Fakrul Islam Tushar
iTRIALSPACE: Programmable Virtual Lesion Trials for Controlled Evaluation of Lung CT Models
May 07, 2026Abstract:We introduce iTRIALSPACE, a programmable evaluation framework for controlled assessment of lung CT models. Standard benchmarks are static retrospective collections that entangle lesion size, lobe prevalence, anatomy, and acquisition context, making it difficult to determine what structurally drives model accuracy. iTRIALSPACE addresses this limitation by composing real clinical CTs and lesion profiles into controlled virtual lesion trials through a four-stage pipeline: multidataset nodule profiling, explicit trial specification, anatomy-aware mask insertion, and ControlNet-conditioned CT synthesis. The framework is built on a unified 54-attribute nodule-profile dataset spanning 13,140 annotated nodules from seven public CT sources and instantiated as 13 trial modes. We evaluate iTRIALSPACE in a 55,469-sample Virtual Lesion Study spanning three medical VLMs, four spatialguidance conditions, and three clinical tasks. Across all 13 modes, the synthetic substrate remains within the real-to-real FID baseline, and synthetic performance rankings transfer strongly to real clinical data ($ρ$ = 0.93, p < 10$^{-15}$). Controlled trial modes expose findings unavailable to fixed-distribution benchmarks, including shortcut-driven size prediction collapse under lobe-equalized sampling and hostto-donor variance ratios of 8.9x and 3.3x in twin-cross analysis. These results position iTRIALSPACE as an auditable evaluation infrastructure for controlled, falsifiable testing beyond static retrospective benchmarks.
Tri-Reader: An Open-Access, Multi-Stage AI Pipeline for First-Pass Lung Nodule Annotation in Screening CT
Jan 27, 2026Abstract:Using multiple open-access models trained on public datasets, we developed Tri-Reader, a comprehensive, freely available pipeline that integrates lung segmentation, nodule detection, and malignancy classification into a unified tri-stage workflow. The pipeline is designed to prioritize sensitivity while reducing the candidate burden for annotators. To ensure accuracy and generalizability across diverse practices, we evaluated Tri-Reader on multiple internal and external datasets as compared with expert annotations and dataset-provided reference standards.
NodMAISI: Nodule-Oriented Medical AI for Synthetic Imaging
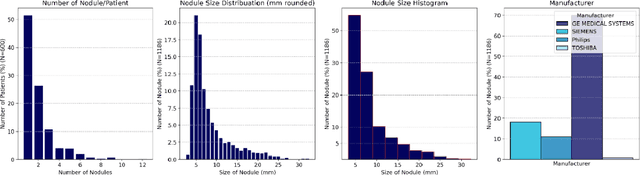
Dec 19, 2025Abstract:Objective: Although medical imaging datasets are increasingly available, abnormal and annotation-intensive findings critical to lung cancer screening, particularly small pulmonary nodules, remain underrepresented and inconsistently curated. Methods: We introduce NodMAISI, an anatomically constrained, nodule-oriented CT synthesis and augmentation framework trained on a unified multi-source cohort (7,042 patients, 8,841 CTs, 14,444 nodules). The framework integrates: (i) a standardized curation and annotation pipeline linking each CT with organ masks and nodule-level annotations, (ii) a ControlNet-conditioned rectified-flow generator built on MAISI-v2's foundational blocks to enforce anatomy- and lesion-consistent synthesis, and (iii) lesion-aware augmentation that perturbs nodule masks (controlled shrinkage) while preserving surrounding anatomy to generate paired CT variants. Results: Across six public test datasets, NodMAISI improved distributional fidelity relative to MAISI-v2 (real-to-synthetic FID range 1.18 to 2.99 vs 1.69 to 5.21). In lesion detectability analysis using a MONAI nodule detector, NodMAISI substantially increased average sensitivity and more closely matched clinical scans (IMD-CT: 0.69 vs 0.39; DLCS24: 0.63 vs 0.20), with the largest gains for sub-centimeter nodules where MAISI-v2 frequently failed to reproduce the conditioned lesion. In downstream nodule-level malignancy classification trained on LUNA25 and externally evaluated on LUNA16, LNDbv4, and DLCS24, NodMAISI augmentation improved AUC by 0.07 to 0.11 at <=20% clinical data and by 0.12 to 0.21 at 10%, consistently narrowing the performance gap under data scarcity.
SYN-LUNGS: Towards Simulating Lung Nodules with Anatomy-Informed Digital Twins for AI Training
Feb 28, 2025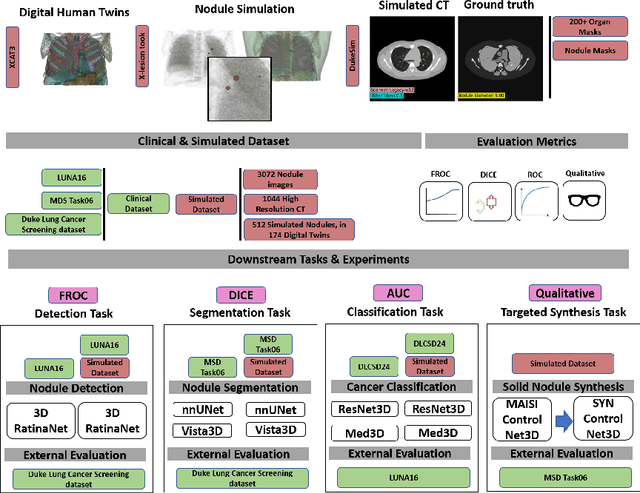
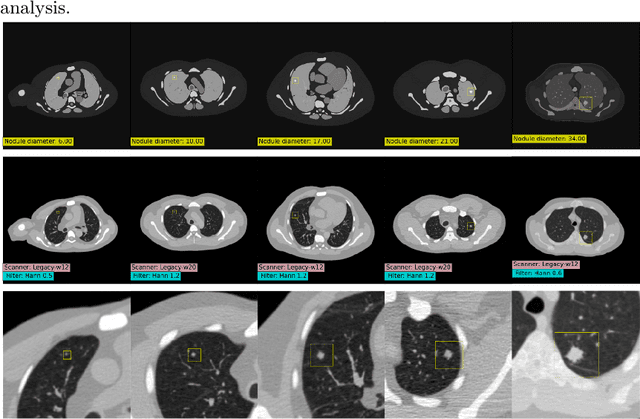
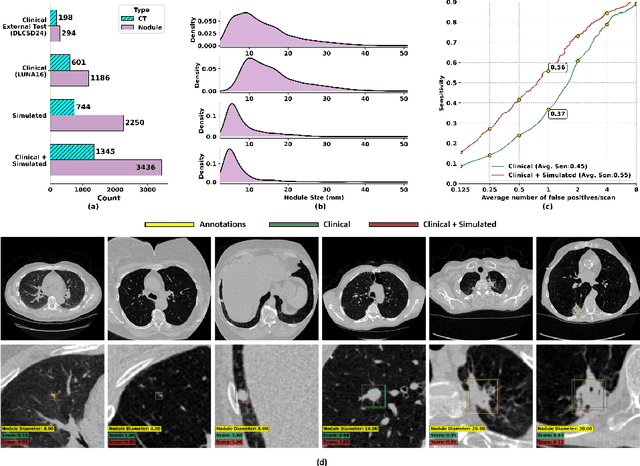
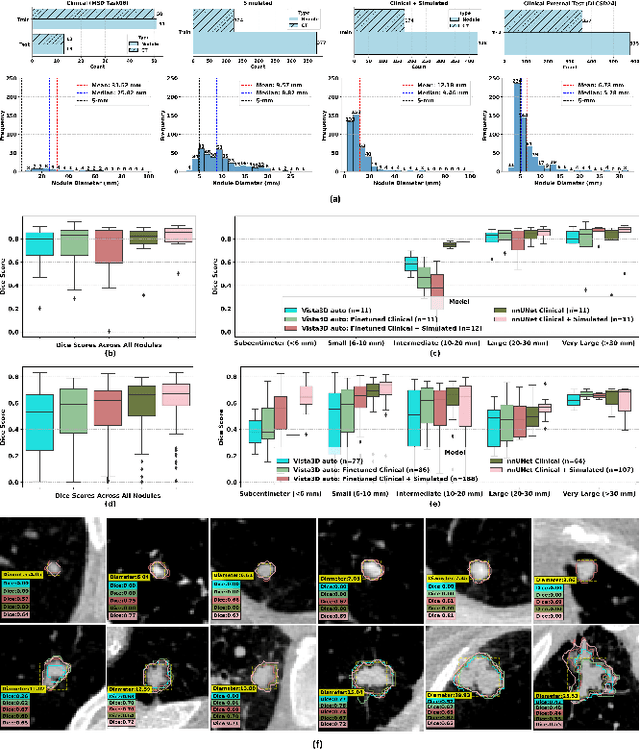
Abstract:AI models for lung cancer screening are limited by data scarcity, impacting generalizability and clinical applicability. Generative models address this issue but are constrained by training data variability. We introduce SYN-LUNGS, a framework for generating high-quality 3D CT images with detailed annotations. SYN-LUNGS integrates XCAT3 phantoms for digital twin generation, X-Lesions for nodule simulation (varying size, location, and appearance), and DukeSim for CT image formation with vendor and parameter variability. The dataset includes 3,072 nodule images from 1,044 simulated CT scans, with 512 lesions and 174 digital twins. Models trained on clinical + simulated data outperform clinical only models, achieving 10% improvement in detection, 2-9% in segmentation and classification, and enhanced synthesis.By incorporating anatomy-informed simulations, SYN-LUNGS provides a scalable approach for AI model development, particularly in rare disease representation and improving model reliability.
Refining Focus in AI for Lung Cancer: Comparing Lesion-Centric and Chest-Region Models with Performance Insights from Internal and External Validation
Nov 25, 2024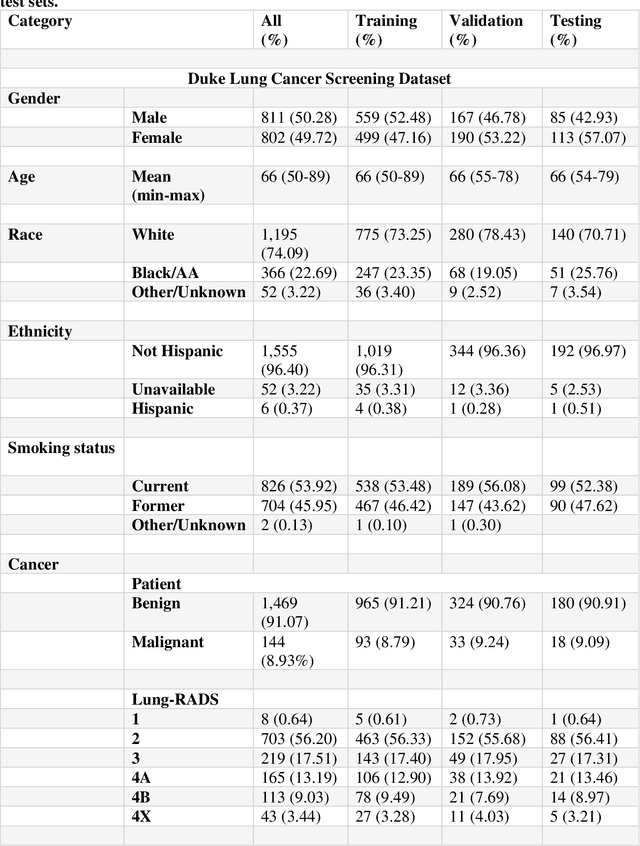

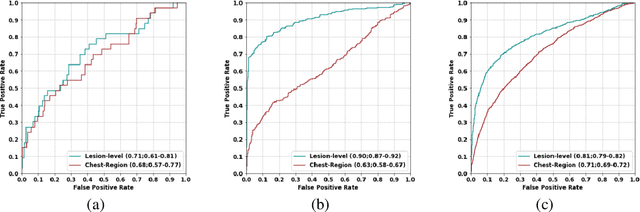

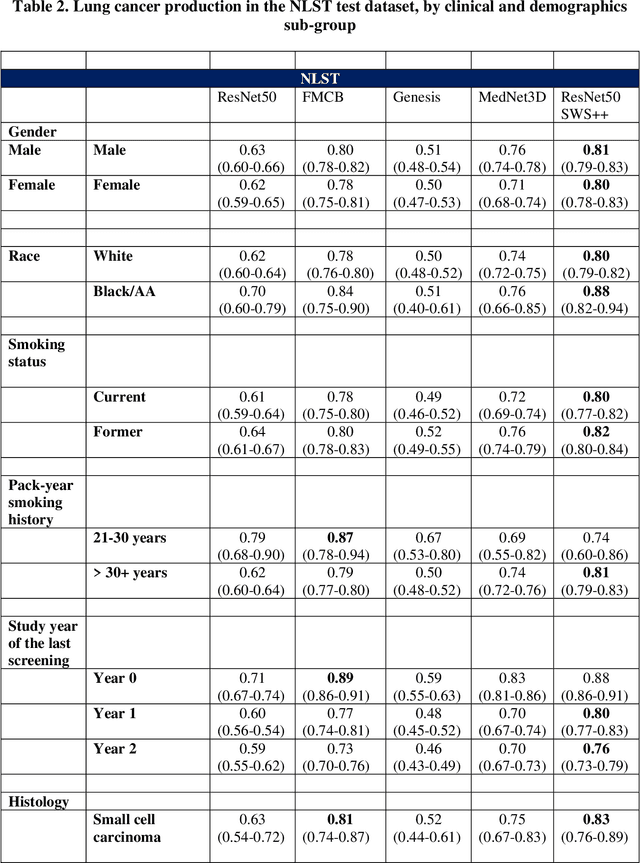
Abstract:Background: AI-based classification models are essential for improving lung cancer diagnosis. However, the relative performance of lesion-level versus chest-region models in internal and external datasets remains unclear. Purpose: This study evaluates the performance of lesion-level and chest-region models for lung cancer classification, comparing their effectiveness across internal Duke Lung Nodule Dataset 2024 (DLND24) and external (LUNA16, NLST) datasets, with a focus on subgroup analyses by demographics, histology, and imaging characteristics. Materials and Methods: Two AI models were trained: one using lesion-centric patches (64,64,64) and the other using chest-region patches (512,512,8). Internal validation was conducted on DLND24, while external validation utilized LUNA16 and NLST datasets. The models performances were assessed using AUC-ROC, with subgroup analyses for demographic, clinical, and imaging factors. Statistical comparisons were performed using DeLongs test. Gradient-based visualizations and probability distribution were further used for analysis. Results: The lesion-level model consistently outperformed the chest-region model across datasets. In internal validation, the lesion-level model achieved an AUC of 0.71(CI: 0.61-0.81), compared to 0.68(0.57-0.77) for the chest-region model. External validation showed similar trends, with AUCs of 0.90(0.87-0.92) and 0.81(0.79-0.82) on LUNA16 and NLST, respectively. Subgroup analyses revealed significant advantages for lesion-level models in certain histological subtypes (adenocarcinoma) and imaging conditions (CT manufacturers). Conclusion: Lesion-level models demonstrate superior classification performance, especially for external datasets and challenging subgroups, suggesting their clinical utility for precision lung cancer diagnostics.
Peritumoral Expansion Radiomics for Improved Lung Cancer Classification
Nov 24, 2024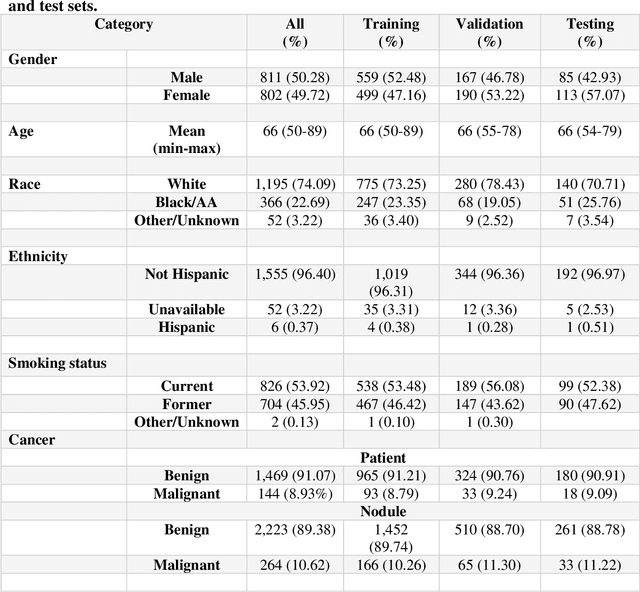
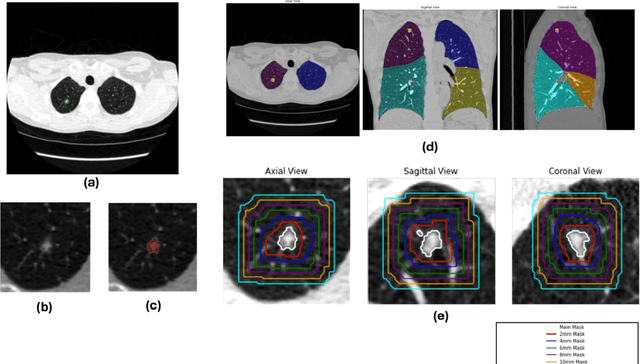
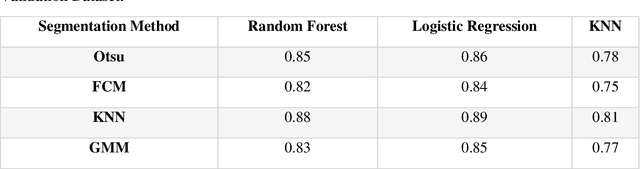
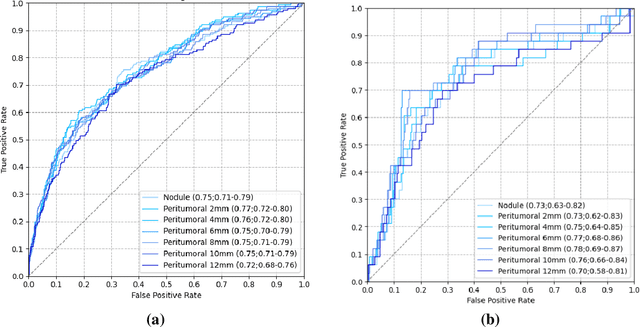
Abstract:Purpose: This study investigated how nodule segmentation and surrounding peritumoral regions influence radionics-based lung cancer classification. Methods: Using 3D CT scans with bounding box annotated nodules, we generated 3D segmentations using four techniques: Otsu, Fuzzy C-Means (FCM), Gaussian Mixture Model (GMM), and K-Nearest Neighbors (KNN). Radiomics features were extracted using the PyRadiomics library, and multiple machine-learning-based classifiers, including Random Forest, Logistic Regression, and KNN, were employed to classify nodules as cancerous or non-cancerous. The best-performing segmentation and model were further analyzed by expanding the initial nodule segmentation into the peritumoral region (2, 4, 6, 8, 10, and 12 mm) to understand the influence of the surrounding area on classification. Additionally, we compared our results to deep learning-based feature extractors Foundation Model for Cancer Biomarkers (FMCB) and other state-of-the-art baseline models. Results: Incorporating peritumoral regions significantly enhanced performance, with the best result obtained at 8 mm expansion (AUC = 0.78). Compared to image-based deep learning models, such as FMCB (AUC = 0.71) and ResNet50-SWS++ (AUC = 0.71), our radiomics-based approach demonstrated superior classification accuracy. Conclusion: The study highlights the importance of peritumoral expansion in improving lung cancer classification using radiomics. These findings can inform the development of more robust AI-driven diagnostic tools.
XCAT-2.0: A Comprehensive Library of Personalized Digital Twins Derived from CT Scans
May 18, 2024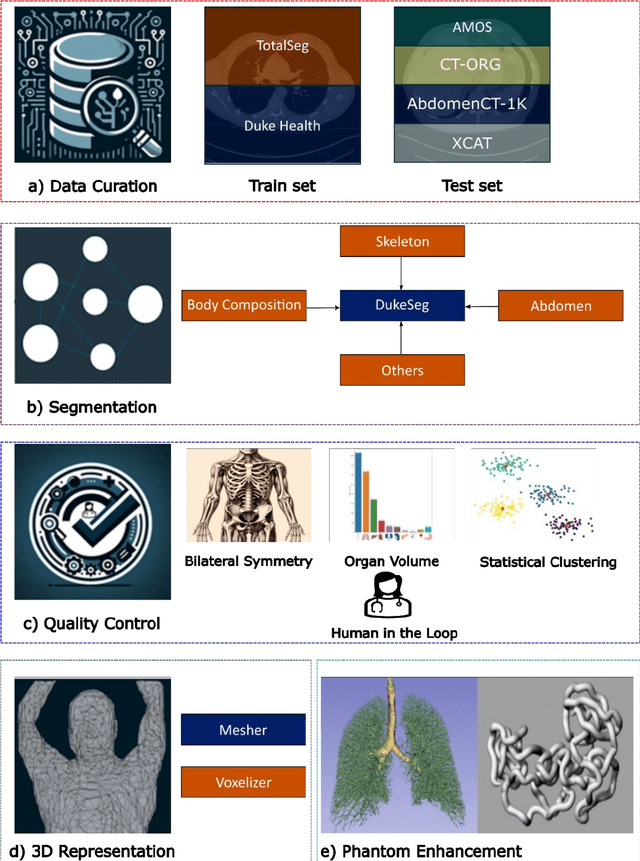
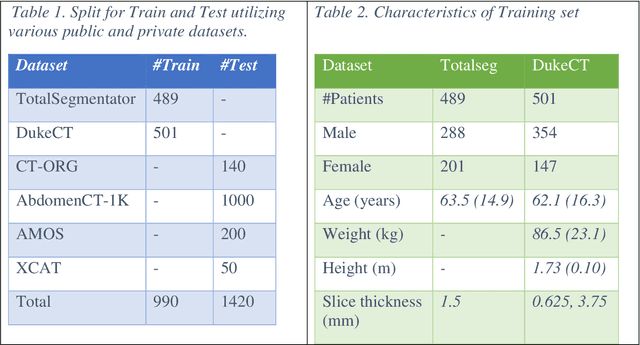
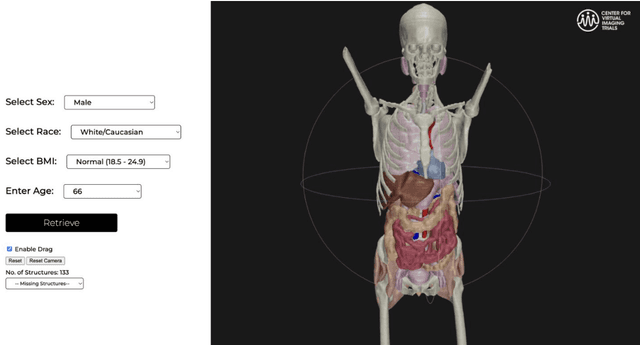
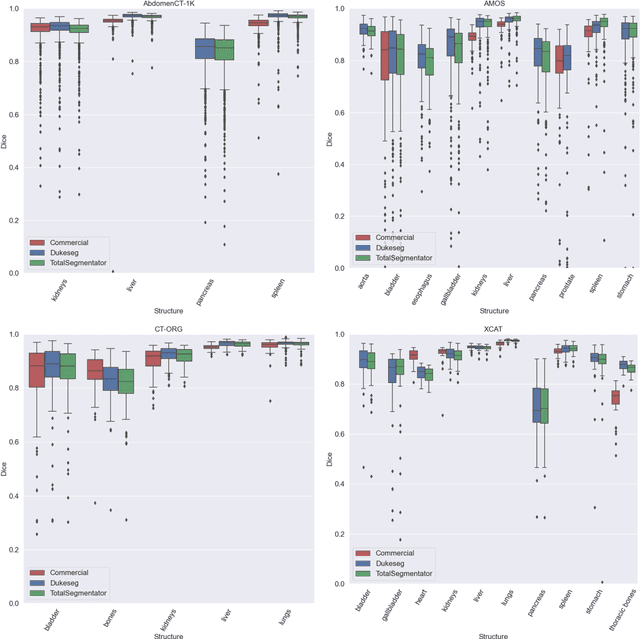
Abstract:Virtual Imaging Trials (VIT) offer a cost-effective and scalable approach for evaluating medical imaging technologies. Computational phantoms, which mimic real patient anatomy and physiology, play a central role in VIT. However, the current libraries of computational phantoms face limitations, particularly in terms of sample size and diversity. Insufficient representation of the population hampers accurate assessment of imaging technologies across different patient groups. Traditionally, phantoms were created by manual segmentation, which is a laborious and time-consuming task, impeding the expansion of phantom libraries. This study presents a framework for realistic computational phantom modeling using a suite of four deep learning segmentation models, followed by three forms of automated organ segmentation quality control. Over 2500 computational phantoms with up to 140 structures illustrating a sophisticated approach to detailed anatomical modeling are released. Phantoms are available in both voxelized and surface mesh formats. The framework is aggregated with an in-house CT scanner simulator to produce realistic CT images. The framework can potentially advance virtual imaging trials, facilitating comprehensive and reliable evaluations of medical imaging technologies. Phantoms may be requested at https://cvit.duke.edu/resources/, code, model weights, and sample CT images are available at https://xcat-2.github.io.
AI in Lung Health: Benchmarking Detection and Diagnostic Models Across Multiple CT Scan Datasets
May 07, 2024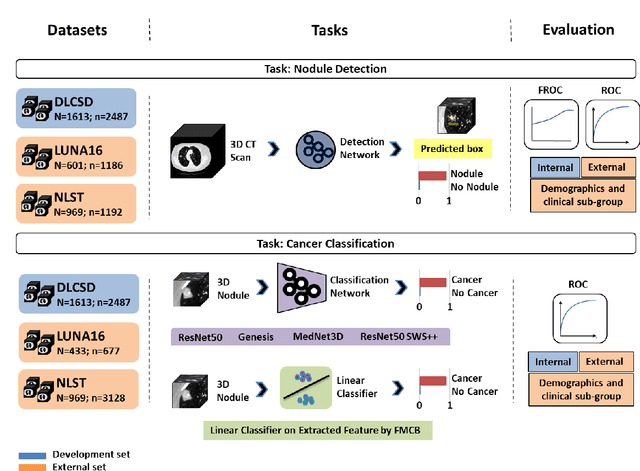
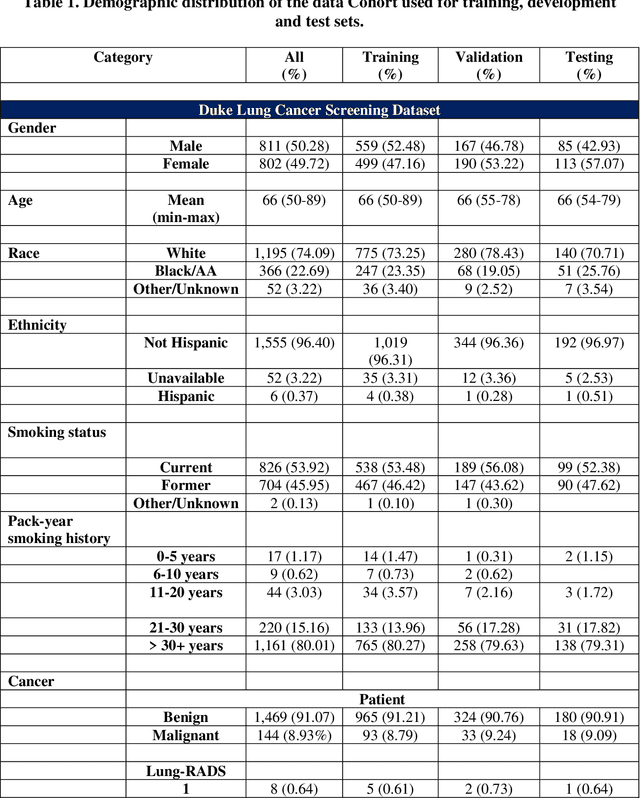
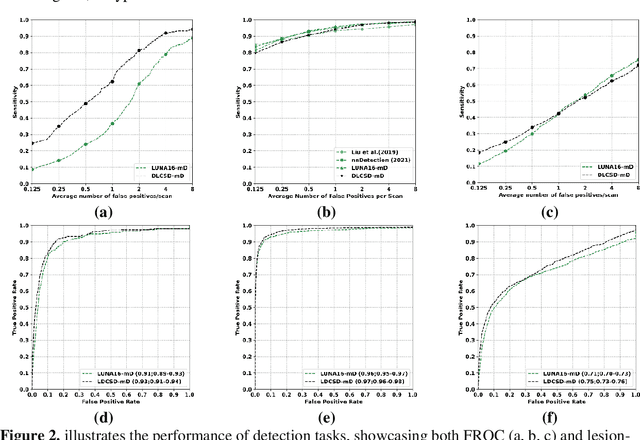
Abstract:BACKGROUND: Lung cancer's high mortality rate can be mitigated by early detection, which is increasingly reliant on artificial intelligence (AI) for diagnostic imaging. However, the performance of AI models is contingent upon the datasets used for their training and validation. METHODS: This study developed and validated the DLCSD-mD and LUNA16-mD models utilizing the Duke Lung Cancer Screening Dataset (DLCSD), encompassing over 2,000 CT scans with more than 3,000 annotations. These models were rigorously evaluated against the internal DLCSD and external LUNA16 and NLST datasets, aiming to establish a benchmark for imaging-based performance. The assessment focused on creating a standardized evaluation framework to facilitate consistent comparison with widely utilized datasets, ensuring a comprehensive validation of the model's efficacy. Diagnostic accuracy was assessed using free-response receiver operating characteristic (FROC) and area under the curve (AUC) analyses. RESULTS: On the internal DLCSD set, the DLCSD-mD model achieved an AUC of 0.93 (95% CI:0.91-0.94), demonstrating high accuracy. Its performance was sustained on the external datasets, with AUCs of 0.97 (95% CI: 0.96-0.98) on LUNA16 and 0.75 (95% CI: 0.73-0.76) on NLST. Similarly, the LUNA16-mD model recorded an AUC of 0.96 (95% CI: 0.95-0.97) on its native dataset and showed transferable diagnostic performance with AUCs of 0.91 (95% CI: 0.89-0.93) on DLCSD and 0.71 (95% CI: 0.70-0.72) on NLST. CONCLUSION: The DLCSD-mD model exhibits reliable performance across different datasets, establishing the DLCSD as a robust benchmark for lung cancer detection and diagnosis. Through the provision of our models and code to the public domain, we aim to accelerate the development of AI-based diagnostic tools and encourage reproducibility and collaborative advancements within the medical machine-learning (ML) field.
VLST: Virtual Lung Screening Trial for Lung Cancer Detection Using Virtual Imaging Trial
Apr 17, 2024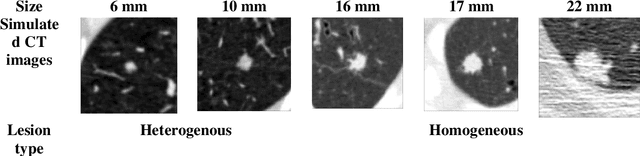
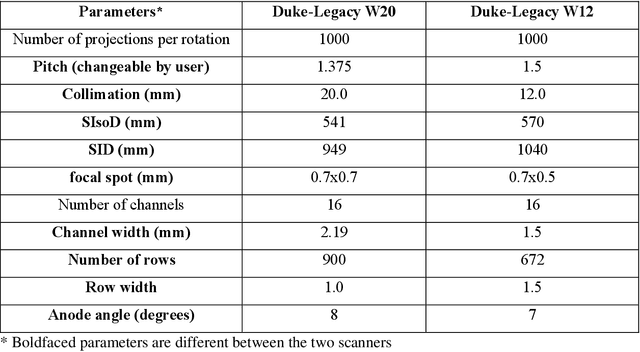

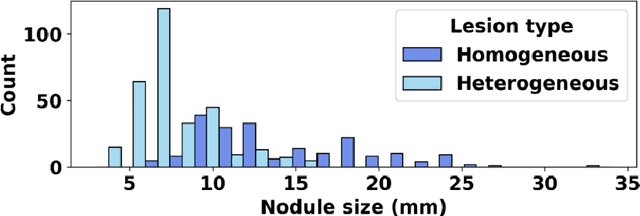
Abstract:Importance: The efficacy of lung cancer screening can be significantly impacted by the imaging modality used. This Virtual Lung Screening Trial (VLST) addresses the critical need for precision in lung cancer diagnostics and the potential for reducing unnecessary radiation exposure in clinical settings. Objectives: To establish a virtual imaging trial (VIT) platform that accurately simulates real-world lung screening trials (LSTs) to assess the diagnostic accuracy of CT and CXR modalities. Design, Setting, and Participants: Utilizing computational models and machine learning algorithms, we created a diverse virtual patient population. The cohort, designed to mirror real-world demographics, was assessed using virtual imaging techniques that reflect historical imaging technologies. Main Outcomes and Measures: The primary outcome was the difference in the Area Under the Curve (AUC) for CT and CXR modalities across lesion types and sizes. Results: The study analyzed 298 CT and 313 CXR simulated images from 313 virtual patients, with a lesion-level AUC of 0.81 (95% CI: 0.78-0.84) for CT and 0.55 (95% CI: 0.53-0.56) for CXR. At the patient level, CT demonstrated an AUC of 0.85 (95% CI: 0.80-0.89), compared to 0.53 (95% CI: 0.47-0.60) for CXR. Subgroup analyses indicated CT's superior performance in detecting homogeneous lesions (AUC of 0.97 for lesion-level) and heterogeneous lesions (AUC of 0.71 for lesion-level) as well as in identifying larger nodules (AUC of 0.98 for nodules > 8 mm). Conclusion and Relevance: The VIT platform validated the superior diagnostic accuracy of CT over CXR, especially for smaller nodules, underscoring its potential to replicate real clinical imaging trials. These findings advocate for the integration of virtual trials in the evaluation and improvement of imaging-based diagnostic tools.
What limits performance of weakly supervised deep learning for chest CT classification?
Feb 06, 2024



Abstract:Weakly supervised learning with noisy data has drawn attention in the medical imaging community due to the sparsity of high-quality disease labels. However, little is known about the limitations of such weakly supervised learning and the effect of these constraints on disease classification performance. In this paper, we test the effects of such weak supervision by examining model tolerance for three conditions. First, we examined model tolerance for noisy data by incrementally increasing error in the labels within the training data. Second, we assessed the impact of dataset size by varying the amount of training data. Third, we compared performance differences between binary and multi-label classification. Results demonstrated that the model could endure up to 10% added label error before experiencing a decline in disease classification performance. Disease classification performance steadily rose as the amount of training data was increased for all disease classes, before experiencing a plateau in performance at 75% of training data. Last, the binary model outperformed the multilabel model in every disease category. However, such interpretations may be misleading, as the binary model was heavily influenced by co-occurring diseases and may not have learned the specific features of the disease in the image. In conclusion, this study may help the medical imaging community understand the benefits and risks of weak supervision with noisy labels. Such studies demonstrate the need to build diverse, large-scale datasets and to develop explainable and responsible AI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge