Mengjiao Hu
Decomposing 3D Neuroimaging into 2+1D Processing for Schizophrenia Recognition
Nov 22, 2022Abstract:Deep learning has been successfully applied to recognizing both natural images and medical images. However, there remains a gap in recognizing 3D neuroimaging data, especially for psychiatric diseases such as schizophrenia and depression that have no visible alteration in specific slices. In this study, we propose to process the 3D data by a 2+1D framework so that we can exploit the powerful deep 2D Convolutional Neural Network (CNN) networks pre-trained on the huge ImageNet dataset for 3D neuroimaging recognition. Specifically, 3D volumes of Magnetic Resonance Imaging (MRI) metrics (grey matter, white matter, and cerebrospinal fluid) are decomposed to 2D slices according to neighboring voxel positions and inputted to 2D CNN models pre-trained on the ImageNet to extract feature maps from three views (axial, coronal, and sagittal). Global pooling is applied to remove redundant information as the activation patterns are sparsely distributed over feature maps. Channel-wise and slice-wise convolutions are proposed to aggregate the contextual information in the third view dimension unprocessed by the 2D CNN model. Multi-metric and multi-view information are fused for final prediction. Our approach outperforms handcrafted feature-based machine learning, deep feature approach with a support vector machine (SVM) classifier and 3D CNN models trained from scratch with better cross-validation results on publicly available Northwestern University Schizophrenia Dataset and the results are replicated on another independent dataset.
Brain MRI-based 3D Convolutional Neural Networks for Classification of Schizophrenia and Controls
Mar 14, 2020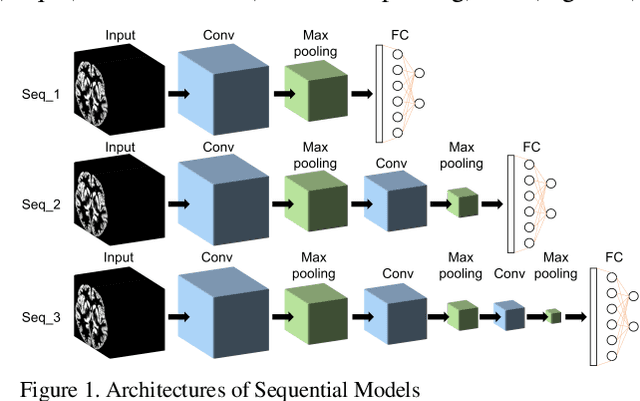

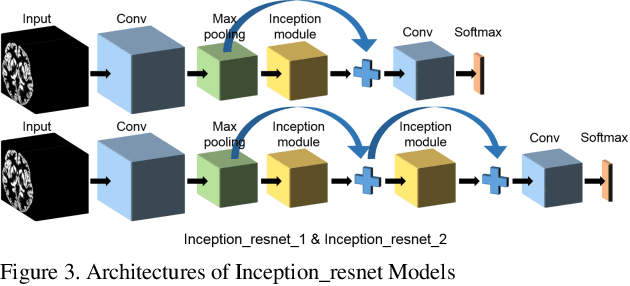
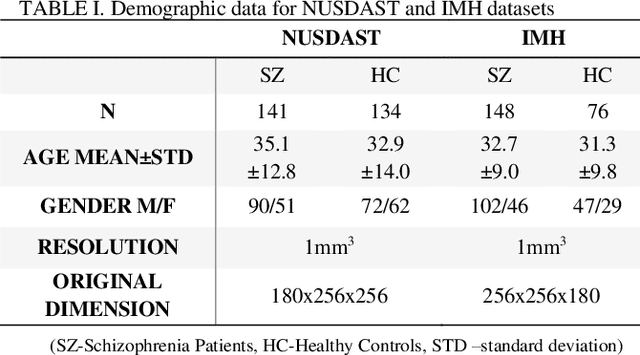
Abstract:Convolutional Neural Network (CNN) has been successfully applied on classification of both natural images and medical images but not yet been applied to differentiating patients with schizophrenia from healthy controls. Given the subtle, mixed, and sparsely distributed brain atrophy patterns of schizophrenia, the capability of automatic feature learning makes CNN a powerful tool for classifying schizophrenia from controls as it removes the subjectivity in selecting relevant spatial features. To examine the feasibility of applying CNN to classification of schizophrenia and controls based on structural Magnetic Resonance Imaging (MRI), we built 3D CNN models with different architectures and compared their performance with a handcrafted feature-based machine learning approach. Support vector machine (SVM) was used as classifier and Voxel-based Morphometry (VBM) was used as feature for handcrafted feature-based machine learning. 3D CNN models with sequential architecture, inception module and residual module were trained from scratch. CNN models achieved higher cross-validation accuracy than handcrafted feature-based machine learning. Moreover, testing on an independent dataset, 3D CNN models greatly outperformed handcrafted feature-based machine learning. This study underscored the potential of CNN for identifying patients with schizophrenia using 3D brain MR images and paved the way for imaging-based individual-level diagnosis and prognosis in psychiatric disorders.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge