Mukund Sudarshan
A dynamic risk score for early prediction of cardiogenic shock using machine learning
Mar 28, 2023



Abstract:Myocardial infarction and heart failure are major cardiovascular diseases that affect millions of people in the US. The morbidity and mortality are highest among patients who develop cardiogenic shock. Early recognition of cardiogenic shock is critical. Prompt implementation of treatment measures can prevent the deleterious spiral of ischemia, low blood pressure, and reduced cardiac output due to cardiogenic shock. However, early identification of cardiogenic shock has been challenging due to human providers' inability to process the enormous amount of data in the cardiac intensive care unit (ICU) and lack of an effective risk stratification tool. We developed a deep learning-based risk stratification tool, called CShock, for patients admitted into the cardiac ICU with acute decompensated heart failure and/or myocardial infarction to predict onset of cardiogenic shock. To develop and validate CShock, we annotated cardiac ICU datasets with physician adjudicated outcomes. CShock achieved an area under the receiver operator characteristic curve (AUROC) of 0.820, which substantially outperformed CardShock (AUROC 0.519), a well-established risk score for cardiogenic shock prognosis. CShock was externally validated in an independent patient cohort and achieved an AUROC of 0.800, demonstrating its generalizability in other cardiac ICUs.
On the Feasibility of Machine Learning Augmented Magnetic Resonance for Point-of-Care Identification of Disease
Feb 02, 2023



Abstract:Early detection of many life-threatening diseases (e.g., prostate and breast cancer) within at-risk population can improve clinical outcomes and reduce cost of care. While numerous disease-specific "screening" tests that are closer to Point-of-Care (POC) are in use for this task, their low specificity results in unnecessary biopsies, leading to avoidable patient trauma and wasteful healthcare spending. On the other hand, despite the high accuracy of Magnetic Resonance (MR) imaging in disease diagnosis, it is not used as a POC disease identification tool because of poor accessibility. The root cause of poor accessibility of MR stems from the requirement to reconstruct high-fidelity images, as it necessitates a lengthy and complex process of acquiring large quantities of high-quality k-space measurements. In this study we explore the feasibility of an ML-augmented MR pipeline that directly infers the disease sidestepping the image reconstruction process. We hypothesise that the disease classification task can be solved using a very small tailored subset of k-space data, compared to image reconstruction. Towards that end, we propose a method that performs two tasks: 1) identifies a subset of the k-space that maximizes disease identification accuracy, and 2) infers the disease directly using the identified k-space subset, bypassing the image reconstruction step. We validate our hypothesis by measuring the performance of the proposed system across multiple diseases and anatomies. We show that comparable performance to image-based classifiers, trained on images reconstructed with full k-space data, can be achieved using small quantities of data: 8% of the data for detecting multiple abnormalities in prostate and brain scans, and 5% of the data for knee abnormalities. To better understand the proposed approach and instigate future research, we provide an extensive analysis and release code.
DIET: Conditional independence testing with marginal dependence measures of residual information
Aug 18, 2022

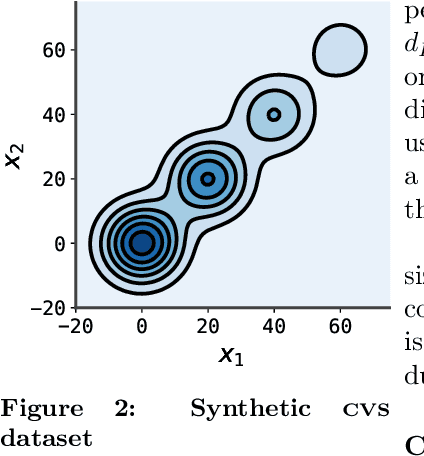

Abstract:Conditional randomization tests (CRTs) assess whether a variable $x$ is predictive of another variable $y$, having observed covariates $z$. CRTs require fitting a large number of predictive models, which is often computationally intractable. Existing solutions to reduce the cost of CRTs typically split the dataset into a train and test portion, or rely on heuristics for interactions, both of which lead to a loss in power. We propose the decoupled independence test (DIET), an algorithm that avoids both of these issues by leveraging marginal independence statistics to test conditional independence relationships. DIET tests the marginal independence of two random variables: $F(x \mid z)$ and $F(y \mid z)$ where $F(\cdot \mid z)$ is a conditional cumulative distribution function (CDF). These variables are termed "information residuals." We give sufficient conditions for DIET to achieve finite sample type-1 error control and power greater than the type-1 error rate. We then prove that when using the mutual information between the information residuals as a test statistic, DIET yields the most powerful conditionally valid test. Finally, we show DIET achieves higher power than other tractable CRTs on several synthetic and real benchmarks.
FastSHAP: Real-Time Shapley Value Estimation
Jul 15, 2021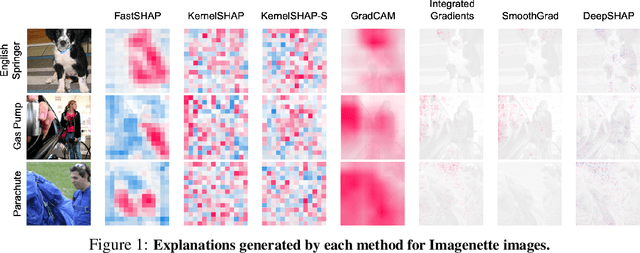
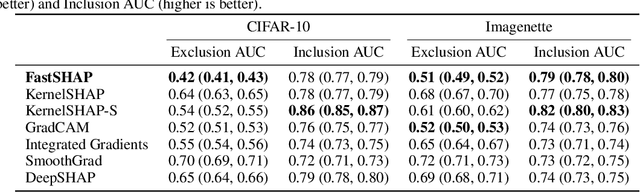
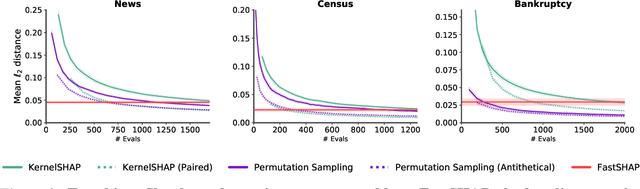
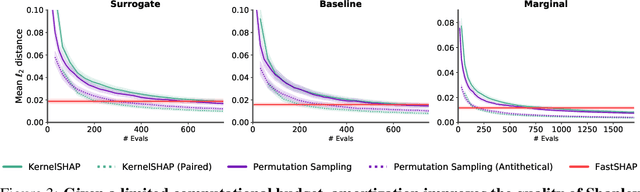
Abstract:Shapley values are widely used to explain black-box models, but they are costly to calculate because they require many model evaluations. We introduce FastSHAP, a method for estimating Shapley values in a single forward pass using a learned explainer model. FastSHAP amortizes the cost of explaining many inputs via a learning approach inspired by the Shapley value's weighted least squares characterization, and it can be trained using standard stochastic gradient optimization. We compare FastSHAP to existing estimation approaches, revealing that it generates high-quality explanations with orders of magnitude speedup.
Have We Learned to Explain?: How Interpretability Methods Can Learn to Encode Predictions in their Interpretations
Mar 02, 2021
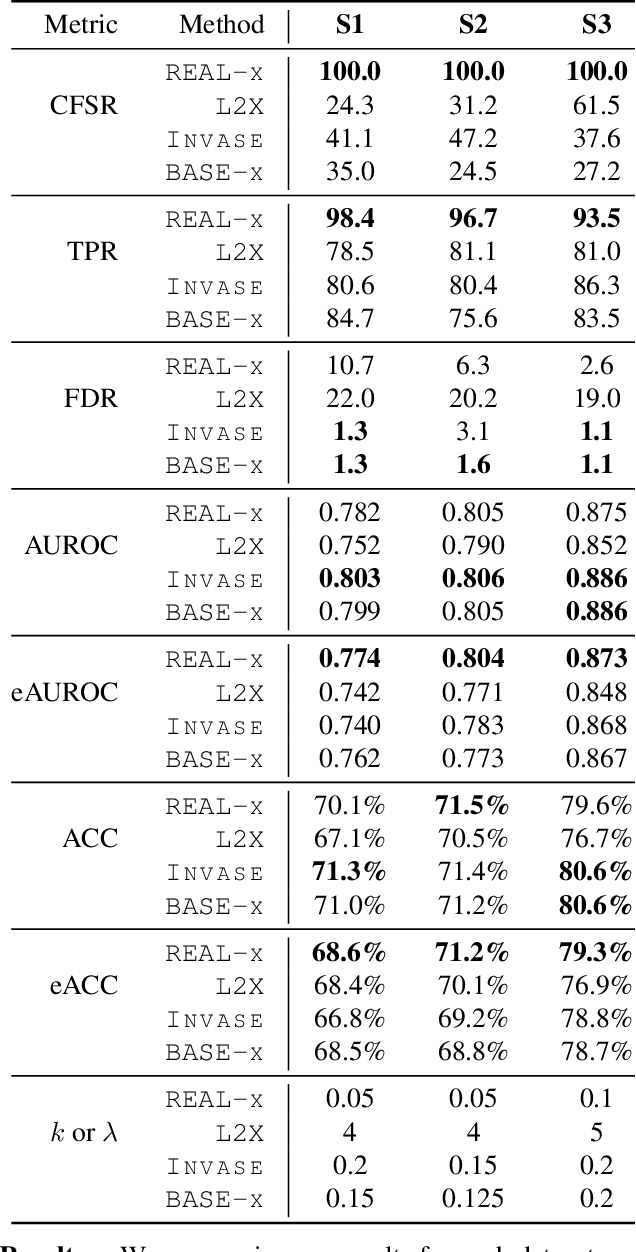
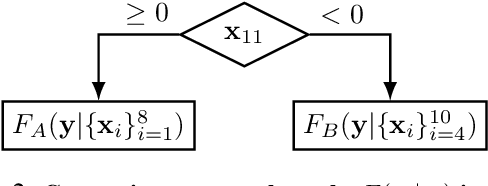
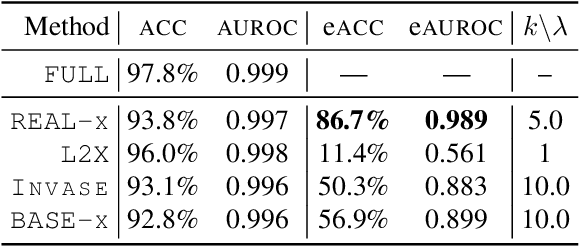
Abstract:While the need for interpretable machine learning has been established, many common approaches are slow, lack fidelity, or hard to evaluate. Amortized explanation methods reduce the cost of providing interpretations by learning a global selector model that returns feature importances for a single instance of data. The selector model is trained to optimize the fidelity of the interpretations, as evaluated by a predictor model for the target. Popular methods learn the selector and predictor model in concert, which we show allows predictions to be encoded within interpretations. We introduce EVAL-X as a method to quantitatively evaluate interpretations and REAL-X as an amortized explanation method, which learn a predictor model that approximates the true data generating distribution given any subset of the input. We show EVAL-X can detect when predictions are encoded in interpretations and show the advantages of REAL-X through quantitative and radiologist evaluation.
* 15 pages, 3 figures, Proceedings of the 24th International Conference on Artificial Intelligence and Statistics (AISTATS) 2021
Deep Direct Likelihood Knockoffs
Jul 31, 2020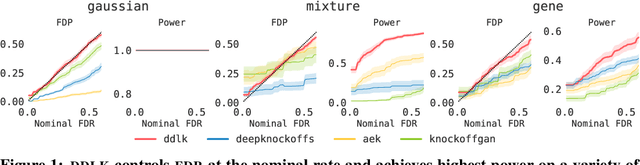



Abstract:Predictive modeling often uses black box machine learning methods, such as deep neural networks, to achieve state-of-the-art performance. In scientific domains, the scientist often wishes to discover which features are actually important for making the predictions. These discoveries may lead to costly follow-up experiments and as such it is important that the error rate on discoveries is not too high. Model-X knockoffs enable important features to be discovered with control of the FDR. However, knockoffs require rich generative models capable of accurately modeling the knockoff features while ensuring they obey the so-called "swap" property. We develop Deep Direct Likelihood Knockoffs (DDLK), which directly minimizes the KL divergence implied by the knockoff swap property. DDLK consists of two stages: it first maximizes the explicit likelihood of the features, then minimizes the KL divergence between the joint distribution of features and knockoffs and any swap between them. To ensure that the generated knockoffs are valid under any possible swap, DDLK uses the Gumbel-Softmax trick to optimize the knockoff generator under the worst-case swap. We find DDLK has higher power than baselines while controlling the false discovery rate on a variety of synthetic and real benchmarks including a task involving a large dataset from one of the epicenters of COVID-19.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge