Lightweight-Clinical-Transformers, respectively, promoting reproducibility of our results.
Using Bottleneck Adapters to Identify Cancer in Clinical Notes under Low-Resource Constraints
Oct 17, 2022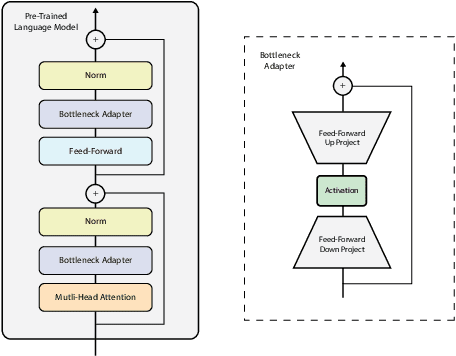
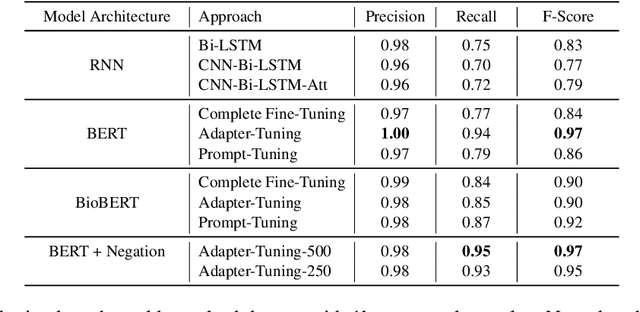
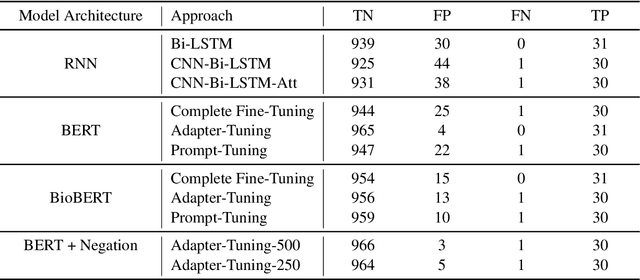
Abstract:Processing information locked within clinical health records is a challenging task that remains an active area of research in biomedical NLP. In this work, we evaluate a broad set of machine learning techniques ranging from simple RNNs to specialised transformers such as BioBERT on a dataset containing clinical notes along with a set of annotations indicating whether a sample is cancer-related or not. Furthermore, we specifically employ efficient fine-tuning methods from NLP, namely, bottleneck adapters and prompt tuning, to adapt the models to our specialised task. Our evaluations suggest that fine-tuning a frozen BERT model pre-trained on natural language and with bottleneck adapters outperforms all other strategies, including full fine-tuning of the specialised BioBERT model. Based on our findings, we suggest that using bottleneck adapters in low-resource situations with limited access to labelled data or processing capacity could be a viable strategy in biomedical text mining. The code used in the experiments are going to be made available at https://github.com/omidrohanian/bottleneck-adapters.
MiniALBERT: Model Distillation via Parameter-Efficient Recursive Transformers
Oct 12, 2022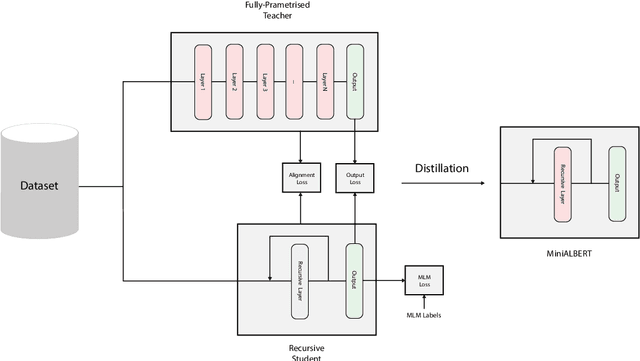
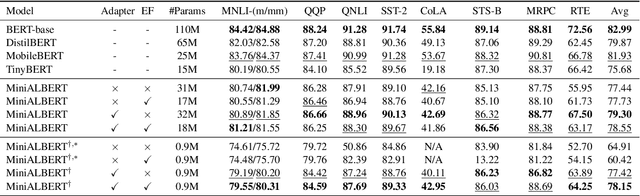
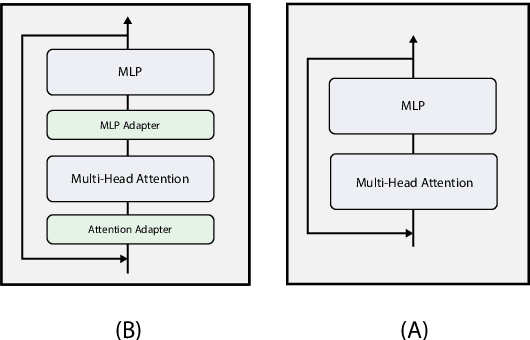
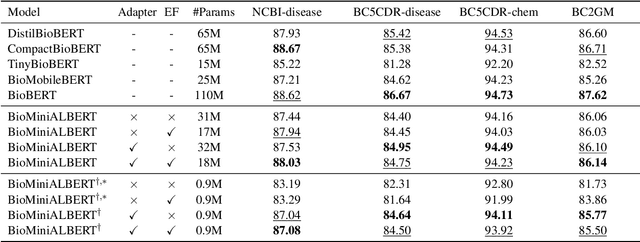
Abstract:Pre-trained Language Models (LMs) have become an integral part of Natural Language Processing (NLP) in recent years, due to their superior performance in downstream applications. In spite of this resounding success, the usability of LMs is constrained by computational and time complexity, along with their increasing size; an issue that has been referred to as `overparameterisation'. Different strategies have been proposed in the literature to alleviate these problems, with the aim to create effective compact models that nearly match the performance of their bloated counterparts with negligible performance losses. One of the most popular techniques in this area of research is model distillation. Another potent but underutilised technique is cross-layer parameter sharing. In this work, we combine these two strategies and present MiniALBERT, a technique for converting the knowledge of fully parameterised LMs (such as BERT) into a compact recursive student. In addition, we investigate the application of bottleneck adapters for layer-wise adaptation of our recursive student, and also explore the efficacy of adapter tuning for fine-tuning of compact models. We test our proposed models on a number of general and biomedical NLP tasks to demonstrate their viability and compare them with the state-of-the-art and other existing compact models. All the codes used in the experiments are available at https://github.com/nlpie-research/MiniALBERT. Our pre-trained compact models can be accessed from https://huggingface.co/nlpie.
On the Effectiveness of Compact Biomedical Transformers
Sep 07, 2022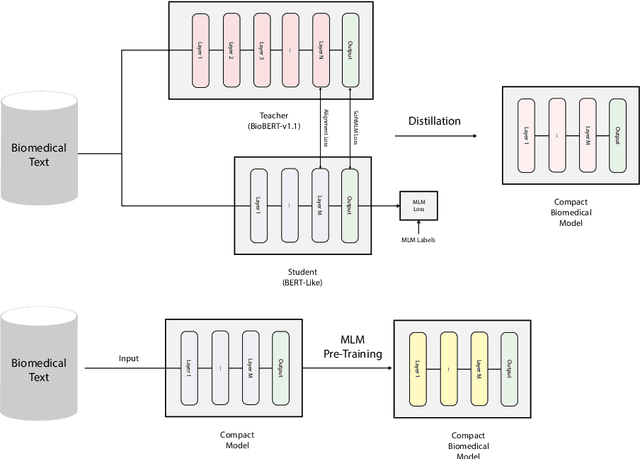
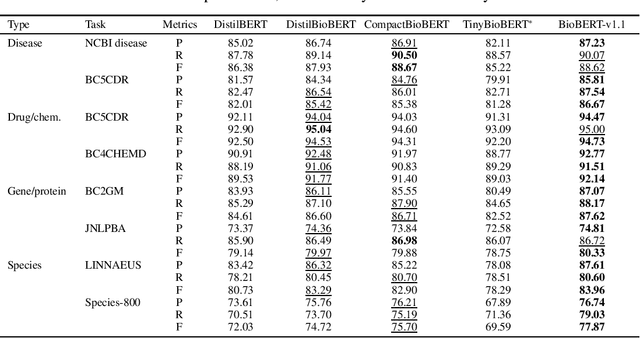
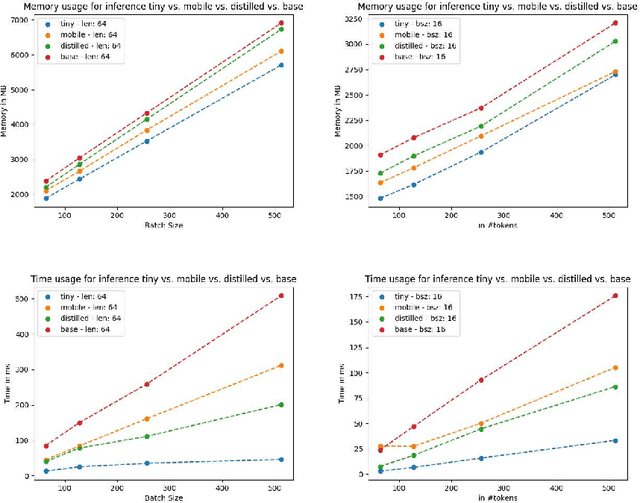
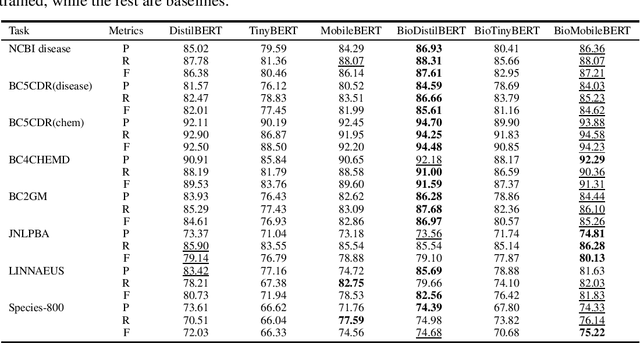
Abstract:Language models pre-trained on biomedical corpora, such as BioBERT, have recently shown promising results on downstream biomedical tasks. Many existing pre-trained models, on the other hand, are resource-intensive and computationally heavy owing to factors such as embedding size, hidden dimension, and number of layers. The natural language processing (NLP) community has developed numerous strategies to compress these models utilising techniques such as pruning, quantisation, and knowledge distillation, resulting in models that are considerably faster, smaller, and subsequently easier to use in practice. By the same token, in this paper we introduce six lightweight models, namely, BioDistilBERT, BioTinyBERT, BioMobileBERT, DistilBioBERT, TinyBioBERT, and CompactBioBERT which are obtained either by knowledge distillation from a biomedical teacher or continual learning on the Pubmed dataset via the Masked Language Modelling (MLM) objective. We evaluate all of our models on three biomedical tasks and compare them with BioBERT-v1.1 to create efficient lightweight models that perform on par with their larger counterparts. All the models will be publicly available on our Huggingface profile at https://huggingface.co/nlpie and the codes used to run the experiments will be available at https://github.com/nlpie-research/Compact-Biomedical-Transformers.
Nowruz at SemEval-2022 Task 7: Tackling Cloze Tests with Transformers and Ordinal Regression
Apr 01, 2022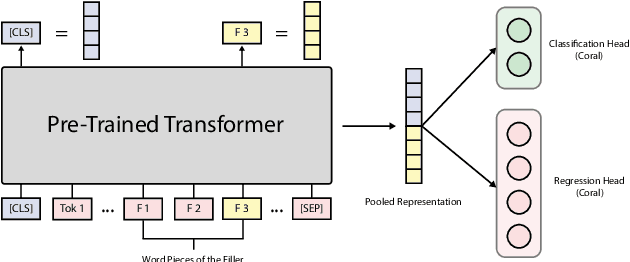

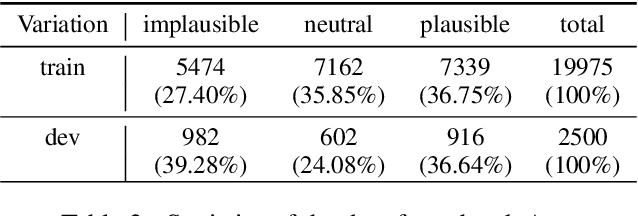
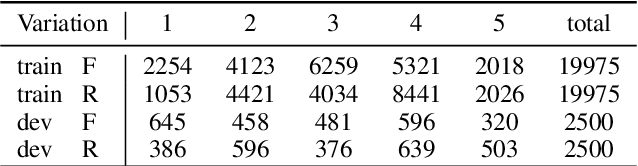
Abstract:This paper outlines the system using which team Nowruz participated in SemEval 2022 Task 7 Identifying Plausible Clarifications of Implicit and Underspecified Phrases for both subtasks A and B. Using a pre-trained transformer as a backbone, the model targeted the task of multi-task classification and ranking in the context of finding the best fillers for a cloze task related to instructional texts on the website Wikihow. The system employed a combination of two ordinal regression components to tackle this task in a multi-task learning scenario. According to the official leaderboard of the shared task, this system was ranked 5th in the ranking and 7th in the classification subtasks out of 21 participating teams. With additional experiments, the models have since been further optimised.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge