Mark Malhotra
Towards a Science of Scaling Agent Systems
Dec 17, 2025Abstract:Agents, language model-based systems that are capable of reasoning, planning, and acting are becoming the dominant paradigm for real-world AI applications. Despite this widespread adoption, the principles that determine their performance remain underexplored. We address this by deriving quantitative scaling principles for agent systems. We first formalize a definition for agentic evaluation and characterize scaling laws as the interplay between agent quantity, coordination structure, model capability, and task properties. We evaluate this across four benchmarks: Finance-Agent, BrowseComp-Plus, PlanCraft, and Workbench. With five canonical agent architectures (Single-Agent and four Multi-Agent Systems: Independent, Centralized, Decentralized, Hybrid), instantiated across three LLM families, we perform a controlled evaluation spanning 180 configurations. We derive a predictive model using coordination metrics, that achieves cross-validated R^2=0.524, enabling prediction on unseen task domains. We identify three effects: (1) a tool-coordination trade-off: under fixed computational budgets, tool-heavy tasks suffer disproportionately from multi-agent overhead. (2) a capability saturation: coordination yields diminishing or negative returns once single-agent baselines exceed ~45%. (3) topology-dependent error amplification: independent agents amplify errors 17.2x, while centralized coordination contains this to 4.4x. Centralized coordination improves performance by 80.8% on parallelizable tasks, while decentralized coordination excels on web navigation (+9.2% vs. +0.2%). Yet for sequential reasoning tasks, every multi-agent variants degraded performance by 39-70%. The framework predicts the optimal coordination strategy for 87% of held-out configurations. Out-of-sample validation on GPT-5.2, achieves MAE=0.071 and confirms four of five scaling principles generalize to unseen frontier models.
Smartphone monitoring of smiling as a behavioral proxy of well-being in everyday life
Dec 10, 2025Abstract:Subjective well-being is a cornerstone of individual and societal health, yet its scientific measurement has traditionally relied on self-report methods prone to recall bias and high participant burden. This has left a gap in our understanding of well-being as it is expressed in everyday life. We hypothesized that candid smiles captured during natural smartphone interactions could serve as a scalable, objective behavioral correlate of positive affect. To test this, we analyzed 405,448 video clips passively recorded from 233 consented participants over one week. Using a deep learning model to quantify smile intensity, we identified distinct diurnal and daily patterns. Daily patterns of smile intensity across the week showed strong correlation with national survey data on happiness (r=0.92), and diurnal rhythms documented close correspondence with established results from the day reconstruction method (r=0.80). Higher daily mean smile intensity was significantly associated with more physical activity (Beta coefficient = 0.043, 95% CI [0.001, 0.085]) and greater light exposure (Beta coefficient = 0.038, [0.013, 0.063]), whereas no significant effects were found for smartphone use. These findings suggest that passive smartphone sensing could serve as a powerful, ecologically valid methodology for studying the dynamics of affective behavior and open the door to understanding this behavior at a population scale.
SensorLM: Learning the Language of Wearable Sensors
Jun 10, 2025Abstract:We present SensorLM, a family of sensor-language foundation models that enable wearable sensor data understanding with natural language. Despite its pervasive nature, aligning and interpreting sensor data with language remains challenging due to the lack of paired, richly annotated sensor-text descriptions in uncurated, real-world wearable data. We introduce a hierarchical caption generation pipeline designed to capture statistical, structural, and semantic information from sensor data. This approach enabled the curation of the largest sensor-language dataset to date, comprising over 59.7 million hours of data from more than 103,000 people. Furthermore, SensorLM extends prominent multimodal pretraining architectures (e.g., CLIP, CoCa) and recovers them as specific variants within a generic architecture. Extensive experiments on real-world tasks in human activity analysis and healthcare verify the superior performance of SensorLM over state-of-the-art in zero-shot recognition, few-shot learning, and cross-modal retrieval. SensorLM also demonstrates intriguing capabilities including scaling behaviors, label efficiency, sensor captioning, and zero-shot generalization to unseen tasks.
RADAR: Benchmarking Language Models on Imperfect Tabular Data
Jun 09, 2025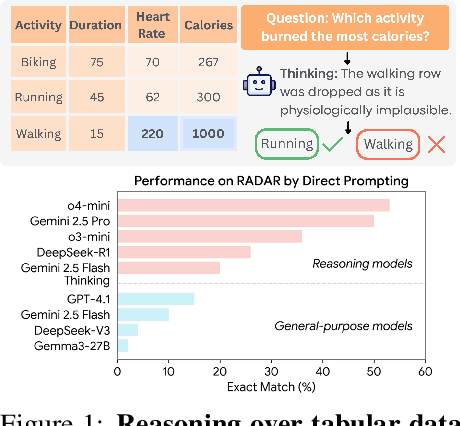
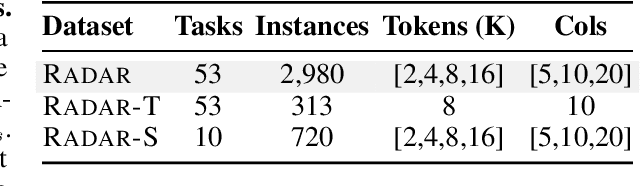
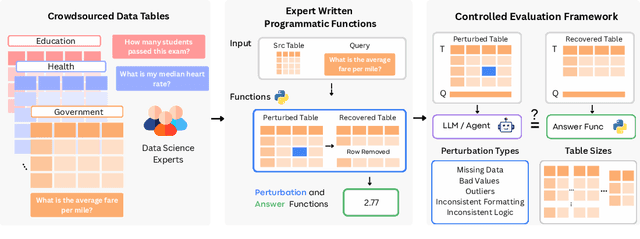
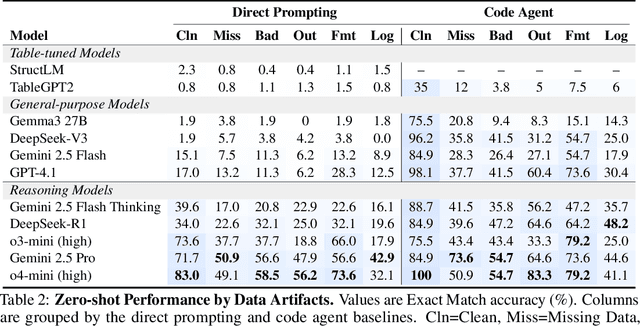
Abstract:Language models (LMs) are increasingly being deployed to perform autonomous data analyses. However, their data awareness -- the ability to recognize, reason over, and appropriately handle data artifacts such as missing values, outliers, and logical inconsistencies -- remains underexplored. These artifacts are especially common in real-world tabular data and, if mishandled, can significantly compromise the validity of analytical conclusions. To address this gap, we present RADAR, a benchmark for systematically evaluating data-aware reasoning on tabular data. We develop a framework to simulate data artifacts via programmatic perturbations to enable targeted evaluation of model behavior. RADAR comprises 2980 table query pairs, grounded in real-world data spanning 9 domains and 5 data artifact types. In addition to evaluating artifact handling, RADAR systematically varies table size to study how reasoning performance holds when increasing table size. Our evaluation reveals that, despite decent performance on tables without data artifacts, frontier models degrade significantly when data artifacts are introduced, exposing critical gaps in their capacity for robust, data-aware analysis. Designed to be flexible and extensible, RADAR supports diverse perturbation types and controllable table sizes, offering a valuable resource for advancing tabular reasoning.
LSM-2: Learning from Incomplete Wearable Sensor Data
Jun 05, 2025



Abstract:Foundation models, a cornerstone of recent advancements in machine learning, have predominantly thrived on complete and well-structured data. Wearable sensor data frequently suffers from significant missingness, posing a substantial challenge for self-supervised learning (SSL) models that typically assume complete data inputs. This paper introduces the second generation of Large Sensor Model (LSM-2) with Adaptive and Inherited Masking (AIM), a novel SSL approach that learns robust representations directly from incomplete data without requiring explicit imputation. AIM's core novelty lies in its use of learnable mask tokens to model both existing ("inherited") and artificially introduced missingness, enabling it to robustly handle fragmented real-world data during inference. Pre-trained on an extensive dataset of 40M hours of day-long multimodal sensor data, our LSM-2 with AIM achieves the best performance across a diverse range of tasks, including classification, regression and generative modeling. Furthermore, LSM-2 with AIM exhibits superior scaling performance, and critically, maintains high performance even under targeted missingness scenarios, reflecting clinically coherent patterns, such as the diagnostic value of nighttime biosignals for hypertension prediction. This makes AIM a more reliable choice for real-world wearable data applications.
A Scalable Framework for Evaluating Health Language Models
Apr 01, 2025Abstract:Large language models (LLMs) have emerged as powerful tools for analyzing complex datasets. Recent studies demonstrate their potential to generate useful, personalized responses when provided with patient-specific health information that encompasses lifestyle, biomarkers, and context. As LLM-driven health applications are increasingly adopted, rigorous and efficient one-sided evaluation methodologies are crucial to ensure response quality across multiple dimensions, including accuracy, personalization and safety. Current evaluation practices for open-ended text responses heavily rely on human experts. This approach introduces human factors and is often cost-prohibitive, labor-intensive, and hinders scalability, especially in complex domains like healthcare where response assessment necessitates domain expertise and considers multifaceted patient data. In this work, we introduce Adaptive Precise Boolean rubrics: an evaluation framework that streamlines human and automated evaluation of open-ended questions by identifying gaps in model responses using a minimal set of targeted rubrics questions. Our approach is based on recent work in more general evaluation settings that contrasts a smaller set of complex evaluation targets with a larger set of more precise, granular targets answerable with simple boolean responses. We validate this approach in metabolic health, a domain encompassing diabetes, cardiovascular disease, and obesity. Our results demonstrate that Adaptive Precise Boolean rubrics yield higher inter-rater agreement among expert and non-expert human evaluators, and in automated assessments, compared to traditional Likert scales, while requiring approximately half the evaluation time of Likert-based methods. This enhanced efficiency, particularly in automated evaluation and non-expert contributions, paves the way for more extensive and cost-effective evaluation of LLMs in health.
Scaling Wearable Foundation Models
Oct 17, 2024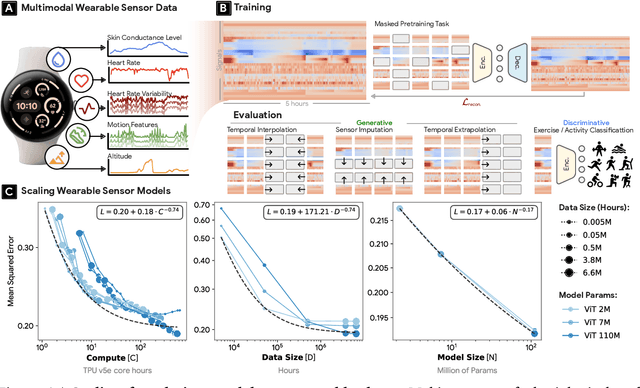
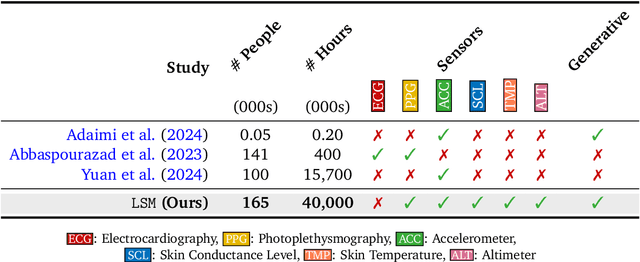


Abstract:Wearable sensors have become ubiquitous thanks to a variety of health tracking features. The resulting continuous and longitudinal measurements from everyday life generate large volumes of data; however, making sense of these observations for scientific and actionable insights is non-trivial. Inspired by the empirical success of generative modeling, where large neural networks learn powerful representations from vast amounts of text, image, video, or audio data, we investigate the scaling properties of sensor foundation models across compute, data, and model size. Using a dataset of up to 40 million hours of in-situ heart rate, heart rate variability, electrodermal activity, accelerometer, skin temperature, and altimeter per-minute data from over 165,000 people, we create LSM, a multimodal foundation model built on the largest wearable-signals dataset with the most extensive range of sensor modalities to date. Our results establish the scaling laws of LSM for tasks such as imputation, interpolation and extrapolation, both across time and sensor modalities. Moreover, we highlight how LSM enables sample-efficient downstream learning for tasks like exercise and activity recognition.
Transforming Wearable Data into Health Insights using Large Language Model Agents
Jun 11, 2024



Abstract:Despite the proliferation of wearable health trackers and the importance of sleep and exercise to health, deriving actionable personalized insights from wearable data remains a challenge because doing so requires non-trivial open-ended analysis of these data. The recent rise of large language model (LLM) agents, which can use tools to reason about and interact with the world, presents a promising opportunity to enable such personalized analysis at scale. Yet, the application of LLM agents in analyzing personal health is still largely untapped. In this paper, we introduce the Personal Health Insights Agent (PHIA), an agent system that leverages state-of-the-art code generation and information retrieval tools to analyze and interpret behavioral health data from wearables. We curate two benchmark question-answering datasets of over 4000 health insights questions. Based on 650 hours of human and expert evaluation we find that PHIA can accurately address over 84% of factual numerical questions and more than 83% of crowd-sourced open-ended questions. This work has implications for advancing behavioral health across the population, potentially enabling individuals to interpret their own wearable data, and paving the way for a new era of accessible, personalized wellness regimens that are informed by data-driven insights.
Towards a Personal Health Large Language Model
Jun 10, 2024



Abstract:In health, most large language model (LLM) research has focused on clinical tasks. However, mobile and wearable devices, which are rarely integrated into such tasks, provide rich, longitudinal data for personal health monitoring. Here we present Personal Health Large Language Model (PH-LLM), fine-tuned from Gemini for understanding and reasoning over numerical time-series personal health data. We created and curated three datasets that test 1) production of personalized insights and recommendations from sleep patterns, physical activity, and physiological responses, 2) expert domain knowledge, and 3) prediction of self-reported sleep outcomes. For the first task we designed 857 case studies in collaboration with domain experts to assess real-world scenarios in sleep and fitness. Through comprehensive evaluation of domain-specific rubrics, we observed that Gemini Ultra 1.0 and PH-LLM are not statistically different from expert performance in fitness and, while experts remain superior for sleep, fine-tuning PH-LLM provided significant improvements in using relevant domain knowledge and personalizing information for sleep insights. We evaluated PH-LLM domain knowledge using multiple choice sleep medicine and fitness examinations. PH-LLM achieved 79% on sleep and 88% on fitness, exceeding average scores from a sample of human experts. Finally, we trained PH-LLM to predict self-reported sleep quality outcomes from textual and multimodal encoding representations of wearable data, and demonstrate that multimodal encoding is required to match performance of specialized discriminative models. Although further development and evaluation are necessary in the safety-critical personal health domain, these results demonstrate both the broad knowledge and capabilities of Gemini models and the benefit of contextualizing physiological data for personal health applications as done with PH-LLM.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge