George Hripcsak
CEHR-GPT: Generating Electronic Health Records with Chronological Patient Timelines
Feb 06, 2024



Abstract:Synthetic Electronic Health Records (EHR) have emerged as a pivotal tool in advancing healthcare applications and machine learning models, particularly for researchers without direct access to healthcare data. Although existing methods, like rule-based approaches and generative adversarial networks (GANs), generate synthetic data that resembles real-world EHR data, these methods often use a tabular format, disregarding temporal dependencies in patient histories and limiting data replication. Recently, there has been a growing interest in leveraging Generative Pre-trained Transformers (GPT) for EHR data. This enables applications like disease progression analysis, population estimation, counterfactual reasoning, and synthetic data generation. In this work, we focus on synthetic data generation and demonstrate the capability of training a GPT model using a particular patient representation derived from CEHR-BERT, enabling us to generate patient sequences that can be seamlessly converted to the Observational Medical Outcomes Partnership (OMOP) data format.
An Open-Source Knowledge Graph Ecosystem for the Life Sciences
Jul 11, 2023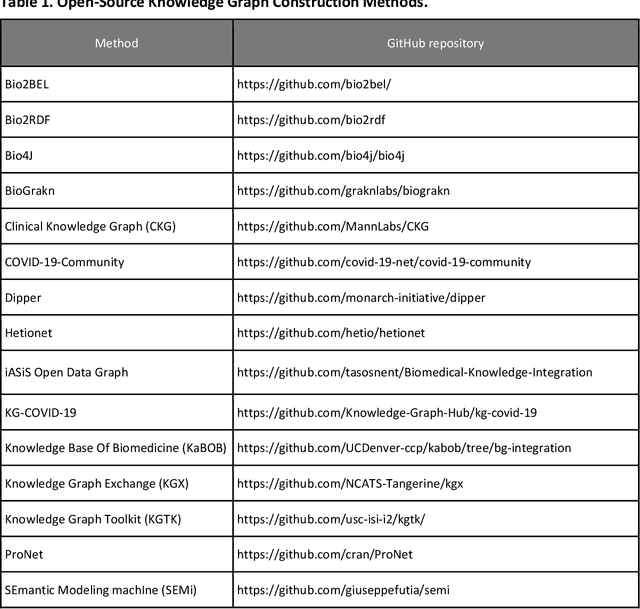
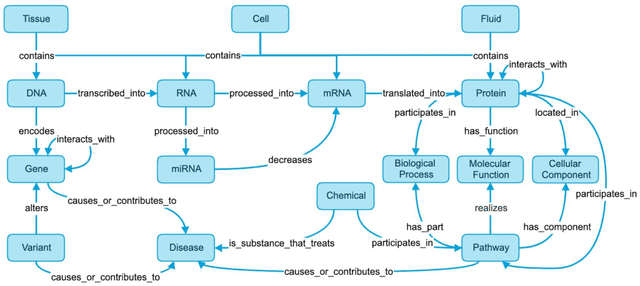
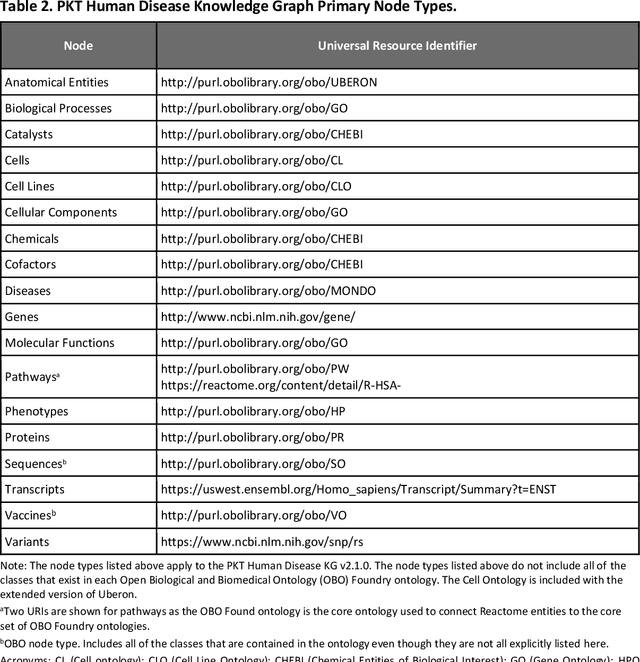
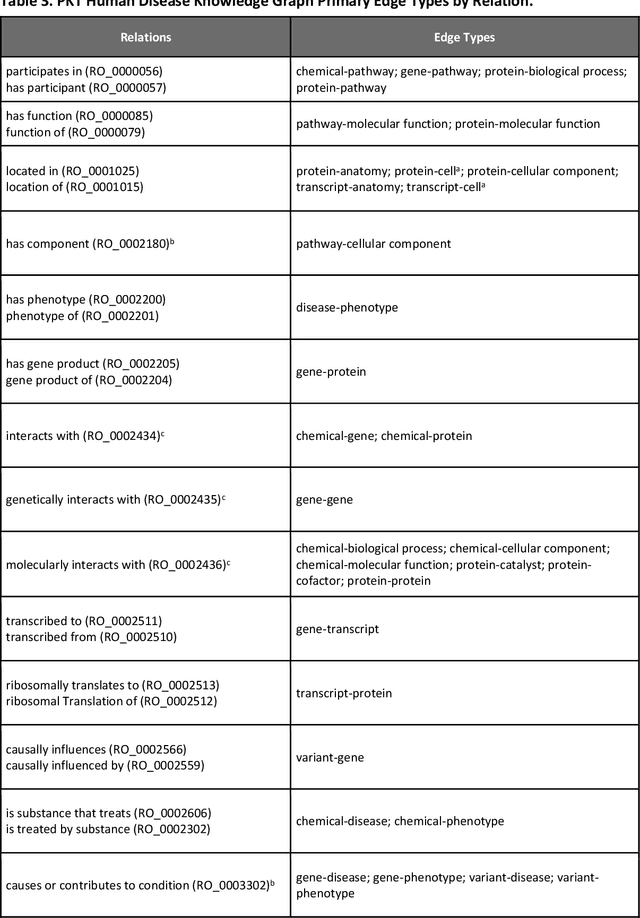
Abstract:Translational research requires data at multiple scales of biological organization. Advancements in sequencing and multi-omics technologies have increased the availability of these data but researchers face significant integration challenges. Knowledge graphs (KGs) are used to model complex phenomena, and methods exist to automatically construct them. However, tackling complex biomedical integration problems requires flexibility in the way knowledge is modeled. Moreover, existing KG construction methods provide robust tooling at the cost of fixed or limited choices among knowledge representation models. PheKnowLator (Phenotype Knowledge Translator) is a semantic ecosystem for automating the FAIR (Findable, Accessible, Interoperable, and Reusable) construction of ontologically grounded KGs with fully customizable knowledge representation. The ecosystem includes KG construction resources (e.g., data preparation APIs), analysis tools (e.g., SPARQL endpoints and abstraction algorithms), and benchmarks (e.g., prebuilt KGs and embeddings). We evaluate the ecosystem by surveying open-source KG construction methods and analyzing its computational performance when constructing 12 large-scale KGs. With flexible knowledge representation, PheKnowLator enables fully customizable KGs without compromising performance or usability.
A Bayesian Causal Inference Approach for Assessing Fairness in Clinical Decision-Making
Nov 21, 2022



Abstract:Fairness in clinical decision-making is a critical element of health equity, but assessing fairness of clinical decisions from observational data is challenging. Recently, many fairness notions have been proposed to quantify fairness in decision-making, among which causality-based fairness notions have gained increasing attention due to its potential in adjusting for confounding and reasoning about bias. However, causal fairness notions remain under-explored in the context of clinical decision-making with large-scale healthcare data. In this work, we propose a Bayesian causal inference approach for assessing a causal fairness notion called principal fairness in clinical settings. We demonstrate our approach using both simulated data and electronic health records (EHR) data.
Ontologizing Health Systems Data at Scale: Making Translational Discovery a Reality
Sep 10, 2022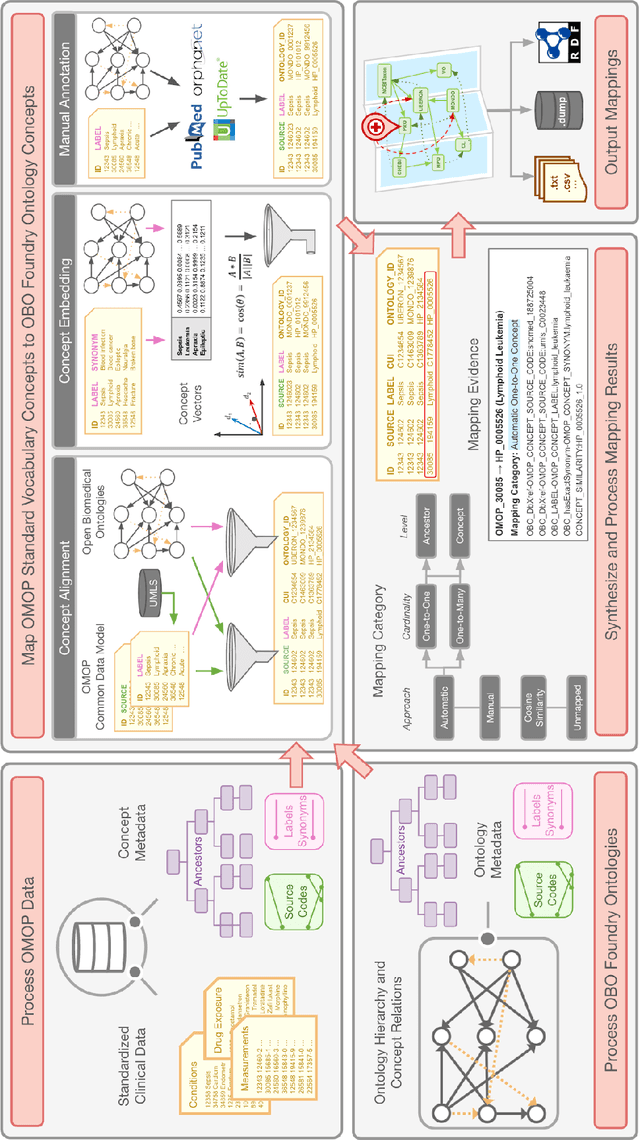
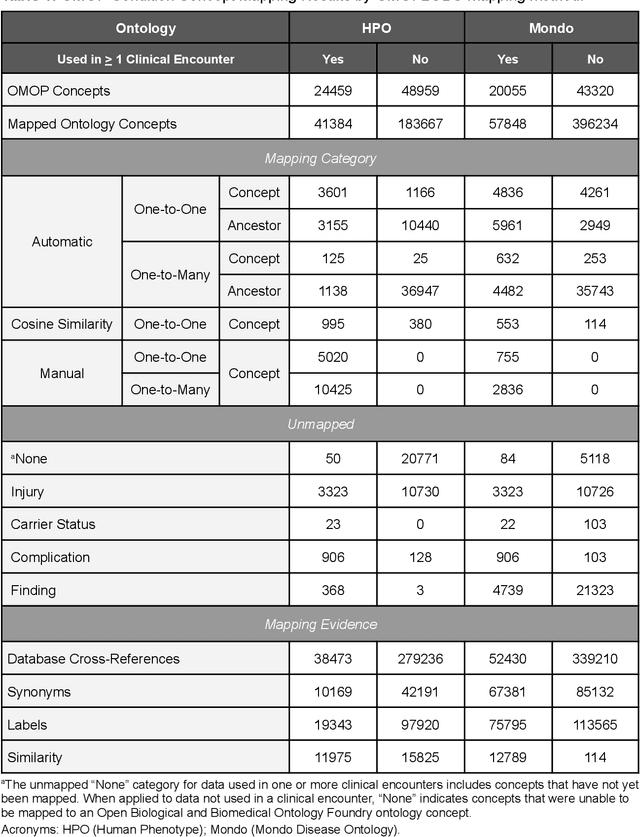
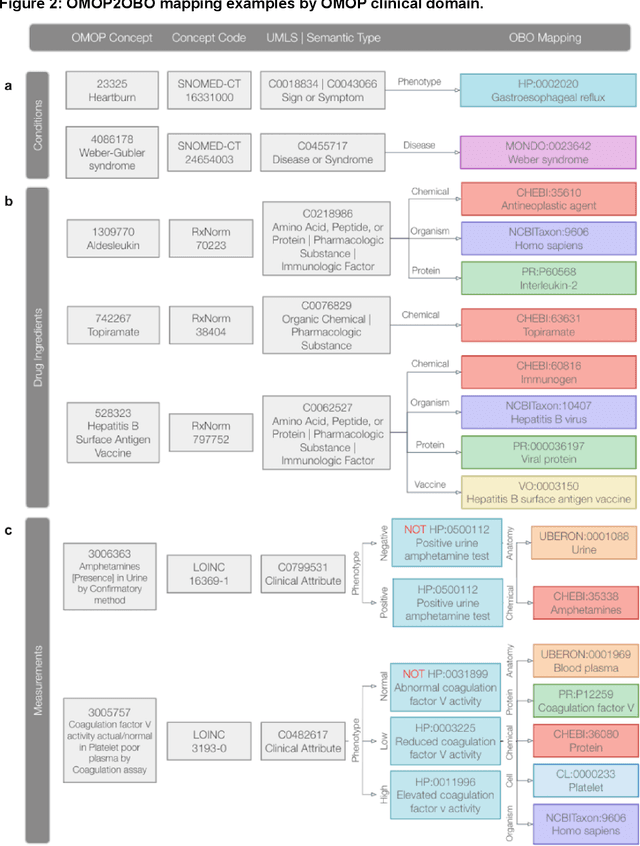
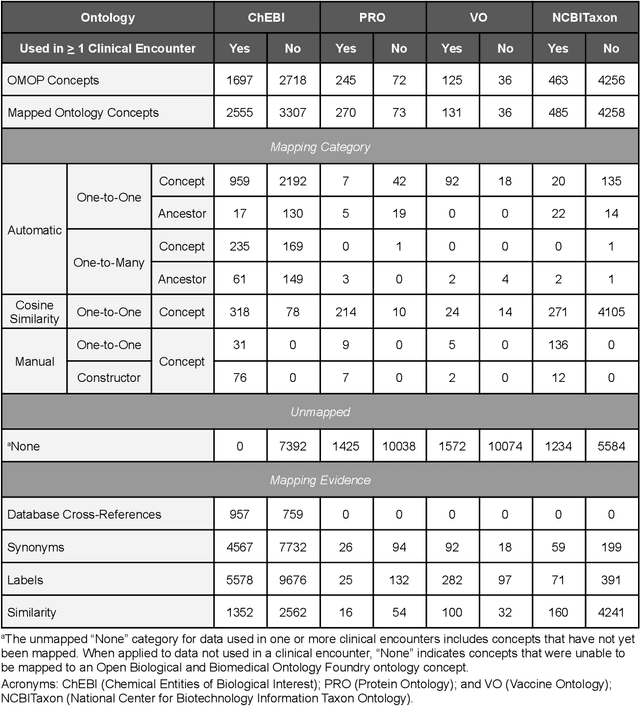
Abstract:Common data models solve many challenges of standardizing electronic health record (EHR) data, but are unable to semantically integrate the resources needed for deep phenotyping. Open Biological and Biomedical Ontology (OBO) Foundry ontologies provide semantically computable representations of biological knowledge and enable the integration of a variety of biomedical data. However, mapping EHR data to OBO Foundry ontologies requires significant manual curation and domain expertise. We introduce a framework for mapping Observational Medical Outcomes Partnership (OMOP) standard vocabularies to OBO Foundry ontologies. Using this framework, we produced mappings for 92,367 conditions, 8,615 drug ingredients, and 10,673 measurement results. Mapping accuracy was verified by domain experts and when examined across 24 hospitals, the mappings covered 99% of conditions and drug ingredients and 68% of measurements. Finally, we demonstrate that OMOP2OBO mappings can aid in the systematic identification of undiagnosed rare disease patients who might benefit from genetic testing.
The Medical Deconfounder: Assessing Treatment Effect with Electronic Health Records (EHRs)
Apr 03, 2019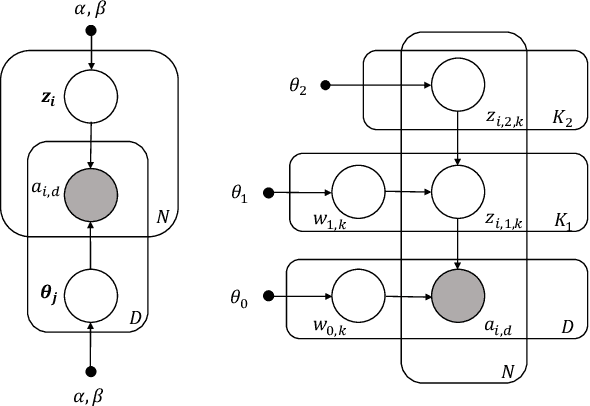
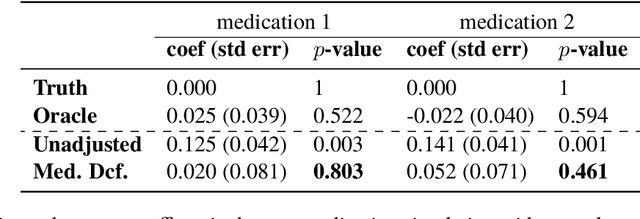
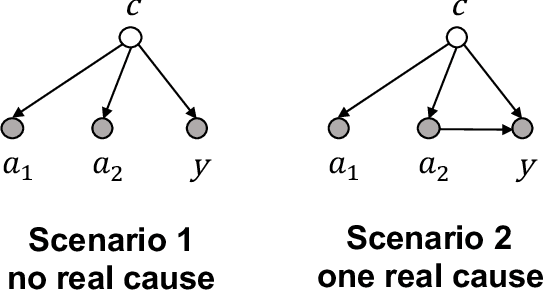
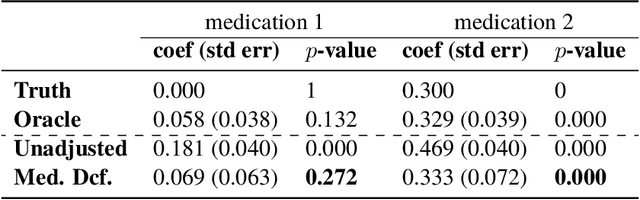
Abstract:Causal estimation of treatment effect has an important role in guiding physicians' decision process for drug prescription. While treatment effect is classically assessed with randomized controlled trials (RCTs), the availability of electronic health records (EHRs) bring an unprecedented opportunity for more efficient estimation. However, the presence of unobserved confounders makes treatment effect assessment from EHRs a challenging task. Confounders are the variables that affect both drug prescription and the patient's outcome; examples include a patient's gender, race, social economic status and comorbidities. When these confounders are unobserved, they bias the estimation. To adjust for unobserved confounders, we develop the medical deconfounder, a machine learning algorithm that unbiasedly estimates treatment effect from EHRs. The medical deconfounder first constructs a substitute confounder by modeling which drugs were prescribed to each patient; this substitute confounder is guaranteed to capture all multi-drug confounders, observed or unobserved (Wang and Blei, 2018). It then uses this substitute confounder to adjust for the confounding bias in the analysis. We validate the medical deconfounder on simulations and two medical data sets. The medical deconfounder produces closer-to-truth estimates in simulations and identifies effective medications that are more consistent with the findings reported in the medical literature compared to classical approaches.
Characterizing Design Patterns of EHR-Driven Phenotype Extraction Algorithms
Nov 15, 2018
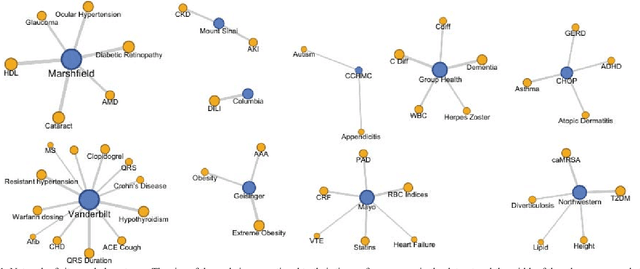
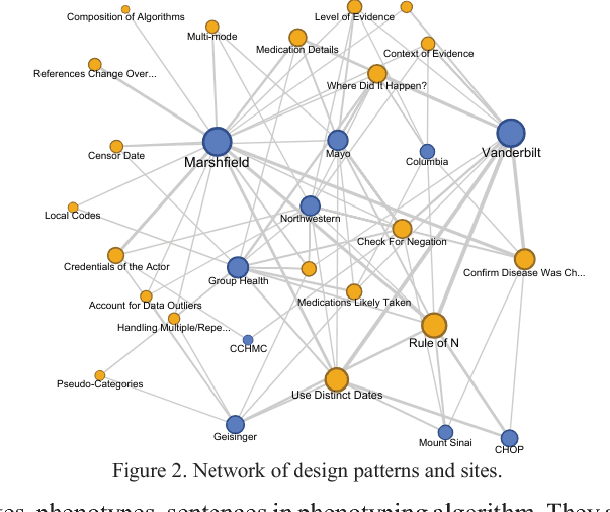
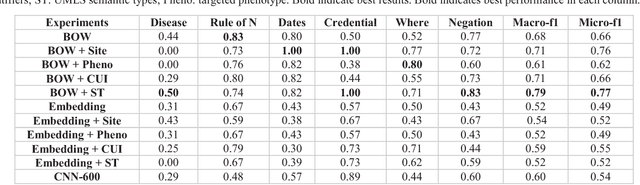
Abstract:The automatic development of phenotype algorithms from Electronic Health Record data with machine learning (ML) techniques is of great interest given the current practice is very time-consuming and resource intensive. The extraction of design patterns from phenotype algorithms is essential to understand their rationale and standard, with great potential to automate the development process. In this pilot study, we perform network visualization on the design patterns and their associations with phenotypes and sites. We classify design patterns using the fragments from previously annotated phenotype algorithms as the ground truth. The classification performance is used as a proxy for coherence at the attribution level. The bag-of-words representation with knowledge-based features generated a good performance in the classification task (0.79 macro-f1 scores). Good classification accuracy with simple features demonstrated the attribution coherence and the feasibility of automatic identification of design patterns. Our results point to both the feasibility and challenges of automatic identification of phenotyping design patterns, which would power the automatic development of phenotype algorithms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge