Gloria Menegaz
Department of Engineering for Innovation Medicine, University of Verona, Verona, Italy
Guidelines For The Choice Of The Baseline in XAI Attribution Methods
Mar 25, 2025Abstract:Given the broad adoption of artificial intelligence, it is essential to provide evidence that AI models are reliable, trustable, and fair. To this end, the emerging field of eXplainable AI develops techniques to probe such requirements, counterbalancing the hype pushing the pervasiveness of this technology. Among the many facets of this issue, this paper focuses on baseline attribution methods, aiming at deriving a feature attribution map at the network input relying on a "neutral" stimulus usually called "baseline". The choice of the baseline is crucial as it determines the explanation of the network behavior. In this framework, this paper has the twofold goal of shedding light on the implications of the choice of the baseline and providing a simple yet effective method for identifying the best baseline for the task. To achieve this, we propose a decision boundary sampling method, since the baseline, by definition, lies on the decision boundary, which naturally becomes the search domain. Experiments are performed on synthetic examples and validated relying on state-of-the-art methods. Despite being limited to the experimental scope, this contribution is relevant as it offers clear guidelines and a simple proxy for baseline selection, reducing ambiguity and enhancing deep models' reliability and trust.
Microscopic Propagator Imaging (MPI) with Diffusion MRI
Feb 28, 2025



Abstract:We propose Microscopic Propagator Imaging (MPI) as a novel method to retrieve the indices of the microscopic propagator which is the probability density function of water displacements due to diffusion within the nervous tissue microstructures. Unlike the Ensemble Average Propagator indices or the Diffusion Tensor Imaging metrics, MPI indices are independent from the mesoscopic organization of the tissue such as the presence of multiple axonal bundle directions and orientation dispersion. As a consequence, MPI indices are more specific to the volumes, sizes, and types of microstructures, like axons and cells, that are present in the tissue. Thus, changes in MPI indices can be more directly linked to alterations in the presence and integrity of microstructures themselves. The methodology behind MPI is rooted on zonal modeling of spherical harmonics, signal simulation, and machine learning regression, and is demonstrated on both synthetic and Human Diffusion MRI data.
Multimodal MRI-based Detection of Amyloid Status in Alzheimer's Disease Continuum
Jun 19, 2024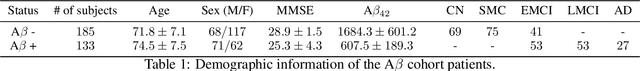
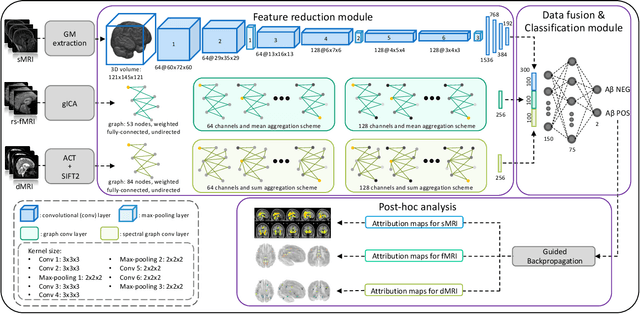

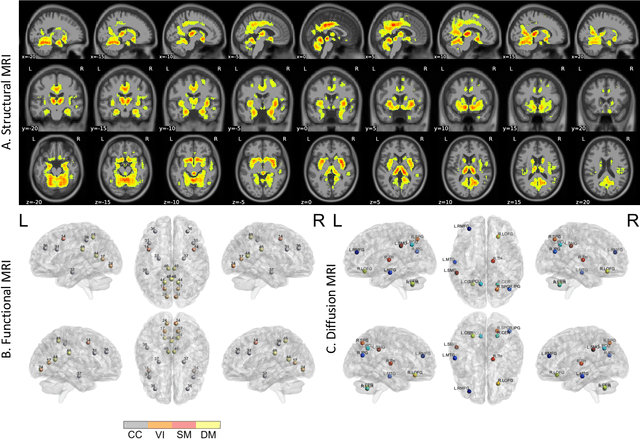
Abstract:Amyloid-$\beta$ (A$\beta$) plaques in conjunction with hyperphosphorylated tau proteins in the form of neurofibrillary tangles are the two neuropathological hallmarks of Alzheimer's disease (AD). In particular, the accumulation of A$\beta$ plaques, as evinced by the A/T/N (amyloid/tau/neurodegeneration) framework, marks the initial stage. Thus, the identification of individuals with A$\beta$ positivity could enable early diagnosis and potentially lead to more effective interventions. Deep learning methods relying mainly on amyloid PET images have been employed to this end. However, PET imaging has some disadvantages, including the need of radiotracers and expensive acquisitions. Hence, in this work, we propose a novel multimodal approach that integrates information from structural, functional, and diffusion MRI data to discriminate A$\beta$ status in the AD continuum. Our method achieved an accuracy of $0.762\pm0.04$. Furthermore, a \textit{post-hoc} explainability analysis (guided backpropagation) was performed to retrieve the brain regions that most influenced the model predictions. This analysis identified some key regions that were common across modalities, some of which were well-established AD-discriminative biomarkers and related to A$\beta$ deposition, such as the hippocampus, thalamus, precuneus, and cingulate gyrus. Hence, our study demonstrates the potential viability of MRI-based characterization of A$\beta$ status, paving the way for further research in this domain.
An interpretable generative multimodal neuroimaging-genomics framework for decoding Alzheimer's disease
Jun 19, 2024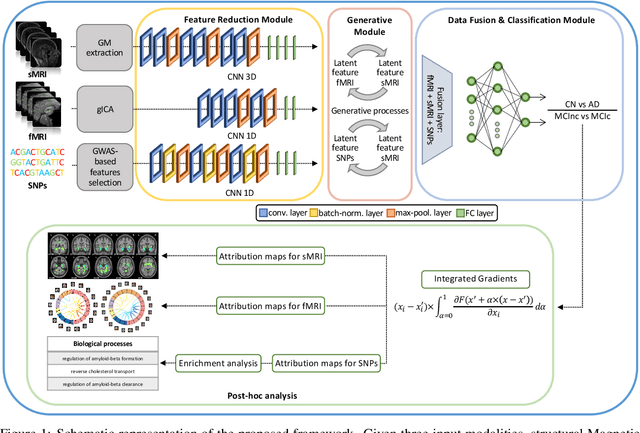

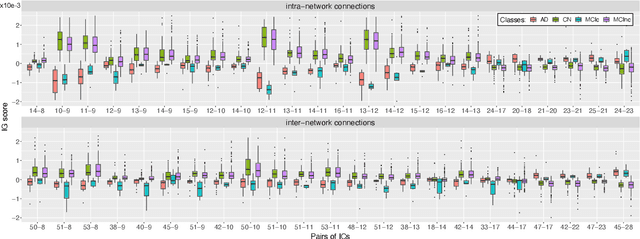

Abstract:Alzheimer's disease (AD) is the most prevalent form of dementia with a progressive decline in cognitive abilities. The AD continuum encompasses a prodormal stage known as Mild Cognitive Impairment (MCI), where patients may either progress to AD or remain stable. In this study, we leveraged structural and functional MRI to investigate the disease-induced grey matter and functional network connectivity changes. Moreover, considering AD's strong genetic component, we introduce SNPs as a third channel. Given such diverse inputs, missing one or more modalities is a typical concern of multimodal methods. We hence propose a novel deep learning-based classification framework where generative module employing Cycle GANs was adopted to impute missing data within the latent space. Additionally, we adopted an Explainable AI method, Integrated Gradients, to extract input features relevance, enhancing our understanding of the learned representations. Two critical tasks were addressed: AD detection and MCI conversion prediction. Experimental results showed that our model was able to reach the SOA in the classification of CN/AD reaching an average test accuracy of $0.926\pm0.02$. For the MCI task, we achieved an average prediction accuracy of $0.711\pm0.01$ using the pre-trained model for CN/AD. The interpretability analysis revealed significant grey matter modulations in cortical and subcortical brain areas well known for their association with AD. Moreover, impairments in sensory-motor and visual resting state network connectivity along the disease continuum, as well as mutations in SNPs defining biological processes linked to amyloid-beta and cholesterol formation clearance and regulation, were identified as contributors to the achieved performance. Overall, our integrative deep learning approach shows promise for AD detection and MCI prediction, while shading light on important biological insights.
Commentary on explainable artificial intelligence methods: SHAP and LIME
May 08, 2023Abstract:eXplainable artificial intelligence (XAI) methods have emerged to convert the black box of machine learning models into a more digestible form. These methods help to communicate how the model works with the aim of making machine learning models more transparent and increasing the trust of end-users into their output. SHapley Additive exPlanations (SHAP) and Local Interpretable Model Agnostic Explanation (LIME) are two widely used XAI methods particularly with tabular data. In this commentary piece, we discuss the way the explainability metrics of these two methods are generated and propose a framework for interpretation of their outputs, highlighting their weaknesses and strengths.
Characterizing the contribution of dependent features in XAI methods
Apr 04, 2023Abstract:Explainable Artificial Intelligence (XAI) provides tools to help understanding how the machine learning models work and reach a specific outcome. It helps to increase the interpretability of models and makes the models more trustworthy and transparent. In this context, many XAI methods were proposed being SHAP and LIME the most popular. However, the proposed methods assume that used predictors in the machine learning models are independent which in general is not necessarily true. Such assumption casts shadows on the robustness of the XAI outcomes such as the list of informative predictors. Here, we propose a simple, yet useful proxy that modifies the outcome of any XAI feature ranking method allowing to account for the dependency among the predictors. The proposed approach has the advantage of being model-agnostic as well as simple to calculate the impact of each predictor in the model in presence of collinearity.
Explainable AI in Biomedical Signal and Image Processing: Promises and Challenges
Jul 09, 2022Abstract:Artificial intelligence has become pervasive across disciplines and fields, and biomedical image and signal processing is no exception. The growing and widespread interest on the topic has triggered a vast research activity that is reflected in an exponential research effort. Through study of massive and diverse biomedical data, machine and deep learning models have revolutionized various tasks such as modeling, segmentation, registration, classification and synthesis, outperforming traditional techniques. However, the difficulty in translating the results into biologically/clinically interpretable information is preventing their full exploitation in the field. Explainable AI (XAI) attempts to fill this translational gap by providing means to make the models interpretable and providing explanations. Different solutions have been proposed so far and are gaining increasing interest from the community. This paper aims at providing an overview on XAI in biomedical data processing and points to an upcoming Special Issue on Deep Learning in Biomedical Image and Signal Processing of the IEEE Signal Processing Magazine that is going to appear in March 2022.
NODDI-SH: a computational efficient NODDI extension for fODF estimation in diffusion MRI
Sep 01, 2017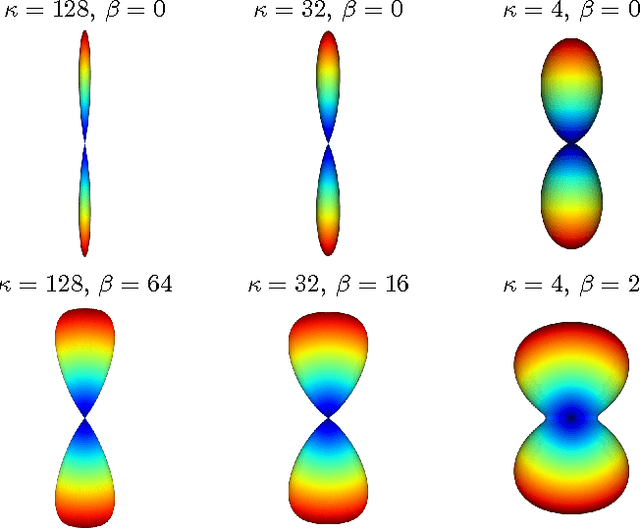
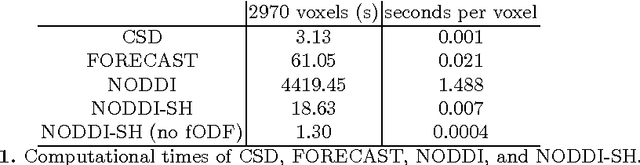
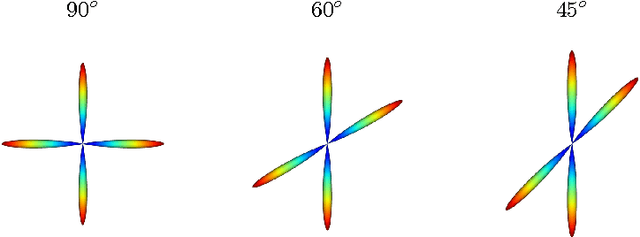
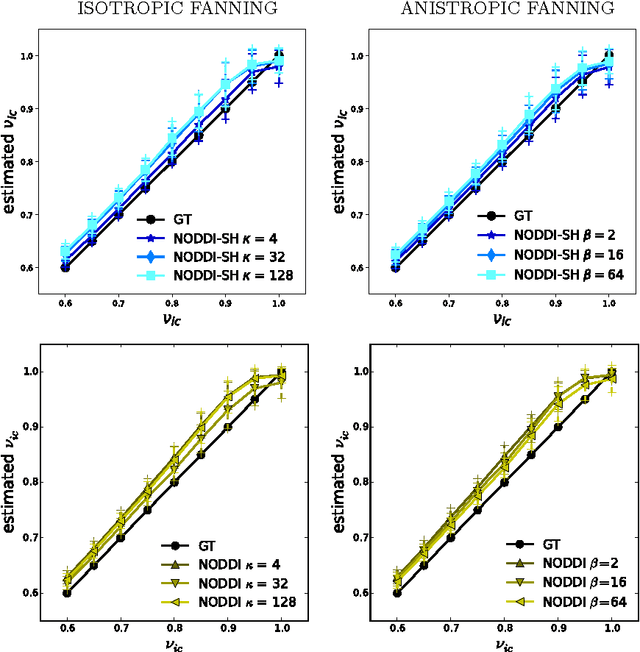
Abstract:Diffusion Magnetic Resonance Imaging (DMRI) is the only non-invasive imaging technique which is able to detect the principal directions of water diffusion as well as neurites density in the human brain. Exploiting the ability of Spherical Harmonics (SH) to model spherical functions, we propose a new reconstruction model for DMRI data which is able to estimate both the fiber Orientation Distribution Function (fODF) and the relative volume fractions of the neurites in each voxel, which is robust to multiple fiber crossings. We consider a Neurite Orientation Dispersion and Density Imaging (NODDI) inspired single fiber diffusion signal to be derived from three compartments: intracellular, extracellular, and cerebrospinal fluid. The model, called NODDI-SH, is derived by convolving the single fiber response with the fODF in each voxel. NODDI-SH embeds the calculation of the fODF and the neurite density in a unified mathematical model providing efficient, robust and accurate results. Results were validated on simulated data and tested on \textit{in-vivo} data of human brain, and compared to and Constrained Spherical Deconvolution (CSD) for benchmarking. Results revealed competitive performance in all respects and inherent adaptivity to local microstructure, while sensibly reducing the computational cost. We also investigated NODDI-SH performance when only a limited number of samples are available for the fitting, demonstrating that 60 samples are enough to obtain reliable results. The fast computational time and the low number of signal samples required, make NODDI-SH feasible for clinical application.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge