Nazim Haouchine
NOIR: Neural Operator mapping for Implicit Representations
Mar 13, 2026Abstract:This paper presents NOIR, a framework that reframes core medical imaging tasks as operator learning between continuous function spaces, challenging the prevailing paradigm of discrete grid-based deep learning. Instead of operating on fixed pixel or voxel grids, NOIR embeds discrete medical signals into shared Implicit Neural Representations and learns a Neural Operator that maps between their latent modulations, enabling resolution-independent function-to-function transformations. We evaluate NOIR across multiple 2D and 3D downstream tasks, including segmentation, shape completion, image-to-image translation, and image synthesis, on several public datasets such as Shenzhen, OASIS-4, SkullBreak, fastMRI, as well as an in-house clinical dataset. It achieves competitive performance at native resolution while demonstrating strong robustness to unseen discretizations, and empirically satisfies key theoretical properties of neural operators. The project page is available here: https://github.com/Sidaty1/NOIR-io.
A Systematic Review on Data-Driven Brain Deformation Modeling for Image-Guided Neurosurgery
Feb 09, 2026Abstract:Accurate compensation of brain deformation is a critical challenge for reliable image-guided neurosurgery, as surgical manipulation and tumor resection induce tissue motion that misaligns preoperative planning images with intraoperative anatomy and longitudinal studies. In this systematic review, we synthesize recent AI-driven approaches developed between January 2020 and April 2025 for modeling and correcting brain deformation. A comprehensive literature search was conducted in PubMed, IEEE Xplore, Scopus, and Web of Science, with predefined inclusion and exclusion criteria focused on computational methods applied to brain deformation compensation for neurosurgical imaging, resulting in 41 studies meeting these criteria. We provide a unified analysis of methodological strategies, including deep learning-based image registration, direct deformation field regression, synthesis-driven multimodal alignment, resection-aware architectures addressing missing correspondences, and hybrid models that integrate biomechanical priors. We also examine dataset utilization, reported evaluation metrics, validation protocols, and how uncertainty and generalization have been assessed across studies. While AI-based deformation models demonstrate promising performance and computational efficiency, current approaches exhibit limitations in out-of-distribution robustness, standardized benchmarking, interpretability, and readiness for clinical deployment. Our review highlights these gaps and outlines opportunities for future research aimed at achieving more robust, generalizable, and clinically translatable deformation compensation solutions for neurosurgical guidance. By organizing recent advances and critically evaluating evaluation practices, this work provides a comprehensive foundation for researchers and clinicians engaged in developing and applying AI-based brain deformation methods.
Surgical Neural Radiance Fields from One Image
Jul 01, 2025Abstract:Purpose: Neural Radiance Fields (NeRF) offer exceptional capabilities for 3D reconstruction and view synthesis, yet their reliance on extensive multi-view data limits their application in surgical intraoperative settings where only limited data is available. In particular, collecting such extensive data intraoperatively is impractical due to time constraints. This work addresses this challenge by leveraging a single intraoperative image and preoperative data to train NeRF efficiently for surgical scenarios. Methods: We leverage preoperative MRI data to define the set of camera viewpoints and images needed for robust and unobstructed training. Intraoperatively, the appearance of the surgical image is transferred to the pre-constructed training set through neural style transfer, specifically combining WTC2 and STROTSS to prevent over-stylization. This process enables the creation of a dataset for instant and fast single-image NeRF training. Results: The method is evaluated with four clinical neurosurgical cases. Quantitative comparisons to NeRF models trained on real surgical microscope images demonstrate strong synthesis agreement, with similarity metrics indicating high reconstruction fidelity and stylistic alignment. When compared with ground truth, our method demonstrates high structural similarity, confirming good reconstruction quality and texture preservation. Conclusion: Our approach demonstrates the feasibility of single-image NeRF training in surgical settings, overcoming the limitations of traditional multi-view methods.
Path and Bone-Contour Regularized Unpaired MRI-to-CT Translation
May 06, 2025



Abstract:Accurate MRI-to-CT translation promises the integration of complementary imaging information without the need for additional imaging sessions. Given the practical challenges associated with acquiring paired MRI and CT scans, the development of robust methods capable of leveraging unpaired datasets is essential for advancing the MRI-to-CT translation. Current unpaired MRI-to-CT translation methods, which predominantly rely on cycle consistency and contrastive learning frameworks, frequently encounter challenges in accurately translating anatomical features that are highly discernible on CT but less distinguishable on MRI, such as bone structures. This limitation renders these approaches less suitable for applications in radiation therapy, where precise bone representation is essential for accurate treatment planning. To address this challenge, we propose a path- and bone-contour regularized approach for unpaired MRI-to-CT translation. In our method, MRI and CT images are projected to a shared latent space, where the MRI-to-CT mapping is modeled as a continuous flow governed by neural ordinary differential equations. The optimal mapping is obtained by minimizing the transition path length of the flow. To enhance the accuracy of translated bone structures, we introduce a trainable neural network to generate bone contours from MRI and implement mechanisms to directly and indirectly encourage the model to focus on bone contours and their adjacent regions. Evaluations conducted on three datasets demonstrate that our method outperforms existing unpaired MRI-to-CT translation approaches, achieving lower overall error rates. Moreover, in a downstream bone segmentation task, our approach exhibits superior performance in preserving the fidelity of bone structures. Our code is available at: https://github.com/kennysyp/PaBoT.
3D/2D Registration of Angiograms using Silhouette-based Differentiable Rendering
Jan 24, 2025


Abstract:We present a method for 3D/2D registration of Digital Subtraction Angiography (DSA) images to provide valuable insight into brain hemodynamics and angioarchitecture. Our approach formulates the registration as a pose estimation problem, leveraging both anteroposterior and lateral DSA views and employing differentiable rendering. Preliminary experiments on real and synthetic datasets demonstrate the effectiveness of our method, with both qualitative and quantitative evaluations highlighting its potential for clinical applications. The code is available at https://github.com/taewoonglee17/TwoViewsDSAReg.
Unified Cross-Modal Image Synthesis with Hierarchical Mixture of Product-of-Experts
Oct 25, 2024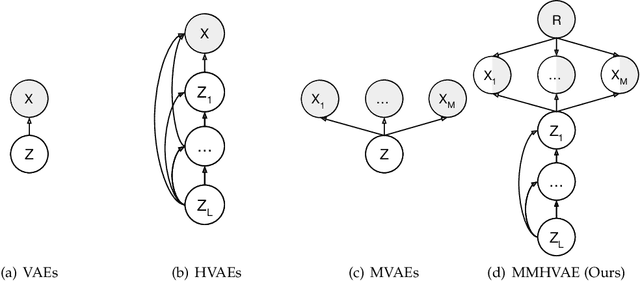



Abstract:We propose a deep mixture of multimodal hierarchical variational auto-encoders called MMHVAE that synthesizes missing images from observed images in different modalities. MMHVAE's design focuses on tackling four challenges: (i) creating a complex latent representation of multimodal data to generate high-resolution images; (ii) encouraging the variational distributions to estimate the missing information needed for cross-modal image synthesis; (iii) learning to fuse multimodal information in the context of missing data; (iv) leveraging dataset-level information to handle incomplete data sets at training time. Extensive experiments are performed on the challenging problem of pre-operative brain multi-parametric magnetic resonance and intra-operative ultrasound imaging.
Intraoperative Registration by Cross-Modal Inverse Neural Rendering
Sep 18, 2024Abstract:We present in this paper a novel approach for 3D/2D intraoperative registration during neurosurgery via cross-modal inverse neural rendering. Our approach separates implicit neural representation into two components, handling anatomical structure preoperatively and appearance intraoperatively. This disentanglement is achieved by controlling a Neural Radiance Field's appearance with a multi-style hypernetwork. Once trained, the implicit neural representation serves as a differentiable rendering engine, which can be used to estimate the surgical camera pose by minimizing the dissimilarity between its rendered images and the target intraoperative image. We tested our method on retrospective patients' data from clinical cases, showing that our method outperforms state-of-the-art while meeting current clinical standards for registration. Code and additional resources can be found at https://maxfehrentz.github.io/style-ngp/.
Learning to Match 2D Keypoints Across Preoperative MR and Intraoperative Ultrasound
Sep 12, 2024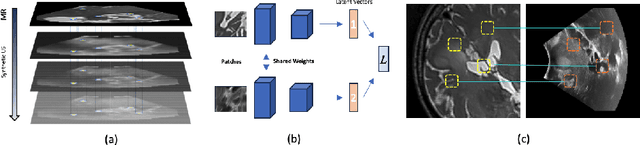



Abstract:We propose in this paper a texture-invariant 2D keypoints descriptor specifically designed for matching preoperative Magnetic Resonance (MR) images with intraoperative Ultrasound (US) images. We introduce a matching-by-synthesis strategy, where intraoperative US images are synthesized from MR images accounting for multiple MR modalities and intraoperative US variability. We build our training set by enforcing keypoints localization over all images then train a patient-specific descriptor network that learns texture-invariant discriminant features in a supervised contrastive manner, leading to robust keypoints descriptors. Our experiments on real cases with ground truth show the effectiveness of the proposed approach, outperforming the state-of-the-art methods and achieving 80.35% matching precision on average.
Patient-Specific Real-Time Segmentation in Trackerless Brain Ultrasound
May 16, 2024



Abstract:Intraoperative ultrasound (iUS) imaging has the potential to improve surgical outcomes in brain surgery. However, its interpretation is challenging, even for expert neurosurgeons. In this work, we designed the first patient-specific framework that performs brain tumor segmentation in trackerless iUS. To disambiguate ultrasound imaging and adapt to the neurosurgeon's surgical objective, a patient-specific real-time network is trained using synthetic ultrasound data generated by simulating virtual iUS sweep acquisitions in pre-operative MR data. Extensive experiments performed in real ultrasound data demonstrate the effectiveness of the proposed approach, allowing for adapting to the surgeon's definition of surgical targets and outperforming non-patient-specific models, neurosurgeon experts, and high-end tracking systems. Our code is available at: \url{https://github.com/ReubenDo/MHVAE-Seg}.
Spatiotemporal Disentanglement of Arteriovenous Malformations in Digital Subtraction Angiography
Feb 15, 2024Abstract:Although Digital Subtraction Angiography (DSA) is the most important imaging for visualizing cerebrovascular anatomy, its interpretation by clinicians remains difficult. This is particularly true when treating arteriovenous malformations (AVMs), where entangled vasculature connecting arteries and veins needs to be carefully identified.The presented method aims to enhance DSA image series by highlighting critical information via automatic classification of vessels using a combination of two learning models: An unsupervised machine learning method based on Independent Component Analysis that decomposes the phases of flow and a convolutional neural network that automatically delineates the vessels in image space. The proposed method was tested on clinical DSA images series and demonstrated efficient differentiation between arteries and veins that provides a viable solution to enhance visualizations for clinical use.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge