Juan Helen Zhou
Standardized Evaluation of Automatic Methods for Perivascular Spaces Segmentation in MRI -- MICCAI 2024 Challenge Results
Dec 20, 2025
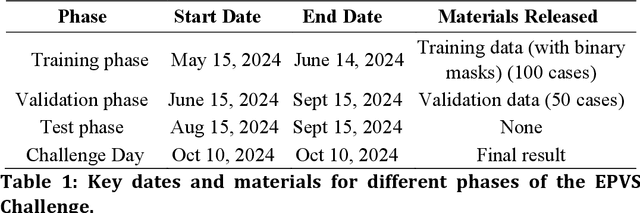
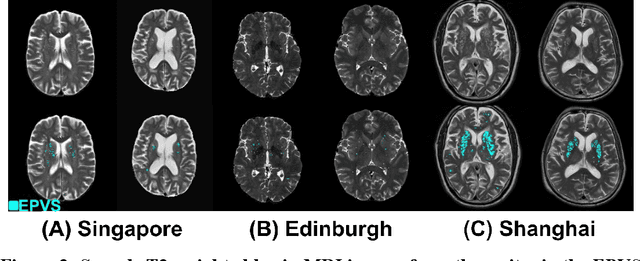
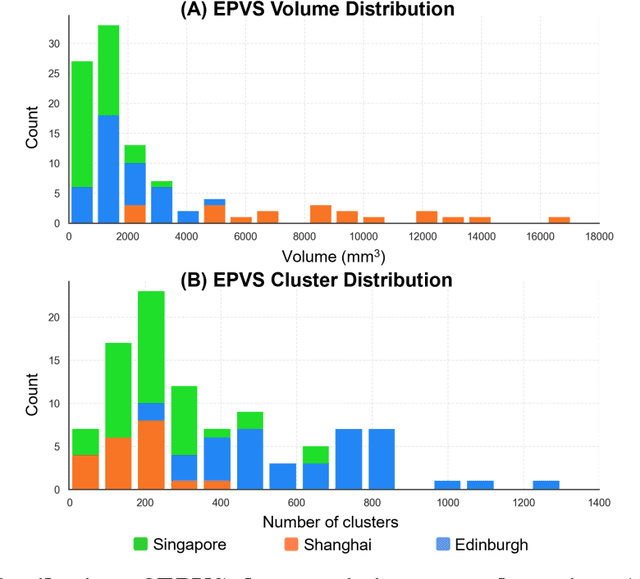
Abstract:Perivascular spaces (PVS), when abnormally enlarged and visible in magnetic resonance imaging (MRI) structural sequences, are important imaging markers of cerebral small vessel disease and potential indicators of neurodegenerative conditions. Despite their clinical significance, automatic enlarged PVS (EPVS) segmentation remains challenging due to their small size, variable morphology, similarity with other pathological features, and limited annotated datasets. This paper presents the EPVS Challenge organized at MICCAI 2024, which aims to advance the development of automated algorithms for EPVS segmentation across multi-site data. We provided a diverse dataset comprising 100 training, 50 validation, and 50 testing scans collected from multiple international sites (UK, Singapore, and China) with varying MRI protocols and demographics. All annotations followed the STRIVE protocol to ensure standardized ground truth and covered the full brain parenchyma. Seven teams completed the full challenge, implementing various deep learning approaches primarily based on U-Net architectures with innovations in multi-modal processing, ensemble strategies, and transformer-based components. Performance was evaluated using dice similarity coefficient, absolute volume difference, recall, and precision metrics. The winning method employed MedNeXt architecture with a dual 2D/3D strategy for handling varying slice thicknesses. The top solutions showed relatively good performance on test data from seen datasets, but significant degradation of performance was observed on the previously unseen Shanghai cohort, highlighting cross-site generalization challenges due to domain shift. This challenge establishes an important benchmark for EPVS segmentation methods and underscores the need for the continued development of robust algorithms that can generalize in diverse clinical settings.
Is an Ultra Large Natural Image-Based Foundation Model Superior to a Retina-Specific Model for Detecting Ocular and Systemic Diseases?
Feb 10, 2025Abstract:The advent of foundation models (FMs) is transforming medical domain. In ophthalmology, RETFound, a retina-specific FM pre-trained sequentially on 1.4 million natural images and 1.6 million retinal images, has demonstrated high adaptability across clinical applications. Conversely, DINOv2, a general-purpose vision FM pre-trained on 142 million natural images, has shown promise in non-medical domains. However, its applicability to clinical tasks remains underexplored. To address this, we conducted head-to-head evaluations by fine-tuning RETFound and three DINOv2 models (large, base, small) for ocular disease detection and systemic disease prediction tasks, across eight standardized open-source ocular datasets, as well as the Moorfields AlzEye and the UK Biobank datasets. DINOv2-large model outperformed RETFound in detecting diabetic retinopathy (AUROC=0.850-0.952 vs 0.823-0.944, across three datasets, all P<=0.007) and multi-class eye diseases (AUROC=0.892 vs. 0.846, P<0.001). In glaucoma, DINOv2-base model outperformed RETFound (AUROC=0.958 vs 0.940, P<0.001). Conversely, RETFound achieved superior performance over all DINOv2 models in predicting heart failure, myocardial infarction, and ischaemic stroke (AUROC=0.732-0.796 vs 0.663-0.771, all P<0.001). These trends persisted even with 10% of the fine-tuning data. These findings showcase the distinct scenarios where general-purpose and domain-specific FMs excel, highlighting the importance of aligning FM selection with task-specific requirements to optimise clinical performance.
Brain-JEPA: Brain Dynamics Foundation Model with Gradient Positioning and Spatiotemporal Masking
Sep 28, 2024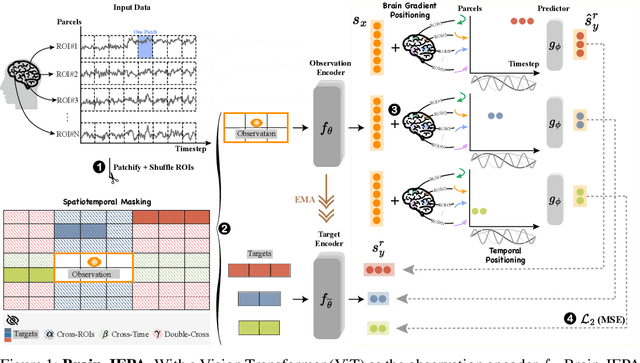
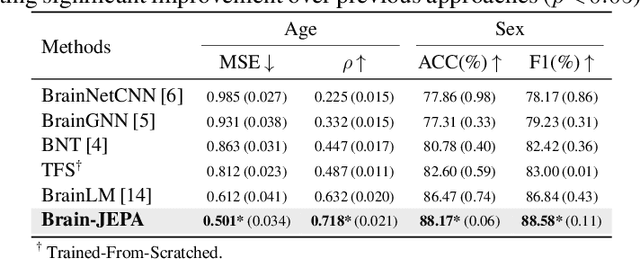
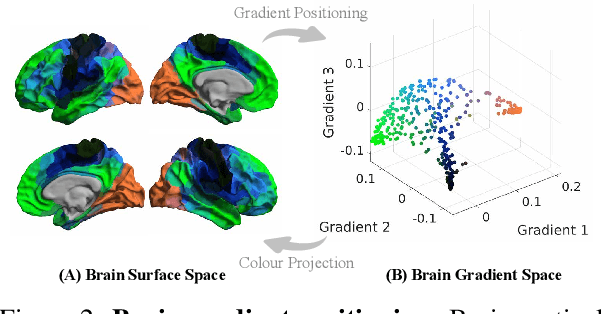
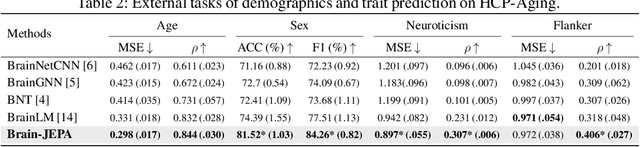
Abstract:We introduce Brain-JEPA, a brain dynamics foundation model with the Joint-Embedding Predictive Architecture (JEPA). This pioneering model achieves state-of-the-art performance in demographic prediction, disease diagnosis/prognosis, and trait prediction through fine-tuning. Furthermore, it excels in off-the-shelf evaluations (e.g., linear probing) and demonstrates superior generalizability across different ethnic groups, surpassing the previous large model for brain activity significantly. Brain-JEPA incorporates two innovative techniques: Brain Gradient Positioning and Spatiotemporal Masking. Brain Gradient Positioning introduces a functional coordinate system for brain functional parcellation, enhancing the positional encoding of different Regions of Interest (ROIs). Spatiotemporal Masking, tailored to the unique characteristics of fMRI data, addresses the challenge of heterogeneous time-series patches. These methodologies enhance model performance and advance our understanding of the neural circuits underlying cognition. Overall, Brain-JEPA is paving the way to address pivotal questions of building brain functional coordinate system and masking brain activity at the AI-neuroscience interface, and setting a potentially new paradigm in brain activity analysis through downstream adaptation.
Prompt Your Brain: Scaffold Prompt Tuning for Efficient Adaptation of fMRI Pre-trained Model
Aug 20, 2024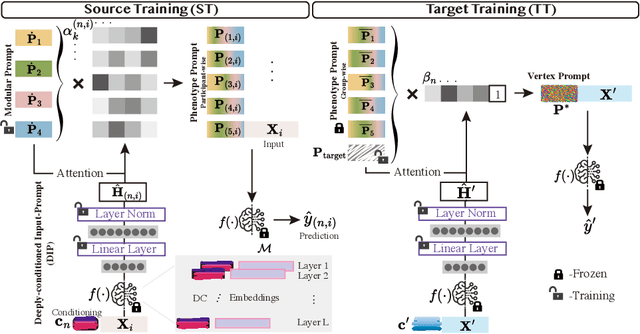
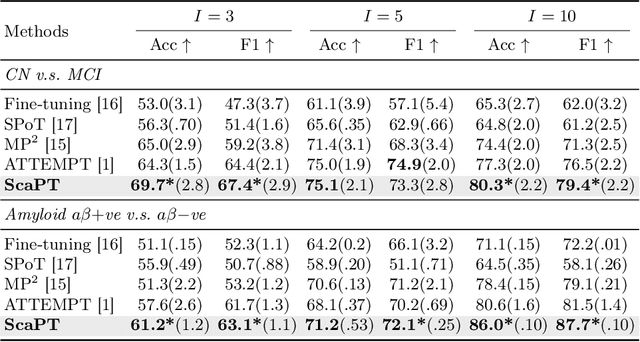
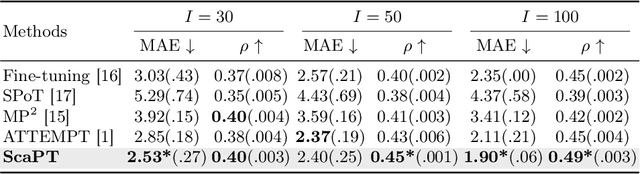
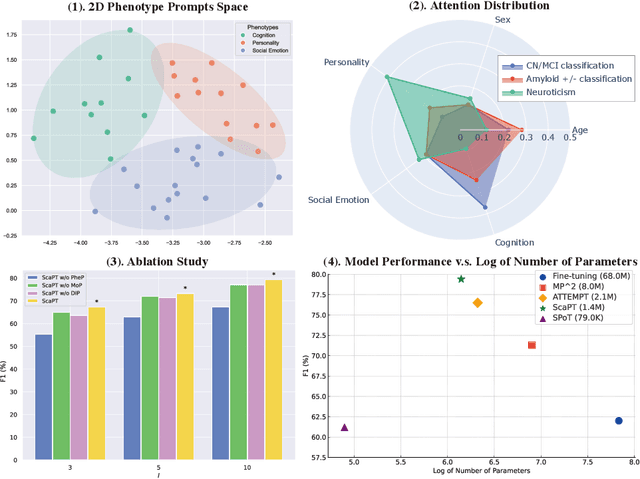
Abstract:We introduce Scaffold Prompt Tuning (ScaPT), a novel prompt-based framework for adapting large-scale functional magnetic resonance imaging (fMRI) pre-trained models to downstream tasks, with high parameter efficiency and improved performance compared to fine-tuning and baselines for prompt tuning. The full fine-tuning updates all pre-trained parameters, which may distort the learned feature space and lead to overfitting with limited training data which is common in fMRI fields. In contrast, we design a hierarchical prompt structure that transfers the knowledge learned from high-resource tasks to low-resource ones. This structure, equipped with a Deeply-conditioned Input-Prompt (DIP) mapping module, allows for efficient adaptation by updating only 2% of the trainable parameters. The framework enhances semantic interpretability through attention mechanisms between inputs and prompts, and it clusters prompts in the latent space in alignment with prior knowledge. Experiments on public resting state fMRI datasets reveal ScaPT outperforms fine-tuning and multitask-based prompt tuning in neurodegenerative diseases diagnosis/prognosis and personality trait prediction, even with fewer than 20 participants. It highlights ScaPT's efficiency in adapting pre-trained fMRI models to low-resource tasks.
Mixup Your Own Pairs
Sep 29, 2023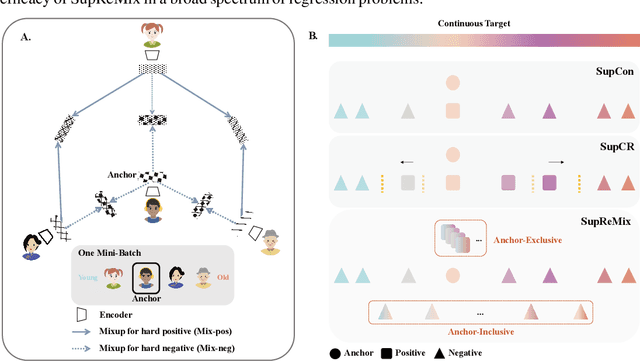
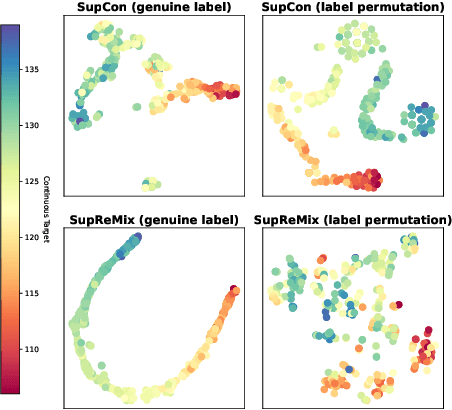
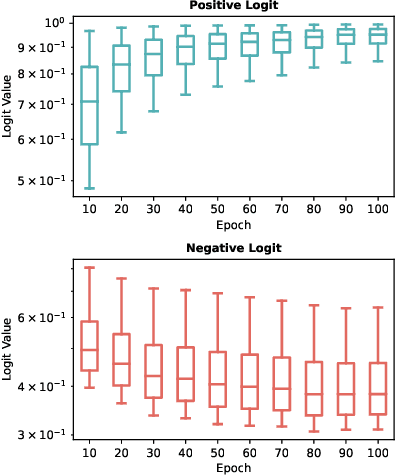
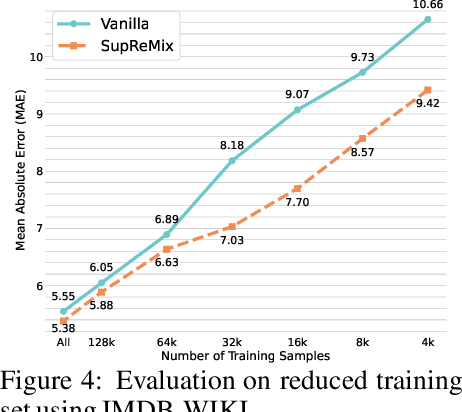
Abstract:In representation learning, regression has traditionally received less attention than classification. Directly applying representation learning techniques designed for classification to regression often results in fragmented representations in the latent space, yielding sub-optimal performance. In this paper, we argue that the potential of contrastive learning for regression has been overshadowed due to the neglect of two crucial aspects: ordinality-awareness and hardness. To address these challenges, we advocate "mixup your own contrastive pairs for supervised contrastive regression", instead of relying solely on real/augmented samples. Specifically, we propose Supervised Contrastive Learning for Regression with Mixup (SupReMix). It takes anchor-inclusive mixtures (mixup of the anchor and a distinct negative sample) as hard negative pairs and anchor-exclusive mixtures (mixup of two distinct negative samples) as hard positive pairs at the embedding level. This strategy formulates harder contrastive pairs by integrating richer ordinal information. Through extensive experiments on six regression datasets including 2D images, volumetric images, text, tabular data, and time-series signals, coupled with theoretical analysis, we demonstrate that SupReMix pre-training fosters continuous ordered representations of regression data, resulting in significant improvement in regression performance. Furthermore, SupReMix is superior to other approaches in a range of regression challenges including transfer learning, imbalanced training data, and scenarios with fewer training samples.
Beyond the Snapshot: Brain Tokenized Graph Transformer for Longitudinal Brain Functional Connectome Embedding
Jul 13, 2023

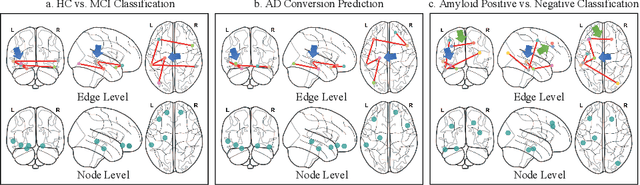
Abstract:Under the framework of network-based neurodegeneration, brain functional connectome (FC)-based Graph Neural Networks (GNN) have emerged as a valuable tool for the diagnosis and prognosis of neurodegenerative diseases such as Alzheimer's disease (AD). However, these models are tailored for brain FC at a single time point instead of characterizing FC trajectory. Discerning how FC evolves with disease progression, particularly at the predementia stages such as cognitively normal individuals with amyloid deposition or individuals with mild cognitive impairment (MCI), is crucial for delineating disease spreading patterns and developing effective strategies to slow down or even halt disease advancement. In this work, we proposed the first interpretable framework for brain FC trajectory embedding with application to neurodegenerative disease diagnosis and prognosis, namely Brain Tokenized Graph Transformer (Brain TokenGT). It consists of two modules: 1) Graph Invariant and Variant Embedding (GIVE) for generation of node and spatio-temporal edge embeddings, which were tokenized for downstream processing; 2) Brain Informed Graph Transformer Readout (BIGTR) which augments previous tokens with trainable type identifiers and non-trainable node identifiers and feeds them into a standard transformer encoder to readout. We conducted extensive experiments on two public longitudinal fMRI datasets of the AD continuum for three tasks, including differentiating MCI from controls, predicting dementia conversion in MCI, and classification of amyloid positive or negative cognitively normal individuals. Based on brain FC trajectory, the proposed Brain TokenGT approach outperformed all the other benchmark models and at the same time provided excellent interpretability. The code is available at https://github.com/ZijianD/Brain-TokenGT.git
Cinematic Mindscapes: High-quality Video Reconstruction from Brain Activity
May 19, 2023
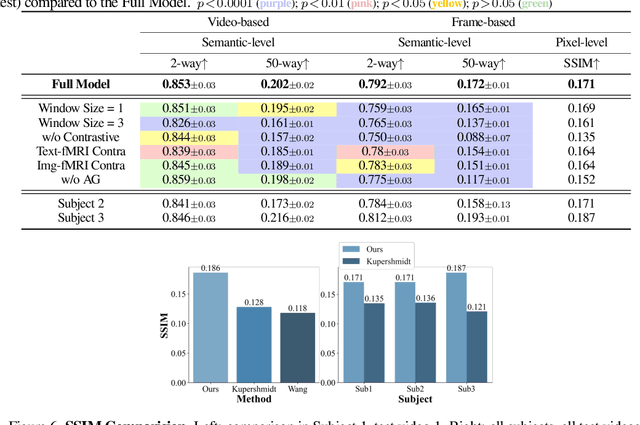
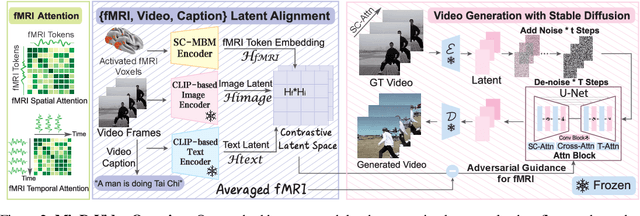
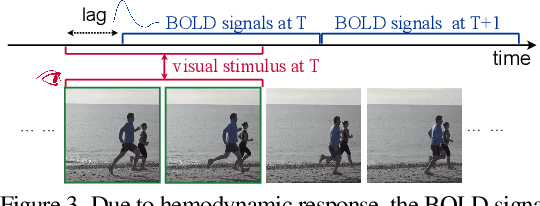
Abstract:Reconstructing human vision from brain activities has been an appealing task that helps to understand our cognitive process. Even though recent research has seen great success in reconstructing static images from non-invasive brain recordings, work on recovering continuous visual experiences in the form of videos is limited. In this work, we propose Mind-Video that learns spatiotemporal information from continuous fMRI data of the cerebral cortex progressively through masked brain modeling, multimodal contrastive learning with spatiotemporal attention, and co-training with an augmented Stable Diffusion model that incorporates network temporal inflation. We show that high-quality videos of arbitrary frame rates can be reconstructed with Mind-Video using adversarial guidance. The recovered videos were evaluated with various semantic and pixel-level metrics. We achieved an average accuracy of 85% in semantic classification tasks and 0.19 in structural similarity index (SSIM), outperforming the previous state-of-the-art by 45%. We also show that our model is biologically plausible and interpretable, reflecting established physiological processes.
Decomposing 3D Neuroimaging into 2+1D Processing for Schizophrenia Recognition
Nov 22, 2022Abstract:Deep learning has been successfully applied to recognizing both natural images and medical images. However, there remains a gap in recognizing 3D neuroimaging data, especially for psychiatric diseases such as schizophrenia and depression that have no visible alteration in specific slices. In this study, we propose to process the 3D data by a 2+1D framework so that we can exploit the powerful deep 2D Convolutional Neural Network (CNN) networks pre-trained on the huge ImageNet dataset for 3D neuroimaging recognition. Specifically, 3D volumes of Magnetic Resonance Imaging (MRI) metrics (grey matter, white matter, and cerebrospinal fluid) are decomposed to 2D slices according to neighboring voxel positions and inputted to 2D CNN models pre-trained on the ImageNet to extract feature maps from three views (axial, coronal, and sagittal). Global pooling is applied to remove redundant information as the activation patterns are sparsely distributed over feature maps. Channel-wise and slice-wise convolutions are proposed to aggregate the contextual information in the third view dimension unprocessed by the 2D CNN model. Multi-metric and multi-view information are fused for final prediction. Our approach outperforms handcrafted feature-based machine learning, deep feature approach with a support vector machine (SVM) classifier and 3D CNN models trained from scratch with better cross-validation results on publicly available Northwestern University Schizophrenia Dataset and the results are replicated on another independent dataset.
Seeing Beyond the Brain: Conditional Diffusion Model with Sparse Masked Modeling for Vision Decoding
Nov 15, 2022


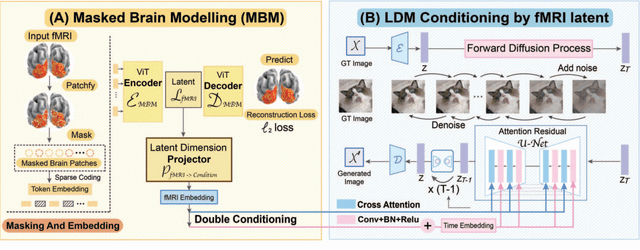
Abstract:Decoding visual stimuli from brain recordings aims to deepen our understanding of the human visual system and build a solid foundation for bridging human and computer vision through the Brain-Computer Interface. However, reconstructing high-quality images with correct semantics from brain recordings is a challenging problem due to the complex underlying representations of brain signals and the scarcity of data annotations. In this work, we present MinD-Vis: Sparse Masked Brain Modeling with Double-Conditioned Latent Diffusion Model for Human Vision Decoding. Firstly, we learn an effective self-supervised representation of fMRI data using mask modeling in a large latent space inspired by the sparse coding of information in the primary visual cortex. Then by augmenting a latent diffusion model with double-conditioning, we show that MinD-Vis can reconstruct highly plausible images with semantically matching details from brain recordings using very few paired annotations. We benchmarked our model qualitatively and quantitatively; the experimental results indicate that our method outperformed state-of-the-art in both semantic mapping (100-way semantic classification) and generation quality (FID) by 66% and 41% respectively. An exhaustive ablation study was also conducted to analyze our framework.
Brain MRI-based 3D Convolutional Neural Networks for Classification of Schizophrenia and Controls
Mar 14, 2020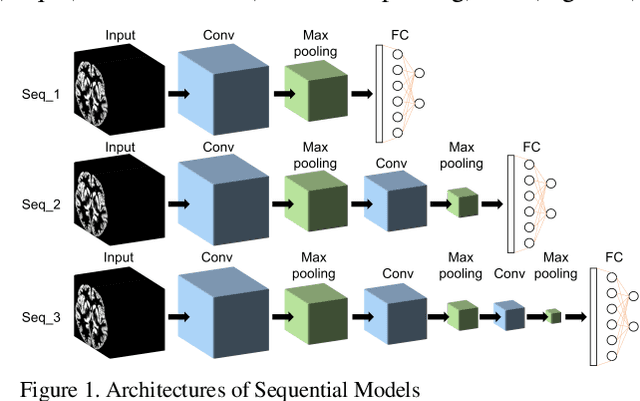

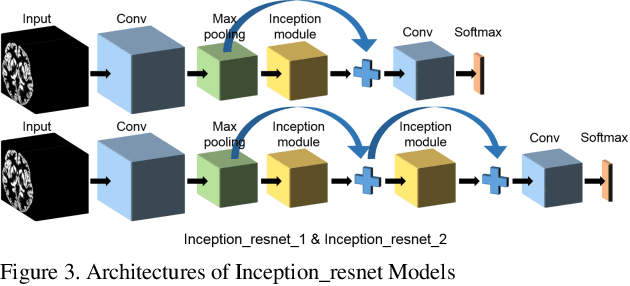
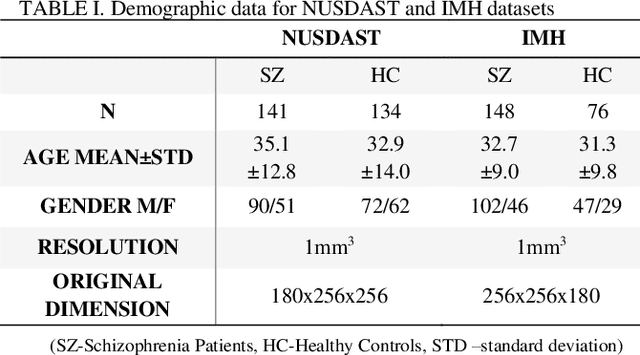
Abstract:Convolutional Neural Network (CNN) has been successfully applied on classification of both natural images and medical images but not yet been applied to differentiating patients with schizophrenia from healthy controls. Given the subtle, mixed, and sparsely distributed brain atrophy patterns of schizophrenia, the capability of automatic feature learning makes CNN a powerful tool for classifying schizophrenia from controls as it removes the subjectivity in selecting relevant spatial features. To examine the feasibility of applying CNN to classification of schizophrenia and controls based on structural Magnetic Resonance Imaging (MRI), we built 3D CNN models with different architectures and compared their performance with a handcrafted feature-based machine learning approach. Support vector machine (SVM) was used as classifier and Voxel-based Morphometry (VBM) was used as feature for handcrafted feature-based machine learning. 3D CNN models with sequential architecture, inception module and residual module were trained from scratch. CNN models achieved higher cross-validation accuracy than handcrafted feature-based machine learning. Moreover, testing on an independent dataset, 3D CNN models greatly outperformed handcrafted feature-based machine learning. This study underscored the potential of CNN for identifying patients with schizophrenia using 3D brain MR images and paved the way for imaging-based individual-level diagnosis and prognosis in psychiatric disorders.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge