Michael R. Harowicz
Transplant-Ready? Evaluating AI Lung Segmentation Models in Candidates with Severe Lung Disease
Sep 18, 2025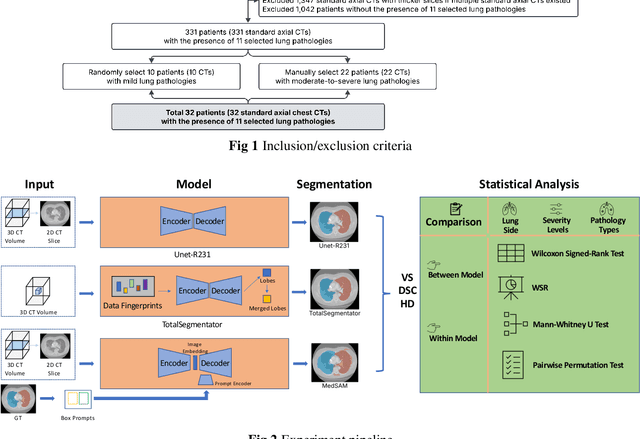
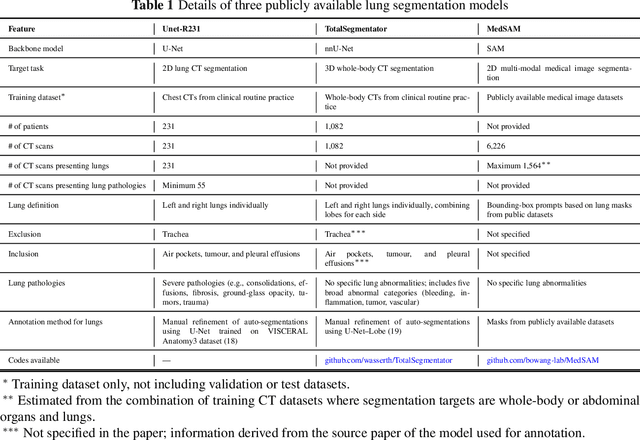
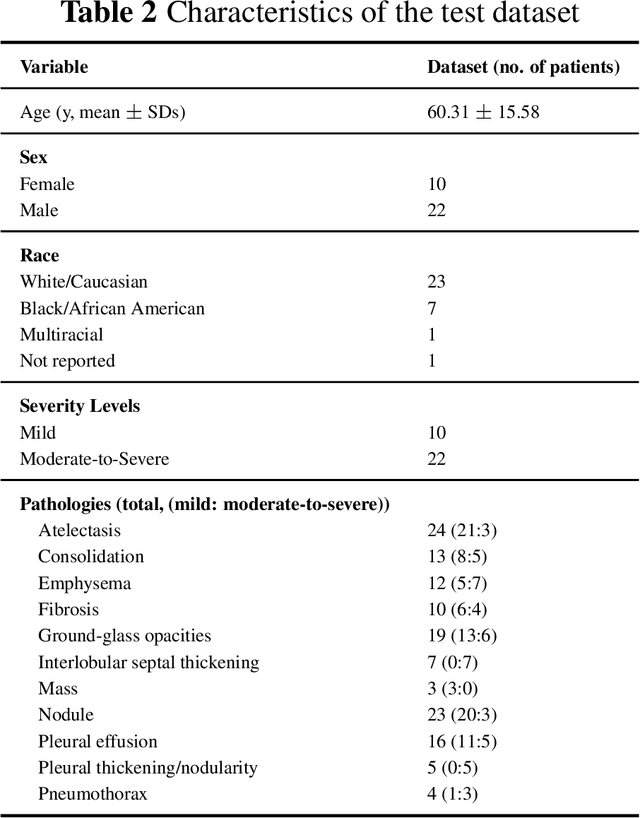
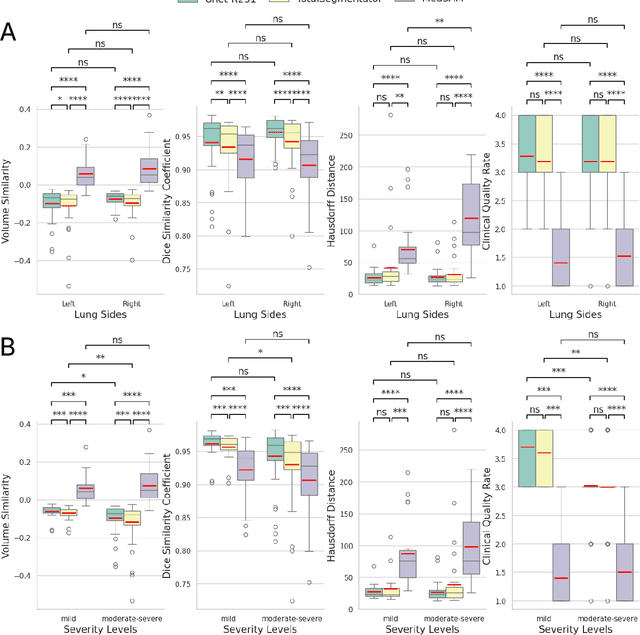
Abstract:This study evaluates publicly available deep-learning based lung segmentation models in transplant-eligible patients to determine their performance across disease severity levels, pathology categories, and lung sides, and to identify limitations impacting their use in preoperative planning in lung transplantation. This retrospective study included 32 patients who underwent chest CT scans at Duke University Health System between 2017 and 2019 (total of 3,645 2D axial slices). Patients with standard axial CT scans were selected based on the presence of two or more lung pathologies of varying severity. Lung segmentation was performed using three previously developed deep learning models: Unet-R231, TotalSegmentator, MedSAM. Performance was assessed using quantitative metrics (volumetric similarity, Dice similarity coefficient, Hausdorff distance) and a qualitative measure (four-point clinical acceptability scale). Unet-R231 consistently outperformed TotalSegmentator and MedSAM in general, for different severity levels, and pathology categories (p<0.05). All models showed significant performance declines from mild to moderate-to-severe cases, particularly in volumetric similarity (p<0.05), without significant differences among lung sides or pathology types. Unet-R231 provided the most accurate automated lung segmentation among evaluated models with TotalSegmentator being a close second, though their performance declined significantly in moderate-to-severe cases, emphasizing the need for specialized model fine-tuning in severe pathology contexts.
AI in Lung Health: Benchmarking Detection and Diagnostic Models Across Multiple CT Scan Datasets
May 07, 2024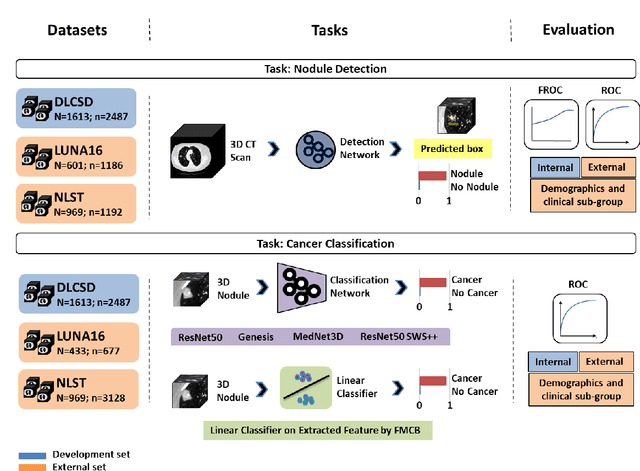
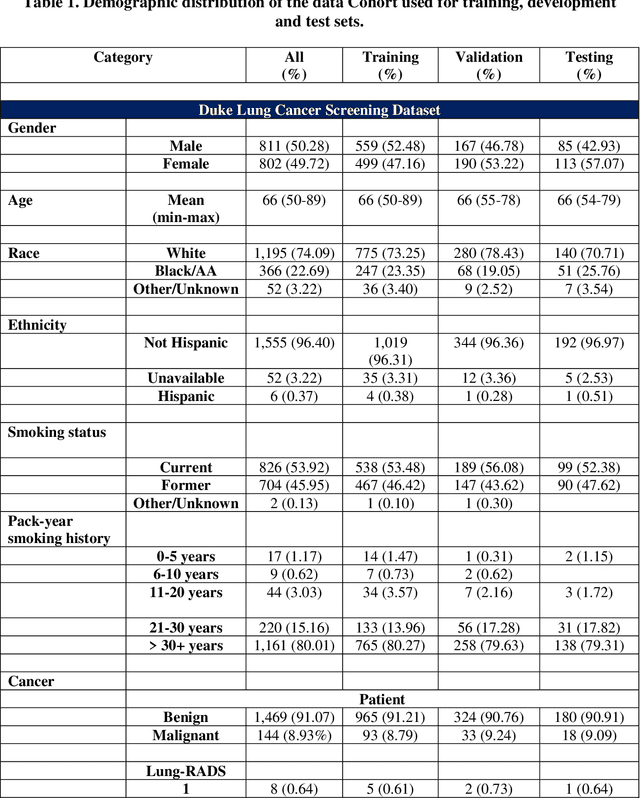
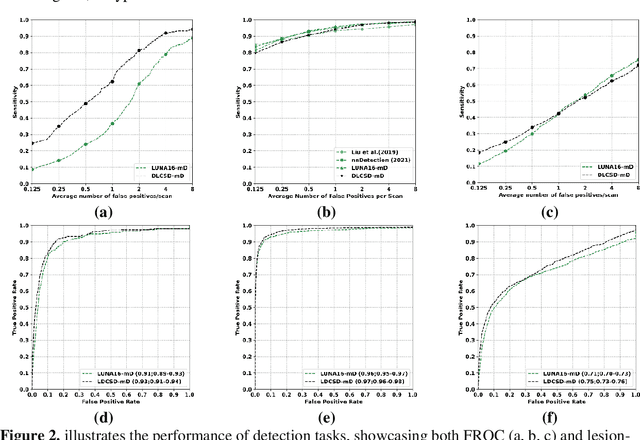
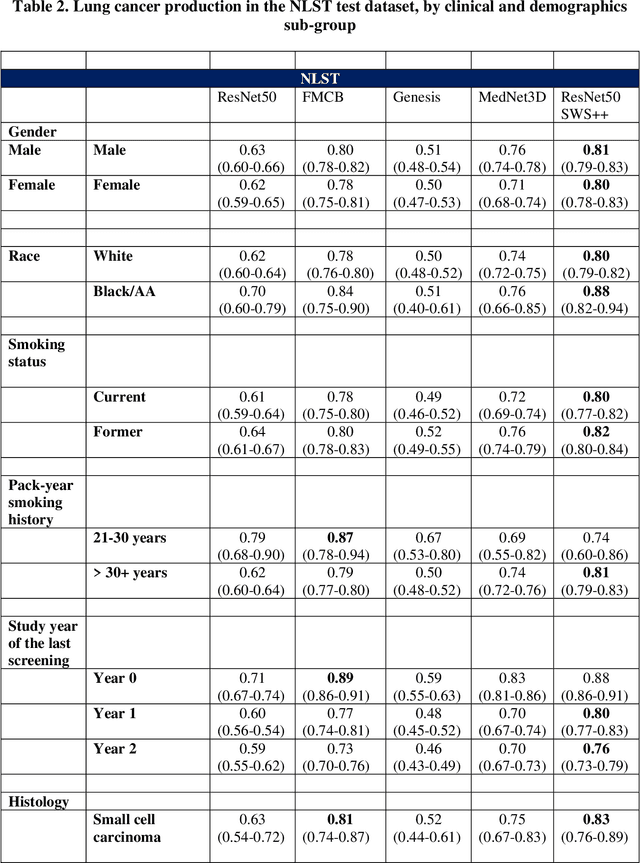
Abstract:BACKGROUND: Lung cancer's high mortality rate can be mitigated by early detection, which is increasingly reliant on artificial intelligence (AI) for diagnostic imaging. However, the performance of AI models is contingent upon the datasets used for their training and validation. METHODS: This study developed and validated the DLCSD-mD and LUNA16-mD models utilizing the Duke Lung Cancer Screening Dataset (DLCSD), encompassing over 2,000 CT scans with more than 3,000 annotations. These models were rigorously evaluated against the internal DLCSD and external LUNA16 and NLST datasets, aiming to establish a benchmark for imaging-based performance. The assessment focused on creating a standardized evaluation framework to facilitate consistent comparison with widely utilized datasets, ensuring a comprehensive validation of the model's efficacy. Diagnostic accuracy was assessed using free-response receiver operating characteristic (FROC) and area under the curve (AUC) analyses. RESULTS: On the internal DLCSD set, the DLCSD-mD model achieved an AUC of 0.93 (95% CI:0.91-0.94), demonstrating high accuracy. Its performance was sustained on the external datasets, with AUCs of 0.97 (95% CI: 0.96-0.98) on LUNA16 and 0.75 (95% CI: 0.73-0.76) on NLST. Similarly, the LUNA16-mD model recorded an AUC of 0.96 (95% CI: 0.95-0.97) on its native dataset and showed transferable diagnostic performance with AUCs of 0.91 (95% CI: 0.89-0.93) on DLCSD and 0.71 (95% CI: 0.70-0.72) on NLST. CONCLUSION: The DLCSD-mD model exhibits reliable performance across different datasets, establishing the DLCSD as a robust benchmark for lung cancer detection and diagnosis. Through the provision of our models and code to the public domain, we aim to accelerate the development of AI-based diagnostic tools and encourage reproducibility and collaborative advancements within the medical machine-learning (ML) field.
Deep Learning for identifying radiogenomic associations in breast cancer
Nov 29, 2017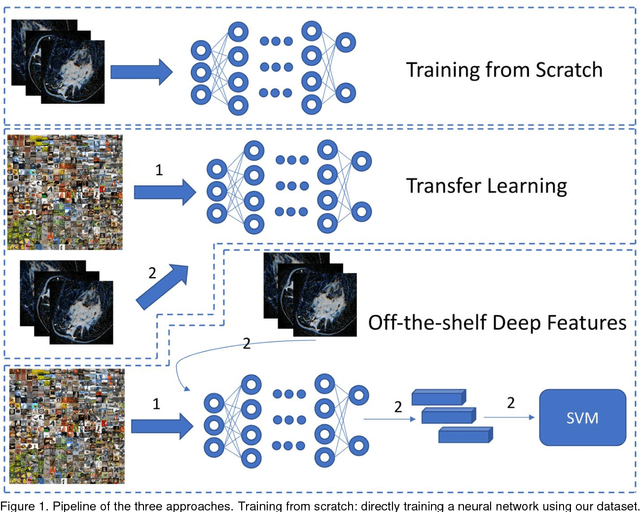
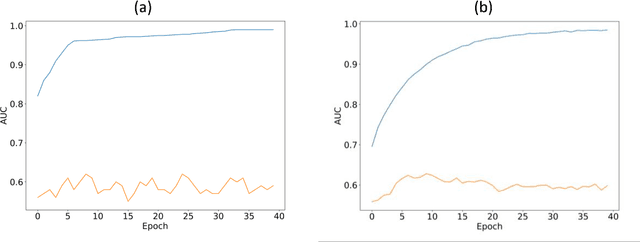

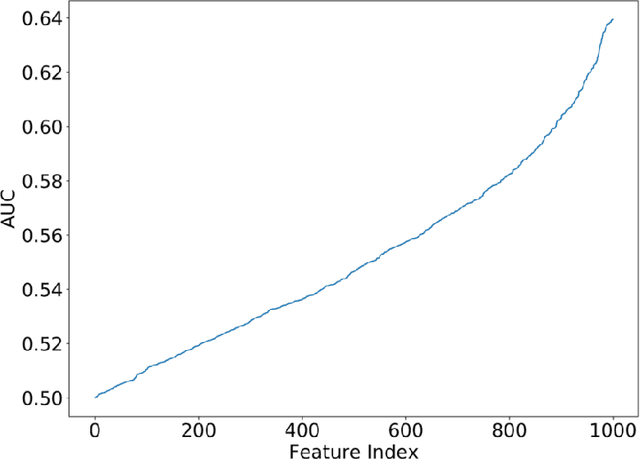
Abstract:Purpose: To determine whether deep learning models can distinguish between breast cancer molecular subtypes based on dynamic contrast-enhanced magnetic resonance imaging (DCE-MRI). Materials and methods: In this institutional review board-approved single-center study, we analyzed DCE-MR images of 270 patients at our institution. Lesions of interest were identified by radiologists. The task was to automatically determine whether the tumor is of the Luminal A subtype or of another subtype based on the MR image patches representing the tumor. Three different deep learning approaches were used to classify the tumor according to their molecular subtypes: learning from scratch where only tumor patches were used for training, transfer learning where networks pre-trained on natural images were fine-tuned using tumor patches, and off-the-shelf deep features where the features extracted by neural networks trained on natural images were used for classification with a support vector machine. Network architectures utilized in our experiments were GoogleNet, VGG, and CIFAR. We used 10-fold crossvalidation method for validation and area under the receiver operating characteristic (AUC) as the measure of performance. Results: The best AUC performance for distinguishing molecular subtypes was 0.65 (95% CI:[0.57,0.71]) and was achieved by the off-the-shelf deep features approach. The highest AUC performance for training from scratch was 0.58 (95% CI:[0.51,0.64]) and the best AUC performance for transfer learning was 0.60 (95% CI:[0.52,0.65]) respectively. For the off-the-shelf approach, the features extracted from the fully connected layer performed the best. Conclusion: Deep learning may play a role in discovering radiogenomic associations in breast cancer.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge