Lauge Sørensen
CARRNN: A Continuous Autoregressive Recurrent Neural Network for Deep Representation Learning from Sporadic Temporal Data
Apr 08, 2021
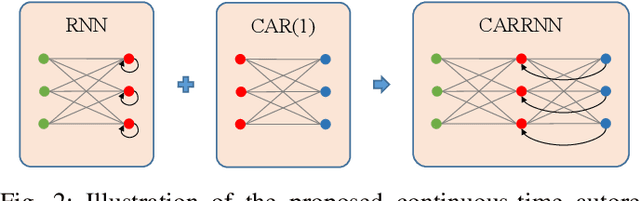
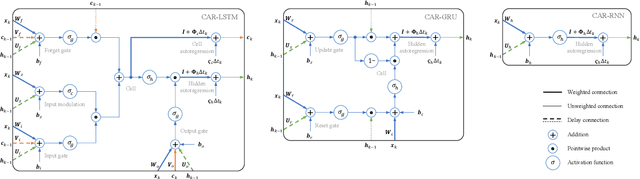
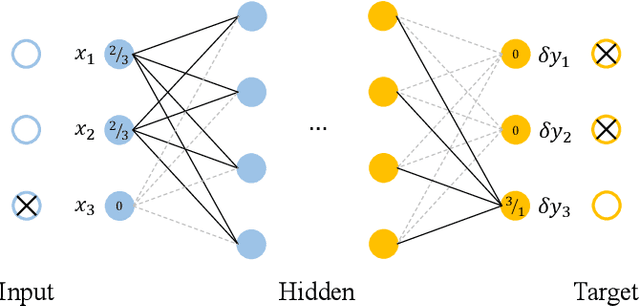
Abstract:Learning temporal patterns from multivariate longitudinal data is challenging especially in cases when data is sporadic, as often seen in, e.g., healthcare applications where the data can suffer from irregularity and asynchronicity as the time between consecutive data points can vary across features and samples, hindering the application of existing deep learning models that are constructed for complete, evenly spaced data with fixed sequence lengths. In this paper, a novel deep learning-based model is developed for modeling multiple temporal features in sporadic data using an integrated deep learning architecture based on a recurrent neural network (RNN) unit and a continuous-time autoregressive (CAR) model. The proposed model, called CARRNN, uses a generalized discrete-time autoregressive model that is trainable end-to-end using neural networks modulated by time lags to describe the changes caused by the irregularity and asynchronicity. It is applied to multivariate time-series regression tasks using data provided for Alzheimer's disease progression modeling and intensive care unit (ICU) mortality rate prediction, where the proposed model based on a gated recurrent unit (GRU) achieves the lowest prediction errors among the proposed RNN-based models and state-of-the-art methods using GRUs and long short-term memory (LSTM) networks in their architecture.
Multi-Domain Adaptation in Brain MRI through Paired Consistency and Adversarial Learning
Sep 17, 2019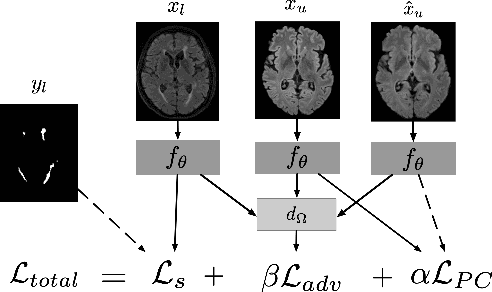
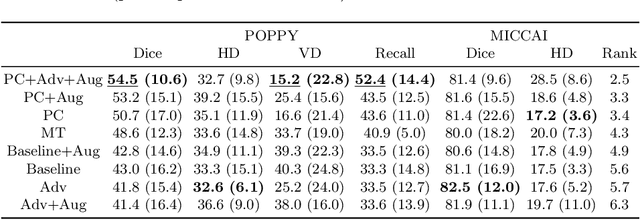
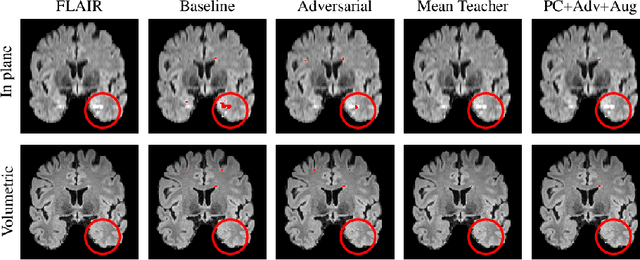
Abstract:Supervised learning algorithms trained on medical images will often fail to generalize across changes in acquisition parameters. Recent work in domain adaptation addresses this challenge and successfully leverages labeled data in a source domain to perform well on an unlabeled target domain. Inspired by recent work in semi-supervised learning we introduce a novel method to adapt from one source domain to $n$ target domains (as long as there is paired data covering all domains). Our multi-domain adaptation method utilises a consistency loss combined with adversarial learning. We provide results on white matter lesion hyperintensity segmentation from brain MRIs using the MICCAI 2017 challenge data as the source domain and two target domains. The proposed method significantly outperforms other domain adaptation baselines.
Knowledge distillation for semi-supervised domain adaptation
Aug 16, 2019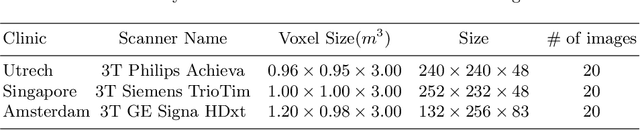
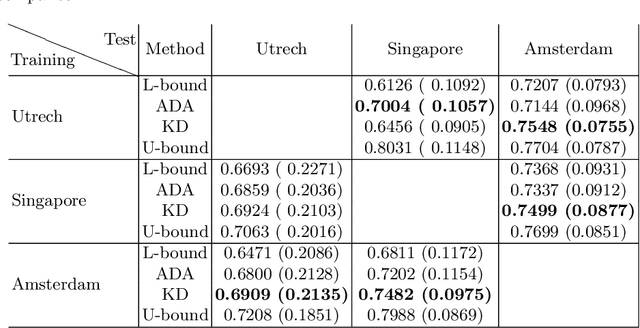
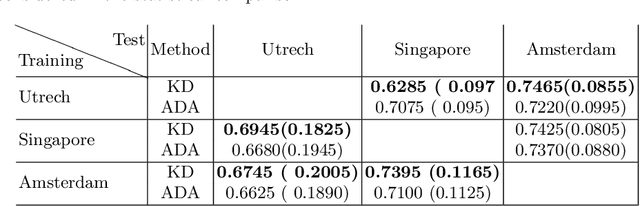
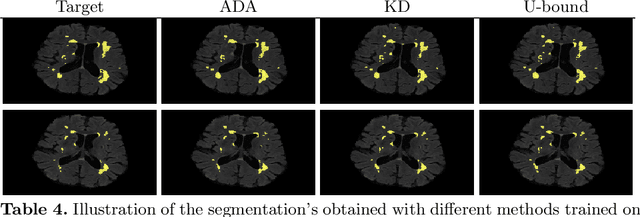
Abstract:In the absence of sufficient data variation (e.g., scanner and protocol variability) in annotated data, deep neural networks (DNNs) tend to overfit during training. As a result, their performance is significantly lower on data from unseen sources compared to the performance on data from the same source as the training data. Semi-supervised domain adaptation methods can alleviate this problem by tuning networks to new target domains without the need for annotated data from these domains. Adversarial domain adaptation (ADA) methods are a popular choice that aim to train networks in such a way that the features generated are domain agnostic. However, these methods require careful dataset-specific selection of hyperparameters such as the complexity of the discriminator in order to achieve a reasonable performance. We propose to use knowledge distillation (KD) -- an efficient way of transferring knowledge between different DNNs -- for semi-supervised domain adaption of DNNs. It does not require dataset-specific hyperparameter tuning, making it generally applicable. The proposed method is compared to ADA for segmentation of white matter hyperintensities (WMH) in magnetic resonance imaging (MRI) scans generated by scanners that are not a part of the training set. Compared with both the baseline DNN (trained on source domain only and without any adaption to target domain) and with using ADA for semi-supervised domain adaptation, the proposed method achieves significantly higher WMH dice scores.
Robust parametric modeling of Alzheimer's disease progression
Aug 14, 2019
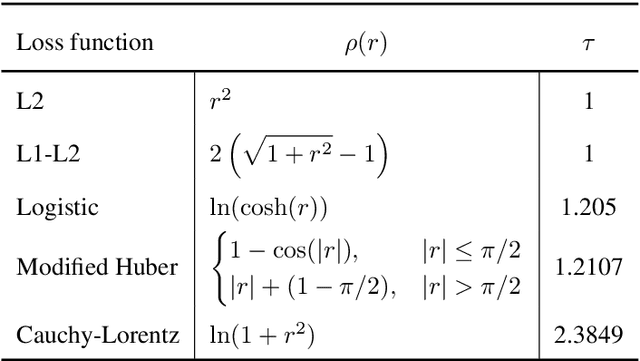
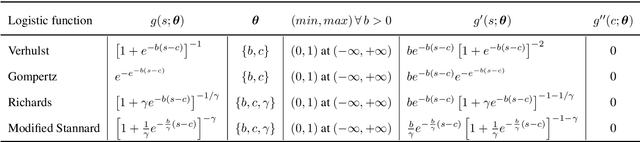


Abstract:Quantitative characterization of disease progression using longitudinal data can provide long-term predictions for the pathological stages of individuals. This work studies robust modeling of Alzheimer's disease progression using parametric methods. The proposed method linearly maps individual's chronological age to a disease progression score (DPS) and robustly fits a constrained generalized logistic function to the longitudinal dynamic of a biomarker as a function of the DPS using M-estimation. Robustness of the estimates is quantified using bootstrapping via Monte Carlo resampling, and the inflection points are used to temporally order the modeled biomarkers in the disease course. Moreover, kernel density estimation is applied to the obtained DPSs for clinical status prediction using a Bayesian classifier. Different M-estimators and logistic functions, including a new generalized type proposed in this study called modified Stannard, are evaluated on the ADNI database for robust modeling of volumetric MRI and PET biomarkers, as well as neuropsychological tests. The results show that the modified Stannard function fitted using the modified Huber loss achieves the best modeling performance with a mean of median absolute errors (MMAE) of 0.059 across all biomarkers and bootstraps. In addition, applied to the ADNI test set, this model achieves a multi-class area under the ROC curve (MAUC) of 0.87 in clinical status prediction, and it significantly outperforms an analogous state-of-the-art method with a biomarker modeling MMAE of 0.059 vs. 0.061 (p < 0.001). Finally, the experiments show that the proposed model, trained using abundant ADNI data, generalizes well to data from the independent NACC database, where both modeling and diagnostic performance are significantly improved (p < 0.001) compared with using a model trained using relatively sparse NACC data.
Training recurrent neural networks robust to incomplete data: application to Alzheimer's disease progression modeling
Mar 17, 2019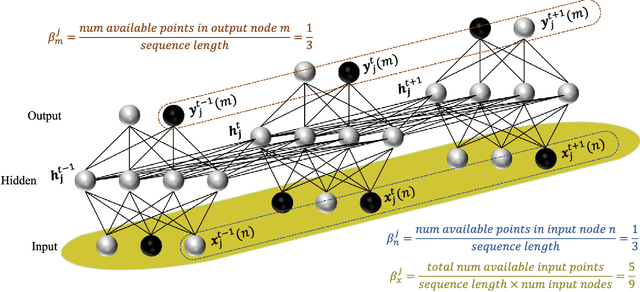
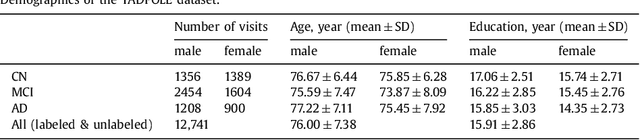
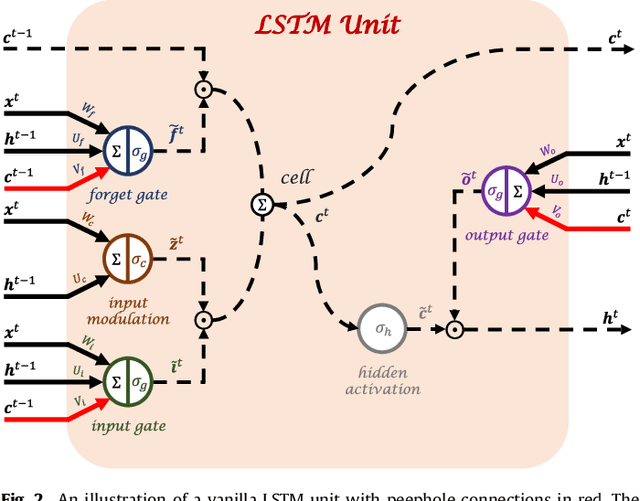
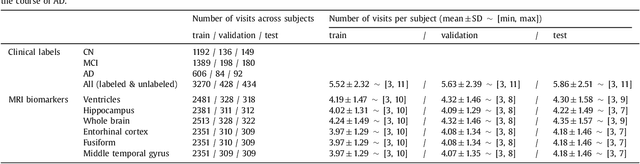
Abstract:Disease progression modeling (DPM) using longitudinal data is a challenging machine learning task. Existing DPM algorithms neglect temporal dependencies among measurements, make parametric assumptions about biomarker trajectories, do not model multiple biomarkers jointly, and need an alignment of subjects' trajectories. In this paper, recurrent neural networks (RNNs) are utilized to address these issues. However, in many cases, longitudinal cohorts contain incomplete data, which hinders the application of standard RNNs and requires a pre-processing step such as imputation of the missing values. Instead, we propose a generalized training rule for the most widely used RNN architecture, long short-term memory (LSTM) networks, that can handle both missing predictor and target values. The proposed LSTM algorithm is applied to model the progression of Alzheimer's disease (AD) using six volumetric magnetic resonance imaging (MRI) biomarkers, i.e., volumes of ventricles, hippocampus, whole brain, fusiform, middle temporal gyrus, and entorhinal cortex, and it is compared to standard LSTM networks with data imputation and a parametric, regression-based DPM method. The results show that the proposed algorithm achieves a significantly lower mean absolute error (MAE) than the alternatives with p < 0.05 using Wilcoxon signed rank test in predicting values of almost all of the MRI biomarkers. Moreover, a linear discriminant analysis (LDA) classifier applied to the predicted biomarker values produces a significantly larger AUC of 0.90 vs. at most 0.84 with p < 0.001 using McNemar's test for clinical diagnosis of AD. Inspection of MAE curves as a function of the amount of missing data reveals that the proposed LSTM algorithm achieves the best performance up until more than 74% missing values. Finally, it is illustrated how the method can successfully be applied to data with varying time intervals.
* arXiv admin note: substantial text overlap with arXiv:1808.05500
PADDIT: Probabilistic Augmentation of Data using Diffeomorphic Image Transformation
Oct 03, 2018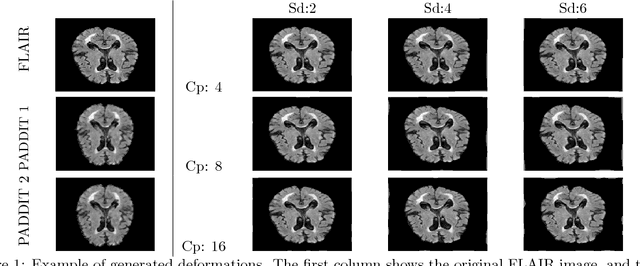
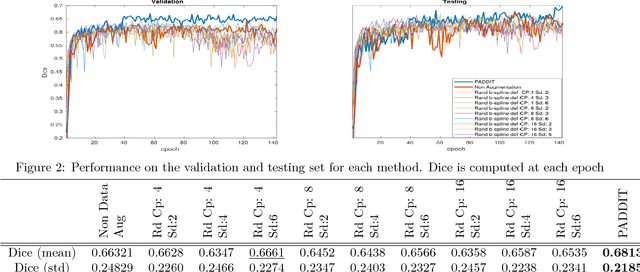
Abstract:For proper generalization performance of convolutional neural networks (CNNs) in medical image segmentation, the learnt features should be invariant under particular non-linear shape variations of the input. To induce invariance in CNNs to such transformations, we propose Probabilistic Augmentation of Data using Diffeomorphic Image Transformation (PADDIT) -- a systematic framework for generating realistic transformations that can be used to augment data for training CNNs. We show that CNNs trained with PADDIT outperforms CNNs trained without augmentation and with generic augmentation in segmenting white matter hyperintensities from T1 and FLAIR brain MRI scans.
Simultaneous synthesis of FLAIR and segmentation of white matter hypointensities from T1 MRIs
Aug 20, 2018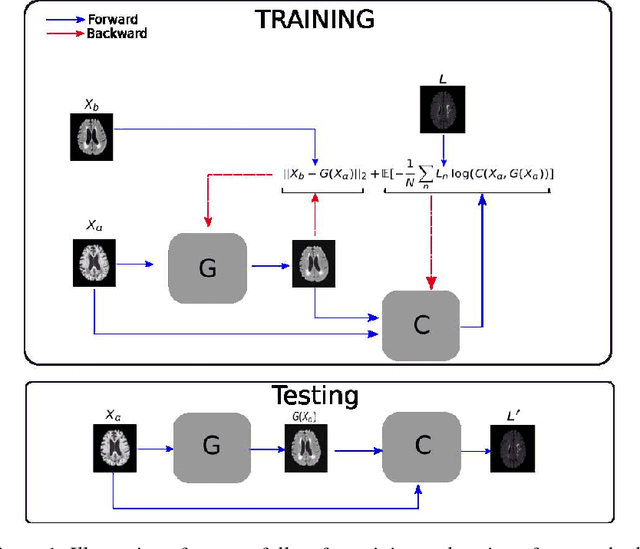
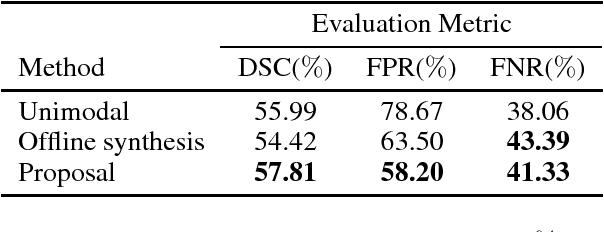
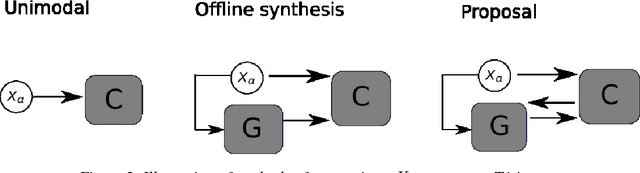
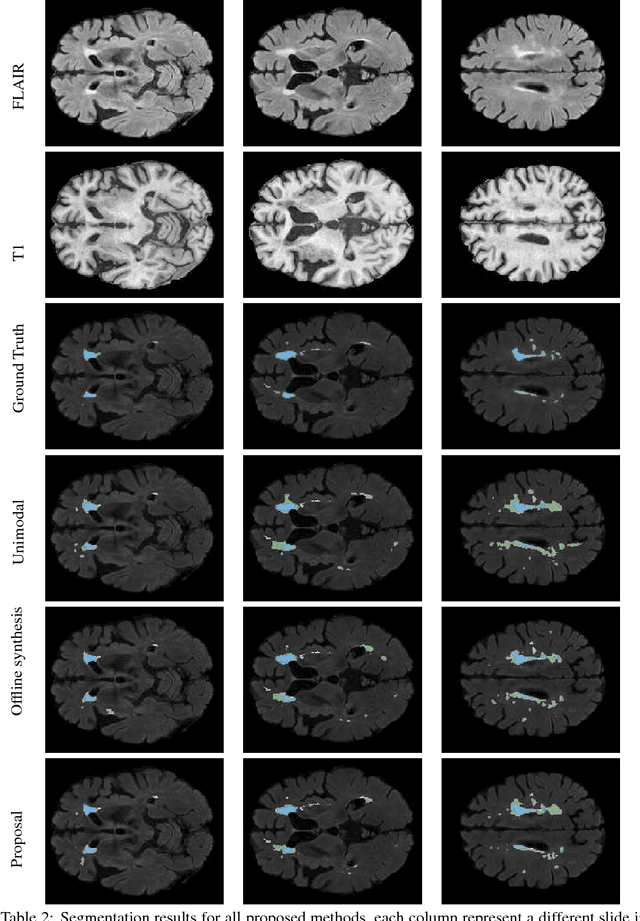
Abstract:Segmenting vascular pathologies such as white matter lesions in Brain magnetic resonance images (MRIs) require acquisition of multiple sequences such as T1-weighted (T1-w) --on which lesions appear hypointense-- and fluid attenuated inversion recovery (FLAIR) sequence --where lesions appear hyperintense--. However, most of the existing retrospective datasets do not consist of FLAIR sequences. Existing missing modality imputation methods separate the process of imputation, and the process of segmentation. In this paper, we propose a method to link both modality imputation and segmentation using convolutional neural networks. We show that by jointly optimizing the imputation network and the segmentation network, the method not only produces more realistic synthetic FLAIR images from T1-w images, but also improves the segmentation of WMH from T1-w images only.
Robust training of recurrent neural networks to handle missing data for disease progression modeling
Aug 16, 2018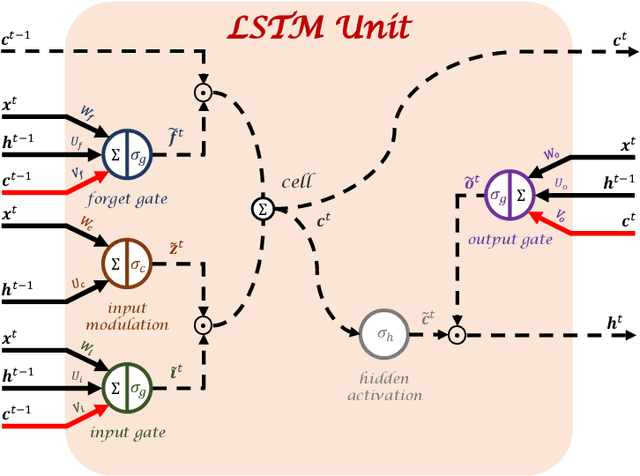
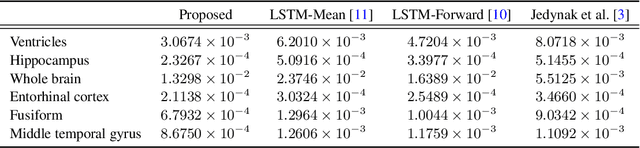
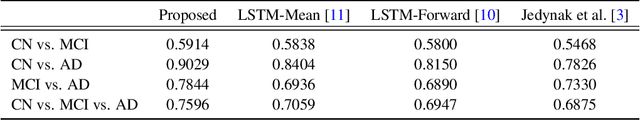



Abstract:Disease progression modeling (DPM) using longitudinal data is a challenging task in machine learning for healthcare that can provide clinicians with better tools for diagnosis and monitoring of disease. Existing DPM algorithms neglect temporal dependencies among measurements and make parametric assumptions about biomarker trajectories. In addition, they do not model multiple biomarkers jointly and need to align subjects' trajectories. In this paper, recurrent neural networks (RNNs) are utilized to address these issues. However, in many cases, longitudinal cohorts contain incomplete data, which hinders the application of standard RNNs and requires a pre-processing step such as imputation of the missing values. We, therefore, propose a generalized training rule for the most widely used RNN architecture, long short-term memory (LSTM) networks, that can handle missing values in both target and predictor variables. This algorithm is applied for modeling the progression of Alzheimer's disease (AD) using magnetic resonance imaging (MRI) biomarkers. The results show that the proposed LSTM algorithm achieves a lower mean absolute error for prediction of measurements across all considered MRI biomarkers compared to using standard LSTM networks with data imputation or using a regression-based DPM method. Moreover, applying linear discriminant analysis to the biomarkers' values predicted by the proposed algorithm results in a larger area under the receiver operating characteristic curve (AUC) for clinical diagnosis of AD compared to the same alternatives, and the AUC is comparable to state-of-the-art AUCs from a recent cross-sectional medical image classification challenge. This paper shows that built-in handling of missing values in LSTM network training paves the way for application of RNNs in disease progression modeling.
Transfer learning for multi-center classification of chronic obstructive pulmonary disease
Nov 23, 2017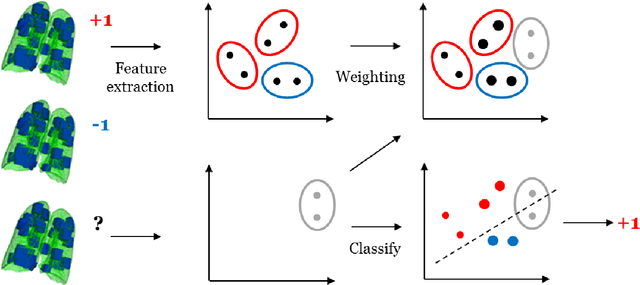
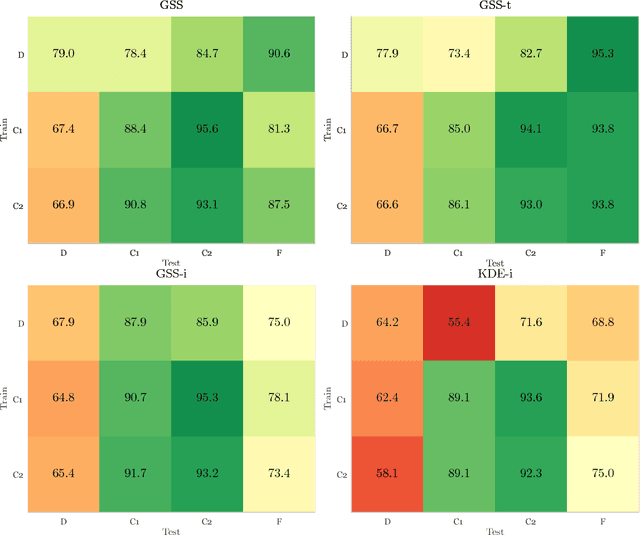
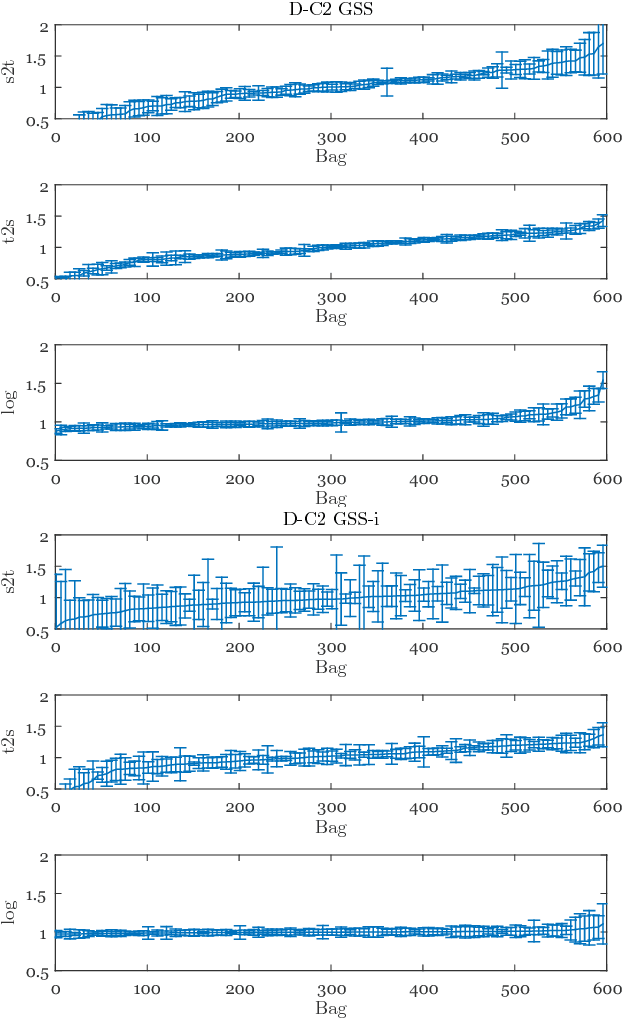
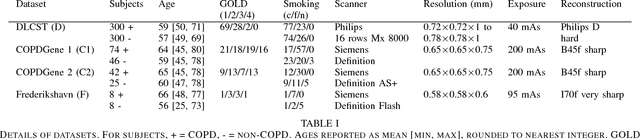
Abstract:Chronic obstructive pulmonary disease (COPD) is a lung disease which can be quantified using chest computed tomography (CT) scans. Recent studies have shown that COPD can be automatically diagnosed using weakly supervised learning of intensity and texture distributions. However, up till now such classifiers have only been evaluated on scans from a single domain, and it is unclear whether they would generalize across domains, such as different scanners or scanning protocols. To address this problem, we investigate classification of COPD in a multi-center dataset with a total of 803 scans from three different centers, four different scanners, with heterogenous subject distributions. Our method is based on Gaussian texture features, and a weighted logistic classifier, which increases the weights of samples similar to the test data. We show that Gaussian texture features outperform intensity features previously used in multi-center classification tasks. We also show that a weighting strategy based on a classifier that is trained to discriminate between scans from different domains, can further improve the results. To encourage further research into transfer learning methods for classification of COPD, upon acceptance of the paper we will release two feature datasets used in this study on http://bigr.nl/research/projects/copd
A Statistical Model for Simultaneous Template Estimation, Bias Correction, and Registration of 3D Brain Images
May 01, 2017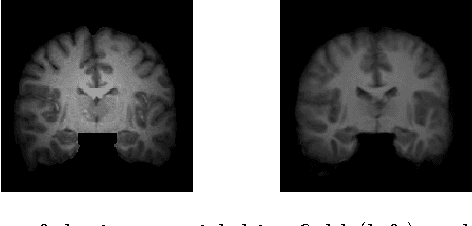
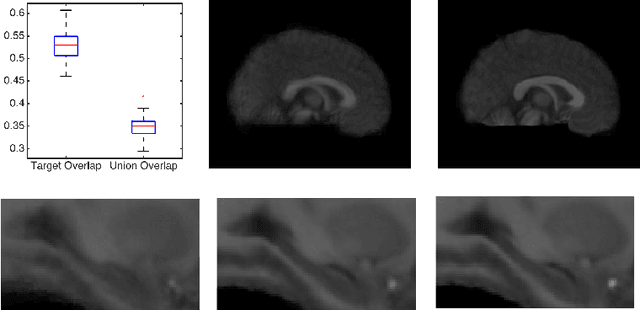
Abstract:Template estimation plays a crucial role in computational anatomy since it provides reference frames for performing statistical analysis of the underlying anatomical population variability. While building models for template estimation, variability in sites and image acquisition protocols need to be accounted for. To account for such variability, we propose a generative template estimation model that makes simultaneous inference of both bias fields in individual images, deformations for image registration, and variance hyperparameters. In contrast, existing maximum a posterori based methods need to rely on either bias-invariant similarity measures or robust image normalization. Results on synthetic and real brain MRI images demonstrate the capability of the model to capture heterogeneity in intensities and provide a reliable template estimation from registration.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge