Kaimin Song
OphNet: A Large-Scale Video Benchmark for Ophthalmic Surgical Workflow Understanding
Jun 12, 2024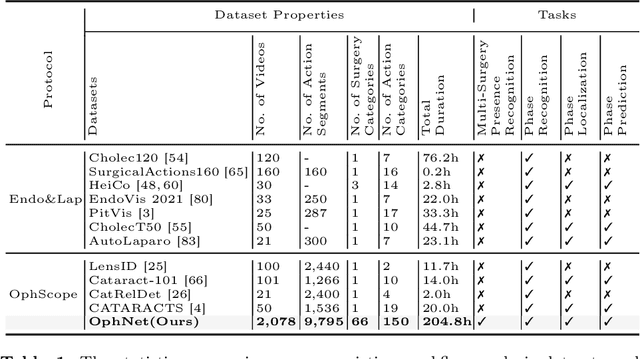
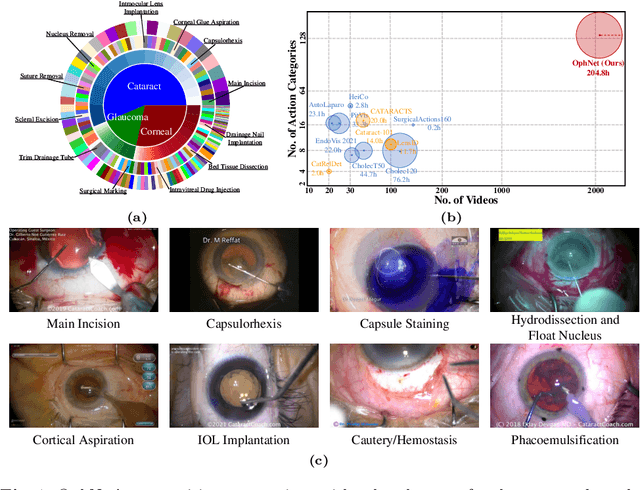

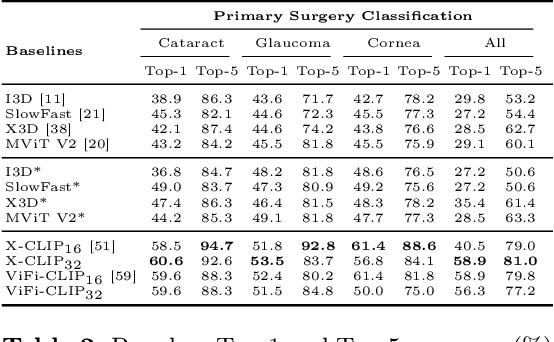
Abstract:Surgical scene perception via videos are critical for advancing robotic surgery, telesurgery, and AI-assisted surgery, particularly in ophthalmology. However, the scarcity of diverse and richly annotated video datasets has hindered the development of intelligent systems for surgical workflow analysis. Existing datasets for surgical workflow analysis, which typically face challenges such as small scale, a lack of diversity in surgery and phase categories, and the absence of time-localized annotations, limit the requirements for action understanding and model generalization validation in complex and diverse real-world surgical scenarios. To address this gap, we introduce OphNet, a large-scale, expert-annotated video benchmark for ophthalmic surgical workflow understanding. OphNet features: 1) A diverse collection of 2,278 surgical videos spanning 66 types of cataract, glaucoma, and corneal surgeries, with detailed annotations for 102 unique surgical phases and 150 granular operations; 2) It offers sequential and hierarchical annotations for each surgery, phase, and operation, enabling comprehensive understanding and improved interpretability; 3) Moreover, OphNet provides time-localized annotations, facilitating temporal localization and prediction tasks within surgical workflows. With approximately 205 hours of surgical videos, OphNet is about 20 times larger than the largest existing surgical workflow analysis benchmark. Our dataset and code have been made available at: \url{https://github.com/minghu0830/OphNet-benchmark}.
Towards Novel Class Discovery: A Study in Novel Skin Lesions Clustering
Sep 28, 2023Abstract:Existing deep learning models have achieved promising performance in recognizing skin diseases from dermoscopic images. However, these models can only recognize samples from predefined categories, when they are deployed in the clinic, data from new unknown categories are constantly emerging. Therefore, it is crucial to automatically discover and identify new semantic categories from new data. In this paper, we propose a new novel class discovery framework for automatically discovering new semantic classes from dermoscopy image datasets based on the knowledge of known classes. Specifically, we first use contrastive learning to learn a robust and unbiased feature representation based on all data from known and unknown categories. We then propose an uncertainty-aware multi-view cross pseudo-supervision strategy, which is trained jointly on all categories of data using pseudo labels generated by a self-labeling strategy. Finally, we further refine the pseudo label by aggregating neighborhood information through local sample similarity to improve the clustering performance of the model for unknown categories. We conducted extensive experiments on the dermatology dataset ISIC 2019, and the experimental results show that our approach can effectively leverage knowledge from known categories to discover new semantic categories. We also further validated the effectiveness of the different modules through extensive ablation experiments. Our code will be released soon.
3D Matting: A Benchmark Study on Soft Segmentation Method for Pulmonary Nodules Applied in Computed Tomography
Oct 11, 2022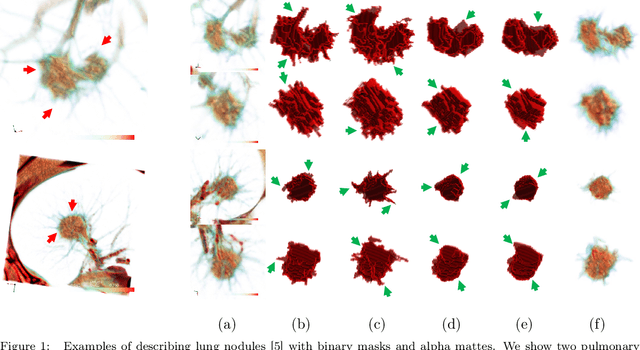
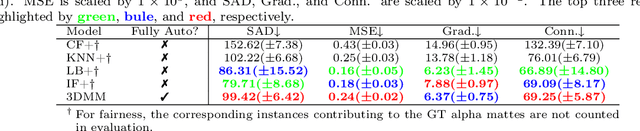
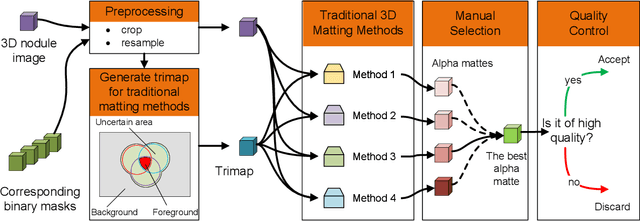
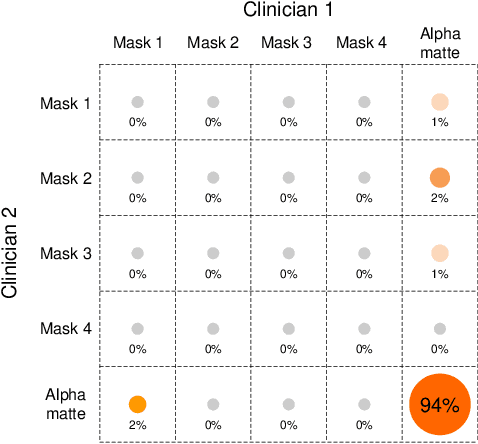
Abstract:Usually, lesions are not isolated but are associated with the surrounding tissues. For example, the growth of a tumour can depend on or infiltrate into the surrounding tissues. Due to the pathological nature of the lesions, it is challenging to distinguish their boundaries in medical imaging. However, these uncertain regions may contain diagnostic information. Therefore, the simple binarization of lesions by traditional binary segmentation can result in the loss of diagnostic information. In this work, we introduce the image matting into the 3D scenes and use the alpha matte, i.e., a soft mask, to describe lesions in a 3D medical image. The traditional soft mask acted as a training trick to compensate for the easily mislabelled or under-labelled ambiguous regions. In contrast, 3D matting uses soft segmentation to characterize the uncertain regions more finely, which means that it retains more structural information for subsequent diagnosis and treatment. The current study of image matting methods in 3D is limited. To address this issue, we conduct a comprehensive study of 3D matting, including both traditional and deep-learning-based methods. We adapt four state-of-the-art 2D image matting algorithms to 3D scenes and further customize the methods for CT images to calibrate the alpha matte with the radiodensity. Moreover, we propose the first end-to-end deep 3D matting network and implement a solid 3D medical image matting benchmark. Its efficient counterparts are also proposed to achieve a good performance-computation balance. Furthermore, there is no high-quality annotated dataset related to 3D matting, slowing down the development of data-driven deep-learning-based methods. To address this issue, we construct the first 3D medical matting dataset. The validity of the dataset was verified through clinicians' assessments and downstream experiments.
3D Matting: A Soft Segmentation Method Applied in Computed Tomography
Sep 16, 2022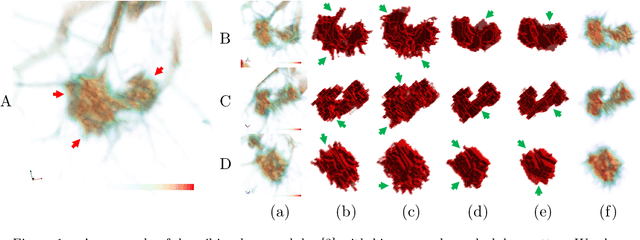

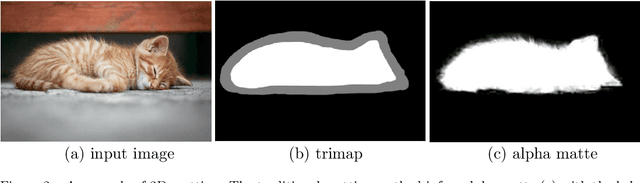
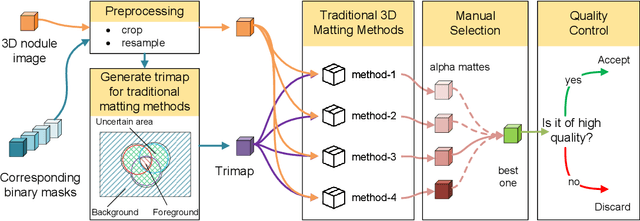
Abstract:Three-dimensional (3D) images, such as CT, MRI, and PET, are common in medical imaging applications and important in clinical diagnosis. Semantic ambiguity is a typical feature of many medical image labels. It can be caused by many factors, such as the imaging properties, pathological anatomy, and the weak representation of the binary masks, which brings challenges to accurate 3D segmentation. In 2D medical images, using soft masks instead of binary masks generated by image matting to characterize lesions can provide rich semantic information, describe the structural characteristics of lesions more comprehensively, and thus benefit the subsequent diagnoses and analyses. In this work, we introduce image matting into the 3D scenes to describe the lesions in 3D medical images. The study of image matting in 3D modality is limited, and there is no high-quality annotated dataset related to 3D matting, therefore slowing down the development of data-driven deep-learning-based methods. To address this issue, we constructed the first 3D medical matting dataset and convincingly verified the validity of the dataset through quality control and downstream experiments in lung nodules classification. We then adapt the four selected state-of-the-art 2D image matting algorithms to 3D scenes and further customize the methods for CT images. Also, we propose the first end-to-end deep 3D matting network and implement a solid 3D medical image matting benchmark, which will be released to encourage further research.
Unsupervised Domain Adaptation for Retinal Vessel Segmentation with Adversarial Learning and Transfer Normalization
Aug 04, 2021
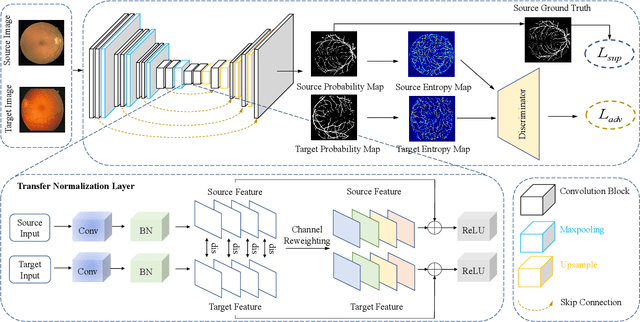
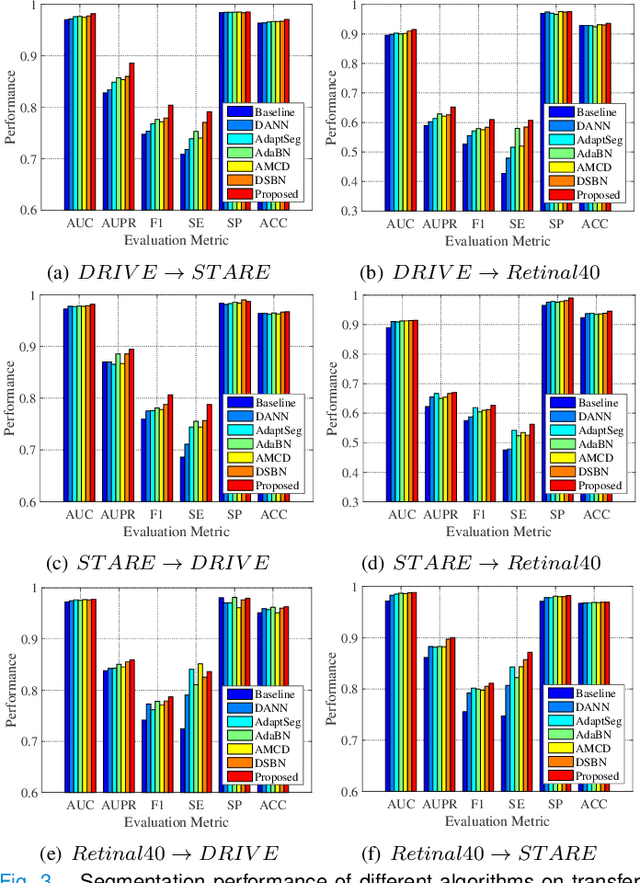

Abstract:Retinal vessel segmentation plays a key role in computer-aided screening, diagnosis, and treatment of various cardiovascular and ophthalmic diseases. Recently, deep learning-based retinal vessel segmentation algorithms have achieved remarkable performance. However, due to the domain shift problem, the performance of these algorithms often degrades when they are applied to new data that is different from the training data. Manually labeling new data for each test domain is often a time-consuming and laborious task. In this work, we explore unsupervised domain adaptation in retinal vessel segmentation by using entropy-based adversarial learning and transfer normalization layer to train a segmentation network, which generalizes well across domains and requires no annotation of the target domain. Specifically, first, an entropy-based adversarial learning strategy is developed to reduce the distribution discrepancy between the source and target domains while also achieving the objective of entropy minimization on the target domain. In addition, a new transfer normalization layer is proposed to further boost the transferability of the deep network. It normalizes the features of each domain separately to compensate for the domain distribution gap. Besides, it also adaptively selects those feature channels that are more transferable between domains, thus further enhancing the generalization performance of the network. We conducted extensive experiments on three regular fundus image datasets and an ultra-widefield fundus image dataset, and the results show that our approach yields significant performance gains compared to other state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge