William P. Segars
Modality-agnostic, patient-specific digital twins modeling temporally varying digestive motion
Jul 03, 2025Abstract:Objective: Clinical implementation of deformable image registration (DIR) requires voxel-based spatial accuracy metrics such as manually identified landmarks, which are challenging to implement for highly mobile gastrointestinal (GI) organs. To address this, patient-specific digital twins (DT) modeling temporally varying motion were created to assess the accuracy of DIR methods. Approach: 21 motion phases simulating digestive GI motion as 4D sequences were generated from static 3D patient scans using published analytical GI motion models through a semi-automated pipeline. Eleven datasets, including six T2w FSE MRI (T2w MRI), two T1w 4D golden-angle stack-of-stars, and three contrast-enhanced CT scans. The motion amplitudes of the DTs were assessed against real patient stomach motion amplitudes extracted from independent 4D MRI datasets. The generated DTs were then used to assess six different DIR methods using target registration error, Dice similarity coefficient, and the 95th percentile Hausdorff distance using summary metrics and voxel-level granular visualizations. Finally, for a subset of T2w MRI scans from patients treated with MR-guided radiation therapy, dose distributions were warped and accumulated to assess dose warping errors, including evaluations of DIR performance in both low- and high-dose regions for patient-specific error estimation. Main results: Our proposed pipeline synthesized DTs modeling realistic GI motion, achieving mean and maximum motion amplitudes and a mean log Jacobian determinant within 0.8 mm and 0.01, respectively, similar to published real-patient gastric motion data. It also enables the extraction of detailed quantitative DIR performance metrics and rigorous validation of dose mapping accuracy. Significance: The pipeline enables rigorously testing DIR tools for dynamic, anatomically complex regions enabling granular spatial and dosimetric accuracies.
Quality or Quantity: Toward a Unified Approach for Multi-organ Segmentation in Body CT
Mar 03, 2022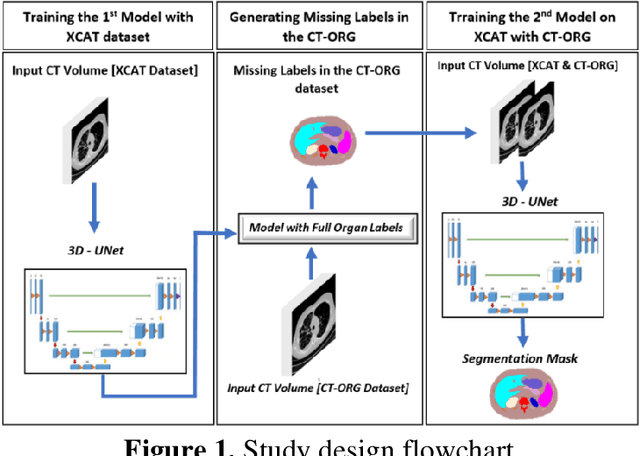
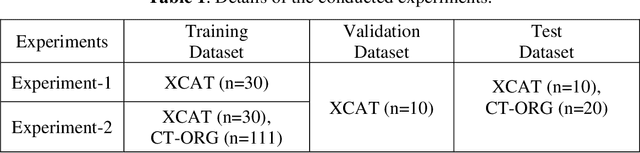
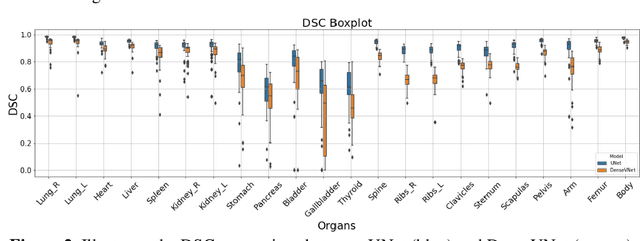
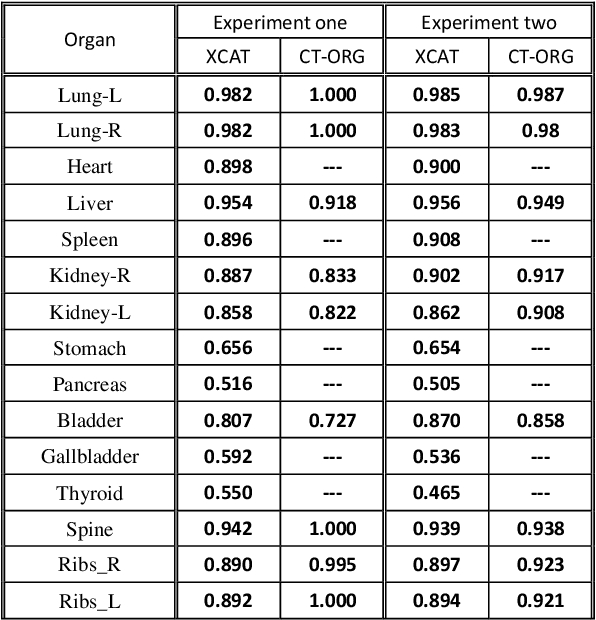
Abstract:Organ segmentation of medical images is a key step in virtual imaging trials. However, organ segmentation datasets are limited in terms of quality (because labels cover only a few organs) and quantity (since case numbers are limited). In this study, we explored the tradeoffs between quality and quantity. Our goal is to create a unified approach for multi-organ segmentation of body CT, which will facilitate the creation of large numbers of accurate virtual phantoms. Initially, we compared two segmentation architectures, 3D-Unet and DenseVNet, which were trained using XCAT data that is fully labeled with 22 organs, and chose the 3D-Unet as the better performing model. We used the XCAT-trained model to generate pseudo-labels for the CT-ORG dataset that has only 7 organs segmented. We performed two experiments: First, we trained 3D-UNet model on the XCAT dataset, representing quality data, and tested it on both XCAT and CT-ORG datasets. Second, we trained 3D-UNet after including the CT-ORG dataset into the training set to have more quantity. Performance improved for segmentation in the organs where we have true labels in both datasets and degraded when relying on pseudo-labels. When organs were labeled in both datasets, Exp-2 improved Average DSC in XCAT and CT-ORG by 1. This demonstrates that quality data is the key to improving the model's performance.
TransMorph: Transformer for unsupervised medical image registration
Nov 23, 2021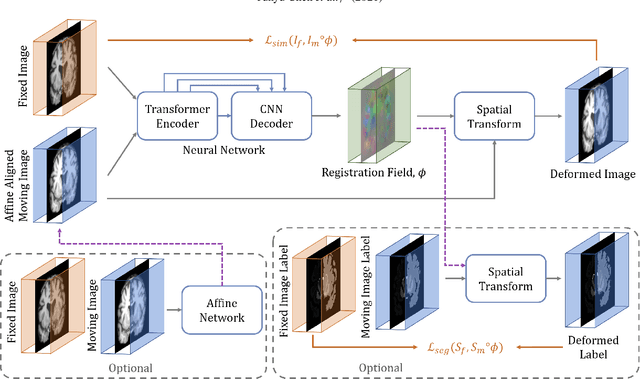

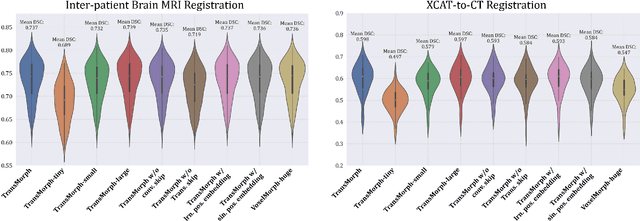
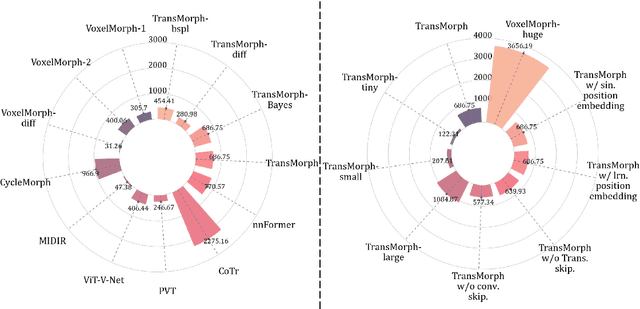
Abstract:In the last decade, convolutional neural networks (ConvNets) have dominated the field of medical image analysis. However, it is found that the performances of ConvNets may still be limited by their inability to model long-range spatial relations between voxels in an image. Numerous vision Transformers have been proposed recently to address the shortcomings of ConvNets, demonstrating state-of-the-art performances in many medical imaging applications. Transformers may be a strong candidate for image registration because their self-attention mechanism enables a more precise comprehension of the spatial correspondence between moving and fixed images. In this paper, we present TransMorph, a hybrid Transformer-ConvNet model for volumetric medical image registration. We also introduce three variants of TransMorph, with two diffeomorphic variants ensuring the topology-preserving deformations and a Bayesian variant producing a well-calibrated registration uncertainty estimate. The proposed models are extensively validated against a variety of existing registration methods and Transformer architectures using volumetric medical images from two applications: inter-patient brain MRI registration and phantom-to-CT registration. Qualitative and quantitative results demonstrate that TransMorph and its variants lead to a substantial performance improvement over the baseline methods, demonstrating the effectiveness of Transformers for medical image registration.
iPhantom: a framework for automated creation of individualized computational phantoms and its application to CT organ dosimetry
Aug 20, 2020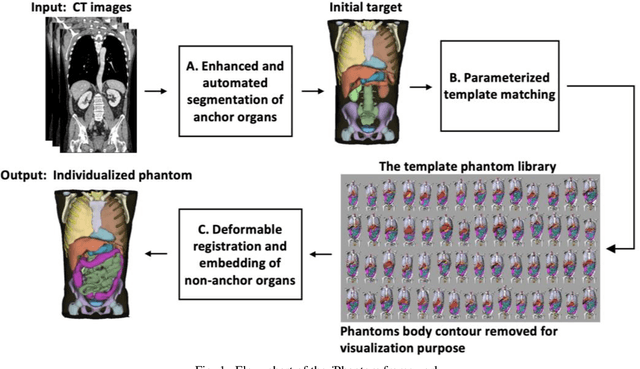
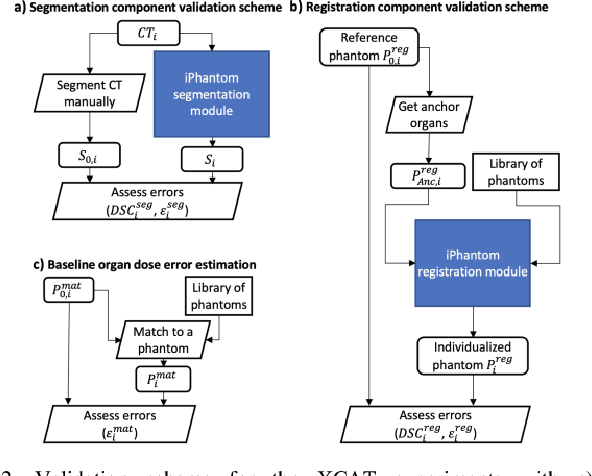
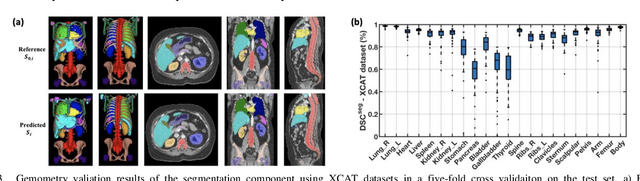
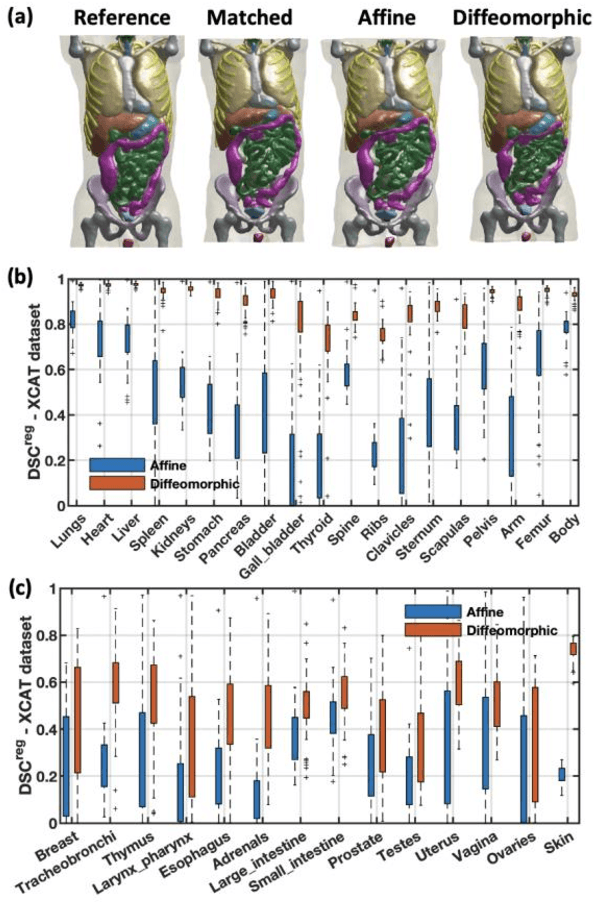
Abstract:Objective: This study aims to develop and validate a novel framework, iPhantom, for automated creation of patient-specific phantoms or digital-twins (DT) using patient medical images. The framework is applied to assess radiation dose to radiosensitive organs in CT imaging of individual patients. Method: From patient CT images, iPhantom segments selected anchor organs (e.g. liver, bones, pancreas) using a learning-based model developed for multi-organ CT segmentation. Organs challenging to segment (e.g. intestines) are incorporated from a matched phantom template, using a diffeomorphic registration model developed for multi-organ phantom-voxels. The resulting full-patient phantoms are used to assess organ doses during routine CT exams. Result: iPhantom was validated on both the XCAT (n=50) and an independent clinical (n=10) dataset with similar accuracy. iPhantom precisely predicted all organ locations with good accuracy of Dice Similarity Coefficients (DSC) >0.6 for anchor organs and DSC of 0.3-0.9 for all other organs. iPhantom showed less than 10% dose errors for the majority of organs, which was notably superior to the state-of-the-art baseline method (20-35% dose errors). Conclusion: iPhantom enables automated and accurate creation of patient-specific phantoms and, for the first time, provides sufficient and automated patient-specific dose estimates for CT dosimetry. Significance: The new framework brings the creation and application of CHPs to the level of individual CHPs through automation, achieving a wider and precise organ localization, paving the way for clinical monitoring, and personalized optimization, and large-scale research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge