Grzegorz Skorupko
Federated nnU-Net for Privacy-Preserving Medical Image Segmentation
Mar 04, 2025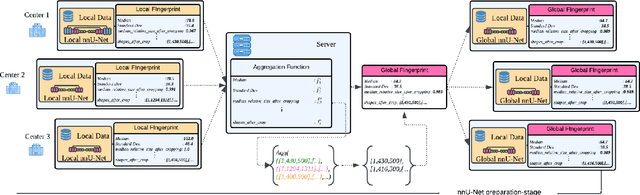
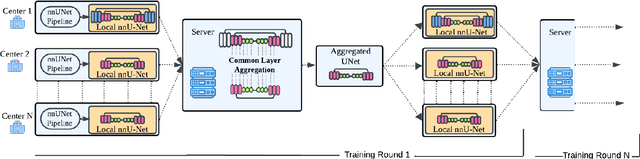
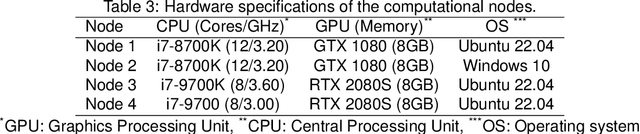
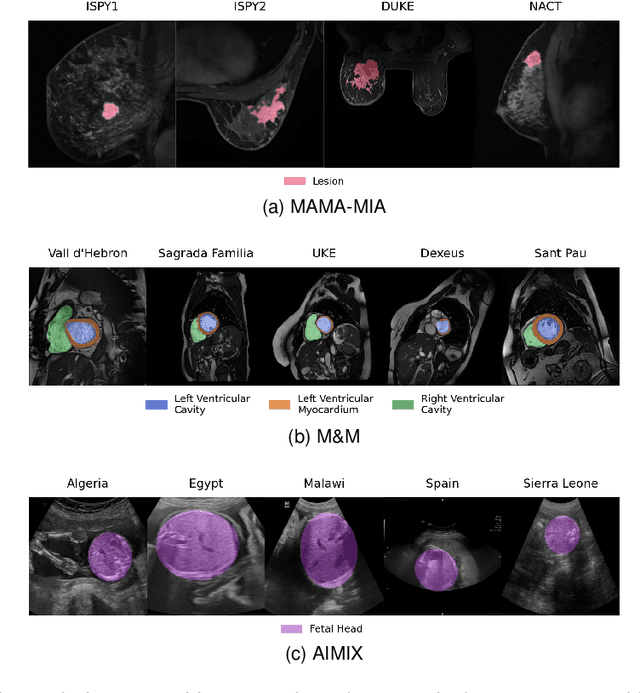
Abstract:The nnU-Net framework has played a crucial role in medical image segmentation and has become the gold standard in multitudes of applications targeting different diseases, organs, and modalities. However, so far it has been used primarily in a centralized approach where the data collected from hospitals are stored in one center and used to train the nnU-Net. This centralized approach has various limitations, such as leakage of sensitive patient information and violation of patient privacy. Federated learning is one of the approaches to train a segmentation model in a decentralized manner that helps preserve patient privacy. In this paper, we propose FednnU-Net, a federated learning extension of nnU-Net. We introduce two novel federated learning methods to the nnU-Net framework - Federated Fingerprint Extraction (FFE) and Asymmetric Federated Averaging (AsymFedAvg) - and experimentally show their consistent performance for breast, cardiac and fetal segmentation using 6 datasets representing samples from 18 institutions. Additionally, to further promote research and deployment of decentralized training in privacy constrained institutions, we make our plug-n-play framework public. The source-code is available at https://github.com/faildeny/FednnUNet .
Enhancing the Utility of Privacy-Preserving Cancer Classification using Synthetic Data
Jul 17, 2024Abstract:Deep learning holds immense promise for aiding radiologists in breast cancer detection. However, achieving optimal model performance is hampered by limitations in availability and sharing of data commonly associated to patient privacy concerns. Such concerns are further exacerbated, as traditional deep learning models can inadvertently leak sensitive training information. This work addresses these challenges exploring and quantifying the utility of privacy-preserving deep learning techniques, concretely, (i) differentially private stochastic gradient descent (DP-SGD) and (ii) fully synthetic training data generated by our proposed malignancy-conditioned generative adversarial network. We assess these methods via downstream malignancy classification of mammography masses using a transformer model. Our experimental results depict that synthetic data augmentation can improve privacy-utility tradeoffs in differentially private model training. Further, model pretraining on synthetic data achieves remarkable performance, which can be further increased with DP-SGD fine-tuning across all privacy guarantees. With this first in-depth exploration of privacy-preserving deep learning in breast imaging, we address current and emerging clinical privacy requirements and pave the way towards the adoption of private high-utility deep diagnostic models. Our reproducible codebase is publicly available at https://github.com/RichardObi/mammo_dp.
Debiasing Cardiac Imaging with Controlled Latent Diffusion Models
Mar 28, 2024

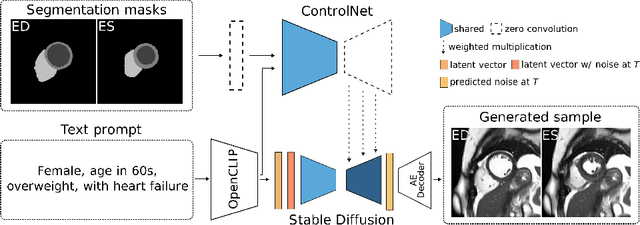

Abstract:The progress in deep learning solutions for disease diagnosis and prognosis based on cardiac magnetic resonance imaging is hindered by highly imbalanced and biased training data. To address this issue, we propose a method to alleviate imbalances inherent in datasets through the generation of synthetic data based on sensitive attributes such as sex, age, body mass index, and health condition. We adopt ControlNet based on a denoising diffusion probabilistic model to condition on text assembled from patient metadata and cardiac geometry derived from segmentation masks using a large-cohort study, specifically, the UK Biobank. We assess our method by evaluating the realism of the generated images using established quantitative metrics. Furthermore, we conduct a downstream classification task aimed at debiasing a classifier by rectifying imbalances within underrepresented groups through synthetically generated samples. Our experiments demonstrate the effectiveness of the proposed approach in mitigating dataset imbalances, such as the scarcity of younger patients or individuals with normal BMI level suffering from heart failure. This work represents a major step towards the adoption of synthetic data for the development of fair and generalizable models for medical classification tasks. Notably, we conduct all our experiments using a single, consumer-level GPU to highlight the feasibility of our approach within resource-constrained environments. Our code is available at https://github.com/faildeny/debiasing-cardiac-mri.
Towards Learning Contrast Kinetics with Multi-Condition Latent Diffusion Models
Mar 20, 2024



Abstract:Contrast agents in dynamic contrast enhanced magnetic resonance imaging allow to localize tumors and observe their contrast kinetics, which is essential for cancer characterization and respective treatment decision-making. However, contrast agent administration is not only associated with adverse health risks, but also restricted for patients during pregnancy, and for those with kidney malfunction, or other adverse reactions. With contrast uptake as key biomarker for lesion malignancy, cancer recurrence risk, and treatment response, it becomes pivotal to reduce the dependency on intravenous contrast agent administration. To this end, we propose a multi-conditional latent diffusion model capable of acquisition time-conditioned image synthesis of DCE-MRI temporal sequences. To evaluate medical image synthesis, we additionally propose and validate the Fr\'echet radiomics distance as an image quality measure based on biomarker variability between synthetic and real imaging data. Our results demonstrate our method's ability to generate realistic multi-sequence fat-saturated breast DCE-MRI and uncover the emerging potential of deep learning based contrast kinetics simulation. We publicly share our accessible codebase at https://github.com/RichardObi/ccnet.
medigan: A Python Library of Pretrained Generative Models for Enriched Data Access in Medical Imaging
Sep 28, 2022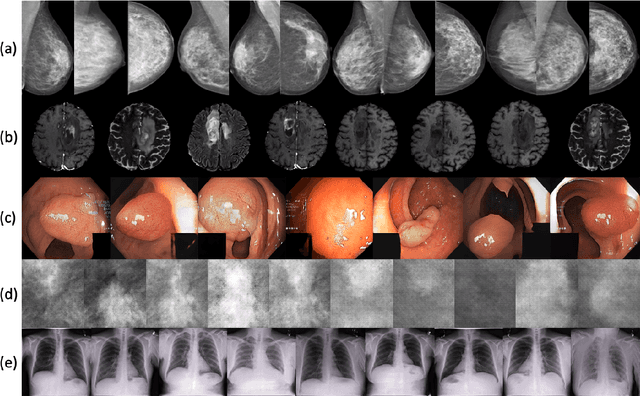
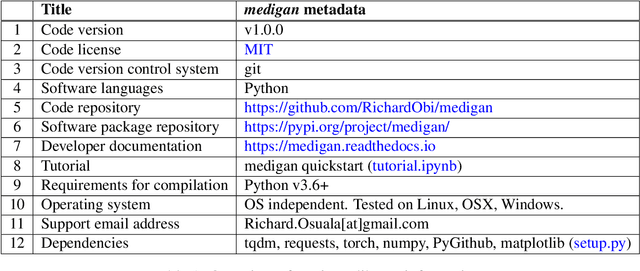
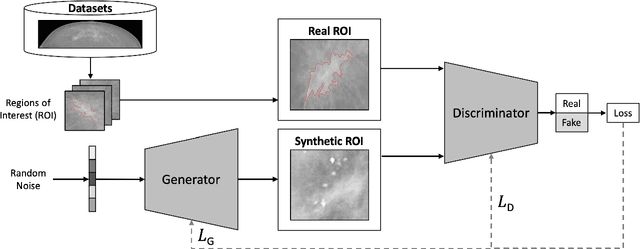
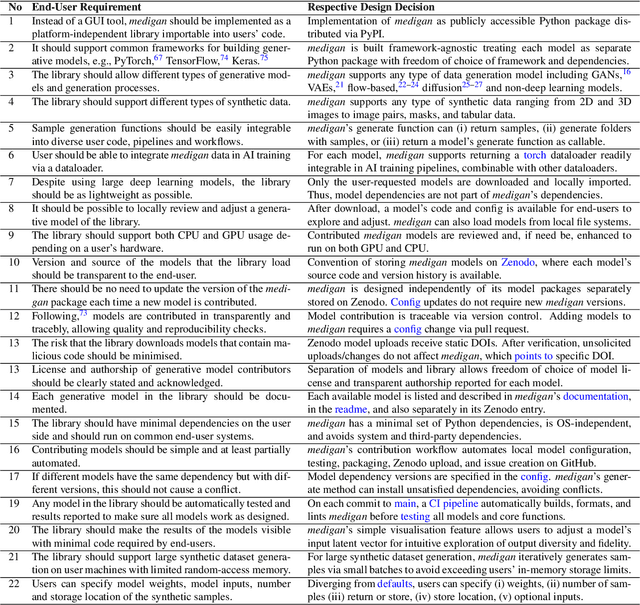
Abstract:Synthetic data generated by generative models can enhance the performance and capabilities of data-hungry deep learning models in medical imaging. However, there is (1) limited availability of (synthetic) datasets and (2) generative models are complex to train, which hinders their adoption in research and clinical applications. To reduce this entry barrier, we propose medigan, a one-stop shop for pretrained generative models implemented as an open-source framework-agnostic Python library. medigan allows researchers and developers to create, increase, and domain-adapt their training data in just a few lines of code. Guided by design decisions based on gathered end-user requirements, we implement medigan based on modular components for generative model (i) execution, (ii) visualisation, (iii) search & ranking, and (iv) contribution. The library's scalability and design is demonstrated by its growing number of integrated and readily-usable pretrained generative models consisting of 21 models utilising 9 different Generative Adversarial Network architectures trained on 11 datasets from 4 domains, namely, mammography, endoscopy, x-ray, and MRI. Furthermore, 3 applications of medigan are analysed in this work, which include (a) enabling community-wide sharing of restricted data, (b) investigating generative model evaluation metrics, and (c) improving clinical downstream tasks. In (b), extending on common medical image synthesis assessment and reporting standards, we show Fr\'echet Inception Distance variability based on image normalisation and radiology-specific feature extraction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge