Giridhar Kaushik Ramachandran
Identifying Imaging Follow-Up in Radiology Reports: A Comparative Analysis of Traditional ML and LLM Approaches
Nov 14, 2025Abstract:Large language models (LLMs) have shown considerable promise in clinical natural language processing, yet few domain-specific datasets exist to rigorously evaluate their performance on radiology tasks. In this work, we introduce an annotated corpus of 6,393 radiology reports from 586 patients, each labeled for follow-up imaging status, to support the development and benchmarking of follow-up adherence detection systems. Using this corpus, we systematically compared traditional machine-learning classifiers, including logistic regression (LR), support vector machines (SVM), Longformer, and a fully fine-tuned Llama3-8B-Instruct, with recent generative LLMs. To evaluate generative LLMs, we tested GPT-4o and the open-source GPT-OSS-20B under two configurations: a baseline (Base) and a task-optimized (Advanced) setting that focused inputs on metadata, recommendation sentences, and their surrounding context. A refined prompt for GPT-OSS-20B further improved reasoning accuracy. Performance was assessed using precision, recall, and F1 scores with 95% confidence intervals estimated via non-parametric bootstrapping. Inter-annotator agreement was high (F1 = 0.846). GPT-4o (Advanced) achieved the best performance (F1 = 0.832), followed closely by GPT-OSS-20B (Advanced; F1 = 0.828). LR and SVM also performed strongly (F1 = 0.776 and 0.775), underscoring that while LLMs approach human-level agreement through prompt optimization, interpretable and resource-efficient models remain valuable baselines.
Adapting Biomedical Abstracts into Plain language using Large Language Models
Jan 26, 2025



Abstract:A vast amount of medical knowledge is available for public use through online health forums, and question-answering platforms on social media. The majority of the population in the United States doesn't have the right amount of health literacy to make the best use of that information. Health literacy means the ability to obtain and comprehend the basic health information to make appropriate health decisions. To build the bridge between this gap, organizations advocate adapting this medical knowledge into plain language. Building robust systems to automate the adaptations helps both medical and non-medical professionals best leverage the available information online. The goal of the Plain Language Adaptation of Biomedical Abstracts (PLABA) track is to adapt the biomedical abstracts in English language extracted from PubMed based on the questions asked in MedlinePlus for the general public using plain language at the sentence level. As part of this track, we leveraged the best open-source Large Language Models suitable and fine-tuned for dialog use cases. We compare and present the results for all of our systems and our ranking among the other participants' submissions. Our top performing GPT-4 based model ranked first in the avg. simplicity measure and 3rd on the avg. accuracy measure.
BioMistral-NLU: Towards More Generalizable Medical Language Understanding through Instruction Tuning
Oct 24, 2024



Abstract:Large language models (LLMs) such as ChatGPT are fine-tuned on large and diverse instruction-following corpora, and can generalize to new tasks. However, those instruction-tuned LLMs often perform poorly in specialized medical natural language understanding (NLU) tasks that require domain knowledge, granular text comprehension, and structured data extraction. To bridge the gap, we: (1) propose a unified prompting format for 7 important NLU tasks, % through span extraction and multi-choice question-answering (QA), (2) curate an instruction-tuning dataset, MNLU-Instruct, utilizing diverse existing open-source medical NLU corpora, and (3) develop BioMistral-NLU, a generalizable medical NLU model, through fine-tuning BioMistral on MNLU-Instruct. We evaluate BioMistral-NLU in a zero-shot setting, across 6 important NLU tasks, from two widely adopted medical NLU benchmarks: Biomedical Language Understanding Evaluation (BLUE) and Biomedical Language Understanding and Reasoning Benchmark (BLURB). Our experiments show that our BioMistral-NLU outperforms the original BioMistral, as well as the proprietary LLMs - ChatGPT and GPT-4. Our dataset-agnostic prompting strategy and instruction tuning step over diverse NLU tasks enhance LLMs' generalizability across diverse medical NLU tasks. Our ablation experiments show that instruction-tuning on a wider variety of tasks, even when the total number of training instances remains constant, enhances downstream zero-shot generalization.
CACER: Clinical Concept Annotations for Cancer Events and Relations
Sep 05, 2024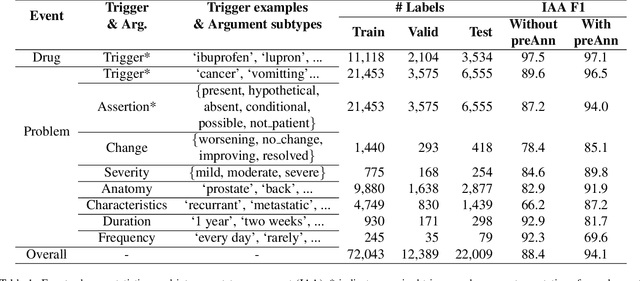
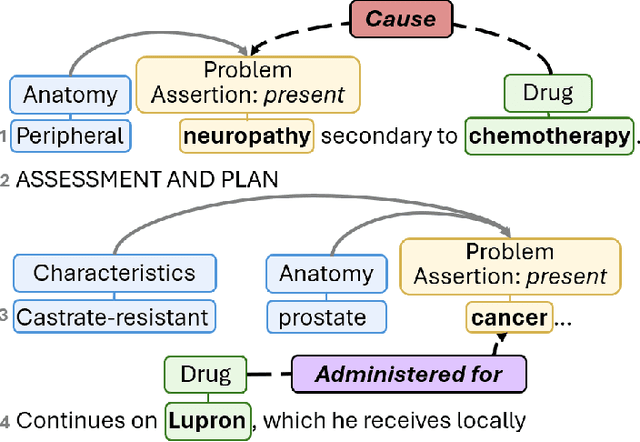
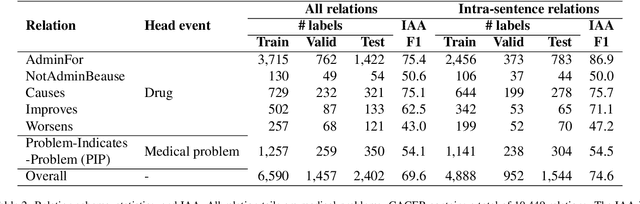
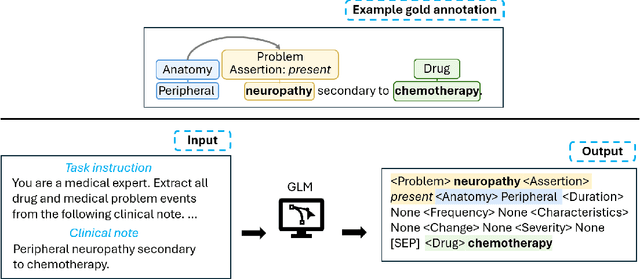
Abstract:Clinical notes contain unstructured representations of patient histories, including the relationships between medical problems and prescription drugs. To investigate the relationship between cancer drugs and their associated symptom burden, we extract structured, semantic representations of medical problem and drug information from the clinical narratives of oncology notes. We present Clinical Concept Annotations for Cancer Events and Relations (CACER), a novel corpus with fine-grained annotations for over 48,000 medical problems and drug events and 10,000 drug-problem and problem-problem relations. Leveraging CACER, we develop and evaluate transformer-based information extraction (IE) models such as BERT, Flan-T5, Llama3, and GPT-4 using fine-tuning and in-context learning (ICL). In event extraction, the fine-tuned BERT and Llama3 models achieved the highest performance at 88.2-88.0 F1, which is comparable to the inter-annotator agreement (IAA) of 88.4 F1. In relation extraction, the fine-tuned BERT, Flan-T5, and Llama3 achieved the highest performance at 61.8-65.3 F1. GPT-4 with ICL achieved the worst performance across both tasks. The fine-tuned models significantly outperformed GPT-4 in ICL, highlighting the importance of annotated training data and model optimization. Furthermore, the BERT models performed similarly to Llama3. For our task, LLMs offer no performance advantage over the smaller BERT models. The results emphasize the need for annotated training data to optimize models. Multiple fine-tuned transformer models achieved performance comparable to IAA for several extraction tasks.
* This is a pre-copy-editing, author-produced PDF of an article accepted for publication in JAMIA following peer review. The definitive publisher-authenticated version is available online at https://academic.oup.com/jamia/advance-article/doi/10.1093/jamia/ocae231/7748302
Extracting Social Determinants of Health from Pediatric Patient Notes Using Large Language Models: Novel Corpus and Methods
Apr 04, 2024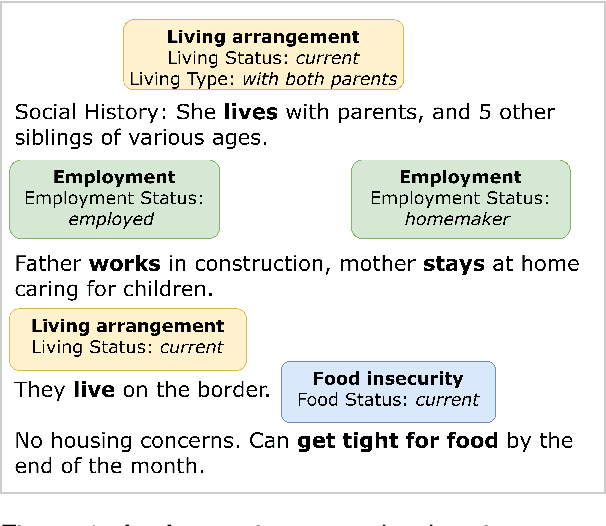
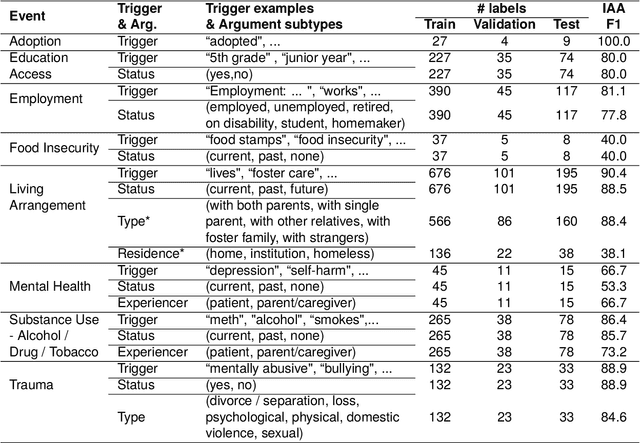
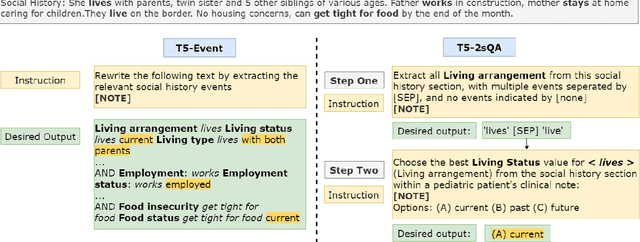
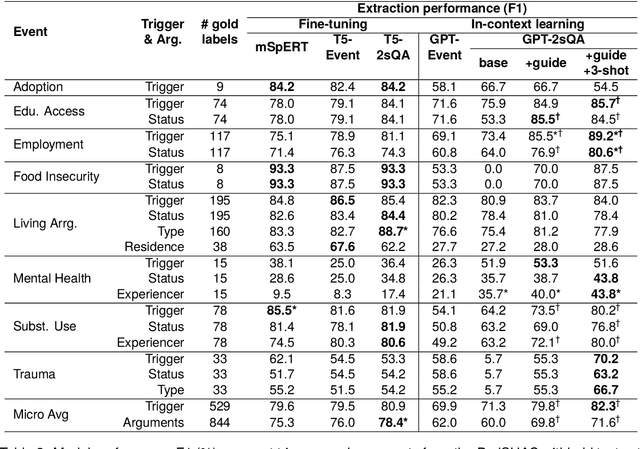
Abstract:Social determinants of health (SDoH) play a critical role in shaping health outcomes, particularly in pediatric populations where interventions can have long-term implications. SDoH are frequently studied in the Electronic Health Record (EHR), which provides a rich repository for diverse patient data. In this work, we present a novel annotated corpus, the Pediatric Social History Annotation Corpus (PedSHAC), and evaluate the automatic extraction of detailed SDoH representations using fine-tuned and in-context learning methods with Large Language Models (LLMs). PedSHAC comprises annotated social history sections from 1,260 clinical notes obtained from pediatric patients within the University of Washington (UW) hospital system. Employing an event-based annotation scheme, PedSHAC captures ten distinct health determinants to encompass living and economic stability, prior trauma, education access, substance use history, and mental health with an overall annotator agreement of 81.9 F1. Our proposed fine-tuning LLM-based extractors achieve high performance at 78.4 F1 for event arguments. In-context learning approaches with GPT-4 demonstrate promise for reliable SDoH extraction with limited annotated examples, with extraction performance at 82.3 F1 for event triggers.
A Novel Corpus of Annotated Medical Imaging Reports and Information Extraction Results Using BERT-based Language Models
Mar 27, 2024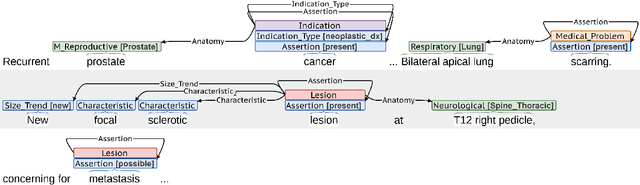
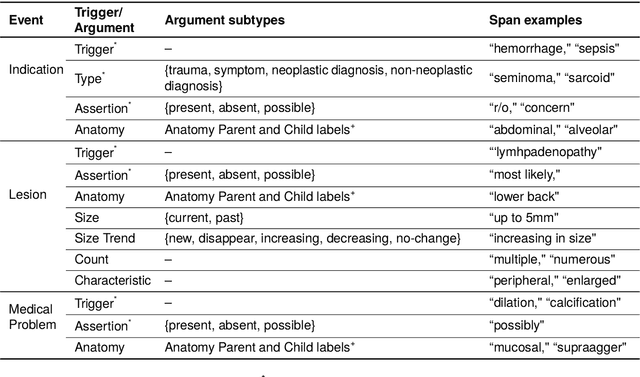
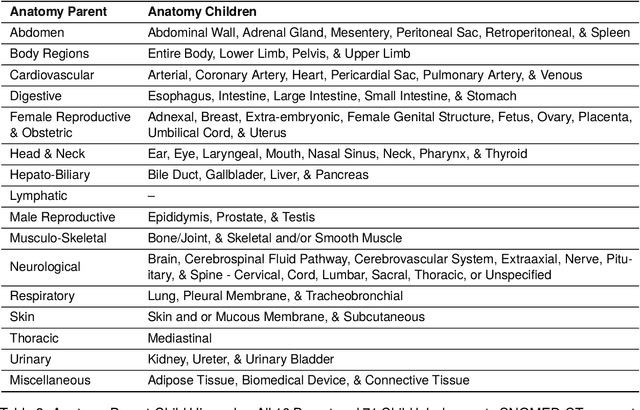
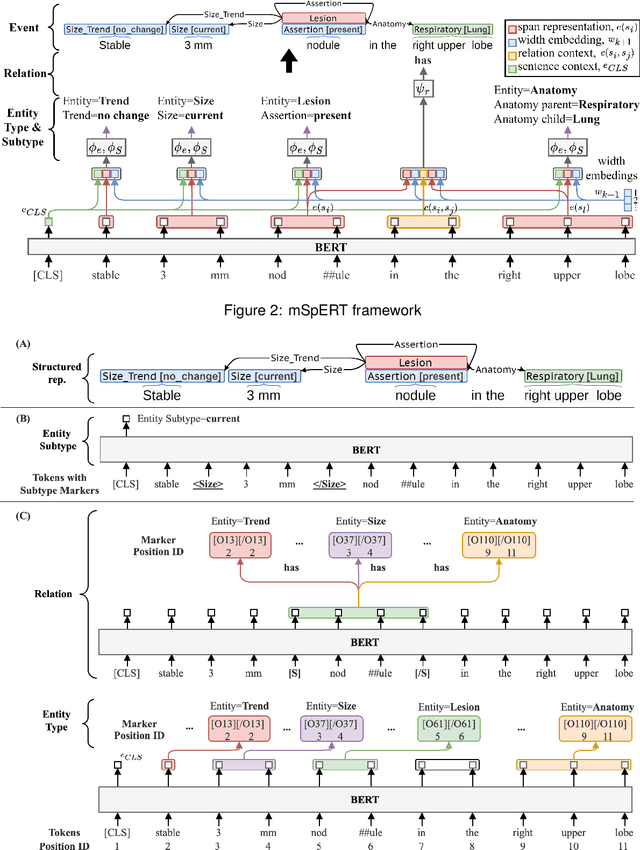
Abstract:Medical imaging is critical to the diagnosis, surveillance, and treatment of many health conditions, including oncological, neurological, cardiovascular, and musculoskeletal disorders, among others. Radiologists interpret these complex, unstructured images and articulate their assessments through narrative reports that remain largely unstructured. This unstructured narrative must be converted into a structured semantic representation to facilitate secondary applications such as retrospective analyses or clinical decision support. Here, we introduce the Corpus of Annotated Medical Imaging Reports (CAMIR), which includes 609 annotated radiology reports from three imaging modality types: Computed Tomography, Magnetic Resonance Imaging, and Positron Emission Tomography-Computed Tomography. Reports were annotated using an event-based schema that captures clinical indications, lesions, and medical problems. Each event consists of a trigger and multiple arguments, and a majority of the argument types, including anatomy, normalize the spans to pre-defined concepts to facilitate secondary use. CAMIR uniquely combines a granular event structure and concept normalization. To extract CAMIR events, we explored two BERT (Bi-directional Encoder Representation from Transformers)-based architectures, including an existing architecture (mSpERT) that jointly extracts all event information and a multi-step approach (PL-Marker++) that we augmented for the CAMIR schema.
Prompt-based Extraction of Social Determinants of Health Using Few-shot Learning
Jun 12, 2023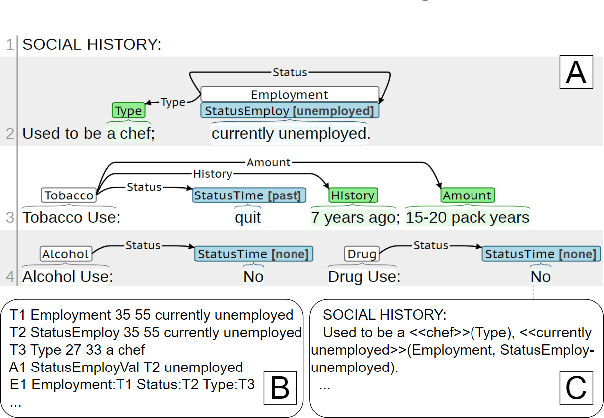
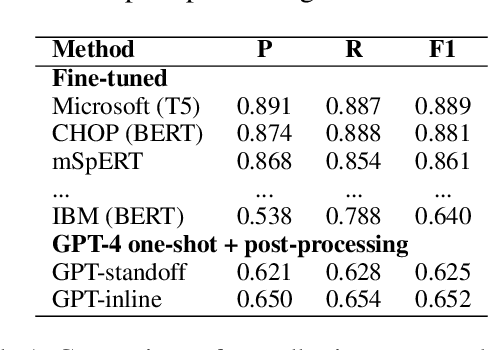
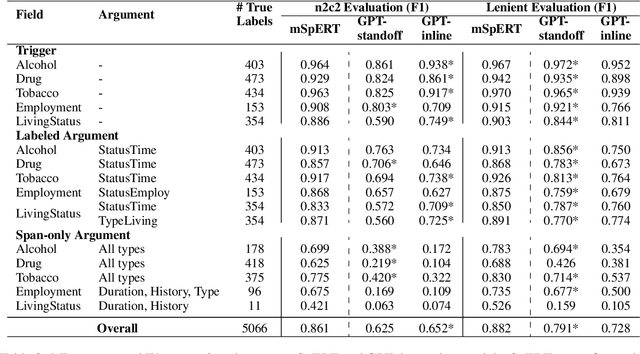
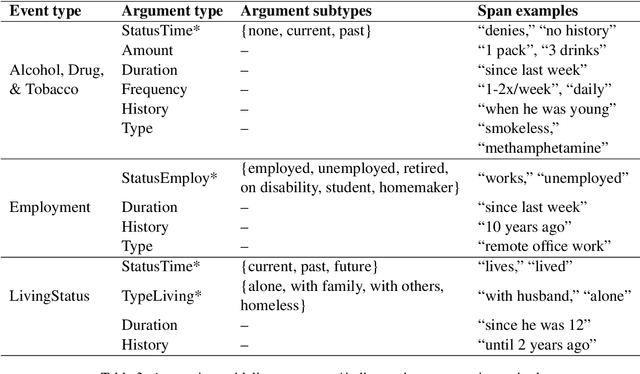
Abstract:Social determinants of health (SDOH) documented in the electronic health record through unstructured text are increasingly being studied to understand how SDOH impacts patient health outcomes. In this work, we utilize the Social History Annotation Corpus (SHAC), a multi-institutional corpus of de-identified social history sections annotated for SDOH, including substance use, employment, and living status information. We explore the automatic extraction of SDOH information with SHAC in both standoff and inline annotation formats using GPT-4 in a one-shot prompting setting. We compare GPT-4 extraction performance with a high-performing supervised approach and perform thorough error analyses. Our prompt-based GPT-4 method achieved an overall 0.652 F1 on the SHAC test set, similar to the 7th best-performing system among all teams in the n2c2 challenge with SHAC.
MasonNLP+ at SemEval-2023 Task 8: Extracting Medical Questions, Experiences and Claims from Social Media using Knowledge-Augmented Pre-trained Language Models
Apr 26, 2023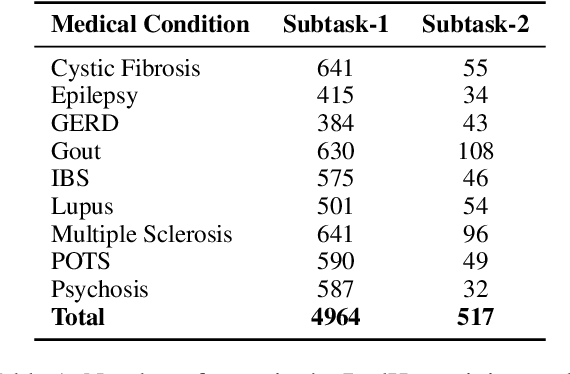



Abstract:In online forums like Reddit, users share their experiences with medical conditions and treatments, including making claims, asking questions, and discussing the effects of treatments on their health. Building systems to understand this information can effectively monitor the spread of misinformation and verify user claims. The Task-8 of the 2023 International Workshop on Semantic Evaluation focused on medical applications, specifically extracting patient experience- and medical condition-related entities from user posts on social media. The Reddit Health Online Talk (RedHot) corpus contains posts from medical condition-related subreddits with annotations characterizing the patient experience and medical conditions. In Subtask-1, patient experience is characterized by personal experience, questions, and claims. In Subtask-2, medical conditions are characterized by population, intervention, and outcome. For the automatic extraction of patient experiences and medical condition information, as a part of the challenge, we proposed language-model-based extraction systems that ranked $3^{rd}$ on both subtasks' leaderboards. In this work, we describe our approach and, in addition, explore the automatic extraction of this information using domain-specific language models and the inclusion of external knowledge.
Extracting Medication Changes in Clinical Narratives using Pre-trained Language Models
Aug 17, 2022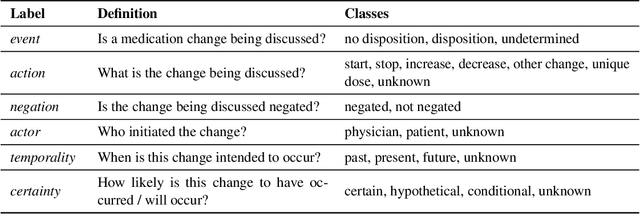
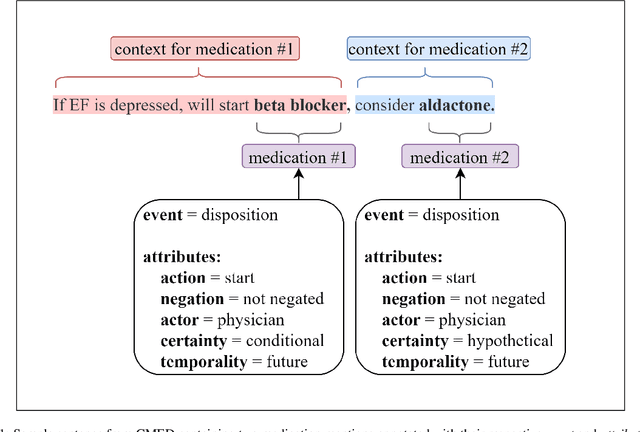
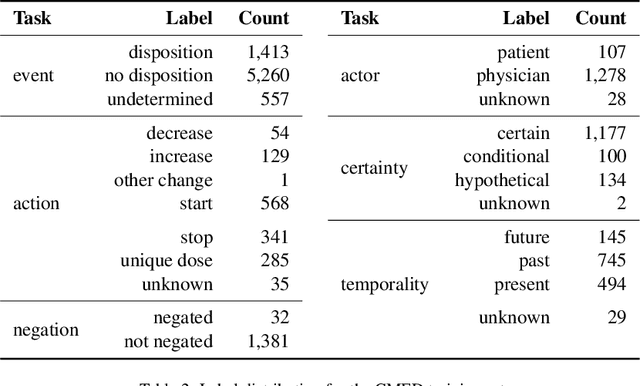
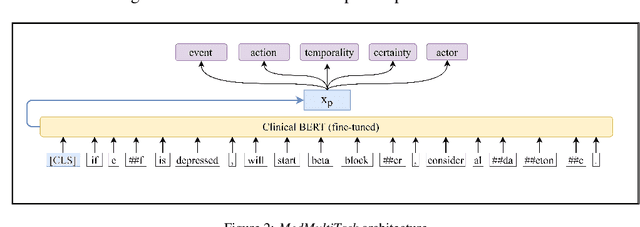
Abstract:An accurate and detailed account of patient medications, including medication changes within the patient timeline, is essential for healthcare providers to provide appropriate patient care. Healthcare providers or the patients themselves may initiate changes to patient medication. Medication changes take many forms, including prescribed medication and associated dosage modification. These changes provide information about the overall health of the patient and the rationale that led to the current care. Future care can then build on the resulting state of the patient. This work explores the automatic extraction of medication change information from free-text clinical notes. The Contextual Medication Event Dataset (CMED) is a corpus of clinical notes with annotations that characterize medication changes through multiple change-related attributes, including the type of change (start, stop, increase, etc.), initiator of the change, temporality, change likelihood, and negation. Using CMED, we identify medication mentions in clinical text and propose three novel high-performing BERT-based systems that resolve the annotated medication change characteristics. We demonstrate that our proposed architectures improve medication change classification performance over the initial work exploring CMED. We identify medication mentions with high performance at 0.959 F1, and our proposed systems classify medication changes and their attributes at an overall average of 0.827 F1.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge