Andreas Nürnberger
Towards Segmenting the Invisible: An End-to-End Registration and Segmentation Framework for Weakly Supervised Tumour Analysis
Feb 05, 2026Abstract:Liver tumour ablation presents a significant clinical challenge: whilst tumours are clearly visible on pre-operative MRI, they are often effectively invisible on intra-operative CT due to minimal contrast between pathological and healthy tissue. This work investigates the feasibility of cross-modality weak supervision for scenarios where pathology is visible in one modality (MRI) but absent in another (CT). We present a hybrid registration-segmentation framework that combines MSCGUNet for inter-modal image registration with a UNet-based segmentation module, enabling registration-assisted pseudo-label generation for CT images. Our evaluation on the CHAOS dataset demonstrates that the pipeline can successfully register and segment healthy liver anatomy, achieving a Dice score of 0.72. However, when applied to clinical data containing tumours, performance degrades substantially (Dice score of 0.16), revealing the fundamental limitations of current registration methods when the target pathology lacks corresponding visual features in the target modality. We analyse the "domain gap" and "feature absence" problems, demonstrating that whilst spatial propagation of labels via registration is feasible for visible structures, segmenting truly invisible pathology remains an open challenge. Our findings highlight that registration-based label transfer cannot compensate for the absence of discriminative features in the target modality, providing important insights for future research in cross-modality medical image analysis. Code an weights are available at: https://github.com/BudhaTronix/Weakly-Supervised-Tumour-Detection
* Accepted for AIBio at ECAI 2025
SMILE-UHURA Challenge -- Small Vessel Segmentation at Mesoscopic Scale from Ultra-High Resolution 7T Magnetic Resonance Angiograms
Nov 14, 2024Abstract:The human brain receives nutrients and oxygen through an intricate network of blood vessels. Pathology affecting small vessels, at the mesoscopic scale, represents a critical vulnerability within the cerebral blood supply and can lead to severe conditions, such as Cerebral Small Vessel Diseases. The advent of 7 Tesla MRI systems has enabled the acquisition of higher spatial resolution images, making it possible to visualise such vessels in the brain. However, the lack of publicly available annotated datasets has impeded the development of robust, machine learning-driven segmentation algorithms. To address this, the SMILE-UHURA challenge was organised. This challenge, held in conjunction with the ISBI 2023, in Cartagena de Indias, Colombia, aimed to provide a platform for researchers working on related topics. The SMILE-UHURA challenge addresses the gap in publicly available annotated datasets by providing an annotated dataset of Time-of-Flight angiography acquired with 7T MRI. This dataset was created through a combination of automated pre-segmentation and extensive manual refinement. In this manuscript, sixteen submitted methods and two baseline methods are compared both quantitatively and qualitatively on two different datasets: held-out test MRAs from the same dataset as the training data (with labels kept secret) and a separate 7T ToF MRA dataset where both input volumes and labels are kept secret. The results demonstrate that most of the submitted deep learning methods, trained on the provided training dataset, achieved reliable segmentation performance. Dice scores reached up to 0.838 $\pm$ 0.066 and 0.716 $\pm$ 0.125 on the respective datasets, with an average performance of up to 0.804 $\pm$ 0.15.
CEAR: Automatic construction of a knowledge graph of chemical entities and roles from scientific literature
Jul 31, 2024


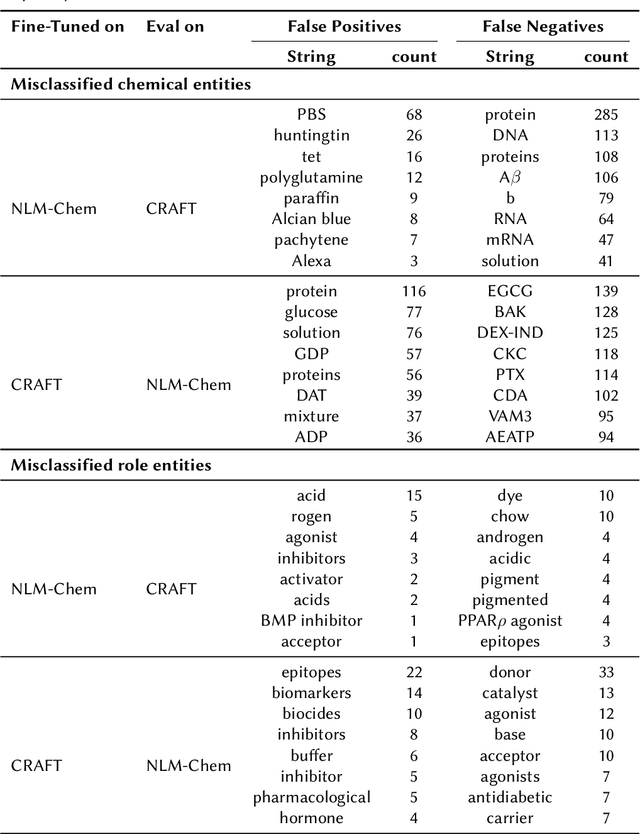
Abstract:Ontologies are formal representations of knowledge in specific domains that provide a structured framework for organizing and understanding complex information. Creating ontologies, however, is a complex and time-consuming endeavor. ChEBI is a well-known ontology in the field of chemistry, which provides a comprehensive resource for defining chemical entities and their properties. However, it covers only a small fraction of the rapidly growing knowledge in chemistry and does not provide references to the scientific literature. To address this, we propose a methodology that involves augmenting existing annotated text corpora with knowledge from Chebi and fine-tuning a large language model (LLM) to recognize chemical entities and their roles in scientific text. Our experiments demonstrate the effectiveness of our approach. By combining ontological knowledge and the language understanding capabilities of LLMs, we achieve high precision and recall rates in identifying both the chemical entities and roles in scientific literature. Furthermore, we extract them from a set of 8,000 ChemRxiv articles, and apply a second LLM to create a knowledge graph (KG) of chemical entities and roles (CEAR), which provides complementary information to ChEBI, and can help to extend it.
SPOCKMIP: Segmentation of Vessels in MRAs with Enhanced Continuity using Maximum Intensity Projection as Loss
Jul 11, 2024



Abstract:Identification of vessel structures of different sizes in biomedical images is crucial in the diagnosis of many neurodegenerative diseases. However, the sparsity of good-quality annotations of such images makes the task of vessel segmentation challenging. Deep learning offers an efficient way to segment vessels of different sizes by learning their high-level feature representations and the spatial continuity of such features across dimensions. Semi-supervised patch-based approaches have been effective in identifying small vessels of one to two voxels in diameter. This study focuses on improving the segmentation quality by considering the spatial correlation of the features using the Maximum Intensity Projection~(MIP) as an additional loss criterion. Two methods are proposed with the incorporation of MIPs of label segmentation on the single~(z-axis) and multiple perceivable axes of the 3D volume. The proposed MIP-based methods produce segmentations with improved vessel continuity, which is evident in visual examinations of ROIs. Patch-based training is improved by introducing an additional loss term, MIP loss, to penalise the predicted discontinuity of vessels. A training set of 14 volumes is selected from the StudyForrest dataset comprising of 18 7-Tesla 3D Time-of-Flight~(ToF) Magnetic Resonance Angiography (MRA) images. The generalisation performance of the method is evaluated using the other unseen volumes in the dataset. It is observed that the proposed method with multi-axes MIP loss produces better quality segmentations with a median Dice of $80.245 \pm 0.129$. Also, the method with single-axis MIP loss produces segmentations with a median Dice of $79.749 \pm 0.109$. Furthermore, a visual comparison of the ROIs in the predicted segmentation reveals a significant improvement in the continuity of the vessels when MIP loss is incorporated into training.
PULASki: Learning inter-rater variability using statistical distances to improve probabilistic segmentation
Dec 25, 2023



Abstract:In the domain of medical imaging, many supervised learning based methods for segmentation face several challenges such as high variability in annotations from multiple experts, paucity of labelled data and class imbalanced datasets. These issues may result in segmentations that lack the requisite precision for clinical analysis and can be misleadingly overconfident without associated uncertainty quantification. We propose the PULASki for biomedical image segmentation that accurately captures variability in expert annotations, even in small datasets. Our approach makes use of an improved loss function based on statistical distances in a conditional variational autoencoder structure (Probabilistic UNet), which improves learning of the conditional decoder compared to the standard cross-entropy particularly in class imbalanced problems. We analyse our method for two structurally different segmentation tasks (intracranial vessel and multiple sclerosis (MS) lesion) and compare our results to four well-established baselines in terms of quantitative metrics and qualitative output. Empirical results demonstrate the PULASKi method outperforms all baselines at the 5\% significance level. The generated segmentations are shown to be much more anatomically plausible than in the 2D case, particularly for the vessel task. Our method can also be applied to a wide range of multi-label segmentation tasks and and is useful for downstream tasks such as hemodynamic modelling (computational fluid dynamics and data assimilation), clinical decision making, and treatment planning.
Fourier PD and PDUNet: Complex-valued networks to speed-up MR Thermometry during Hypterthermia
Oct 02, 2023Abstract:Hyperthermia (HT) in combination with radio- and/or chemotherapy has become an accepted cancer treatment for distinct solid tumour entities. In HT, tumour tissue is exogenously heated to temperatures of 39 to 43 $\degree$C for 60 minutes. Temperature monitoring can be performed noninvasively using dynamic magnetic resonance imaging (MRI). However, the slow nature of MRI leads to motion artefacts in the images due to the movements of patients during image acquisition time. By discarding parts of the data, the speed of the acquisition can be increased - known as Undersampling. However, due to the invalidation of the Nyquist criterion, the acquired images have lower resolution and can also produce artefacts. The aim of this work was, therefore, to reconstruct highly undersampled MR thermometry acquisitions with better resolution and with less artefacts compared to conventional techniques like compressed sensing. The use of deep learning in the medical field has emerged in recent times, and various studies have shown that deep learning has the potential to solve inverse problems such as MR image reconstruction. However, most of the published work only focusses on the magnitude images, while the phase images are ignored, which are fundamental requirements for MR thermometry. This work, for the first time ever, presents deep learning based solutions for reconstructing undersampled MR thermometry data. Two different deep learning models have been employed here, the Fourier Primal-Dual network and Fourier Primal-Dual UNet, to reconstruct highly undersampled complex images of MR thermometry. It was observed that the method was able to reduce the temperature difference between the undersampled MRIs and the fully sampled MRIs from 1.5 $\degree$C to 0.5 $\degree$C.
MedShapeNet -- A Large-Scale Dataset of 3D Medical Shapes for Computer Vision
Sep 12, 2023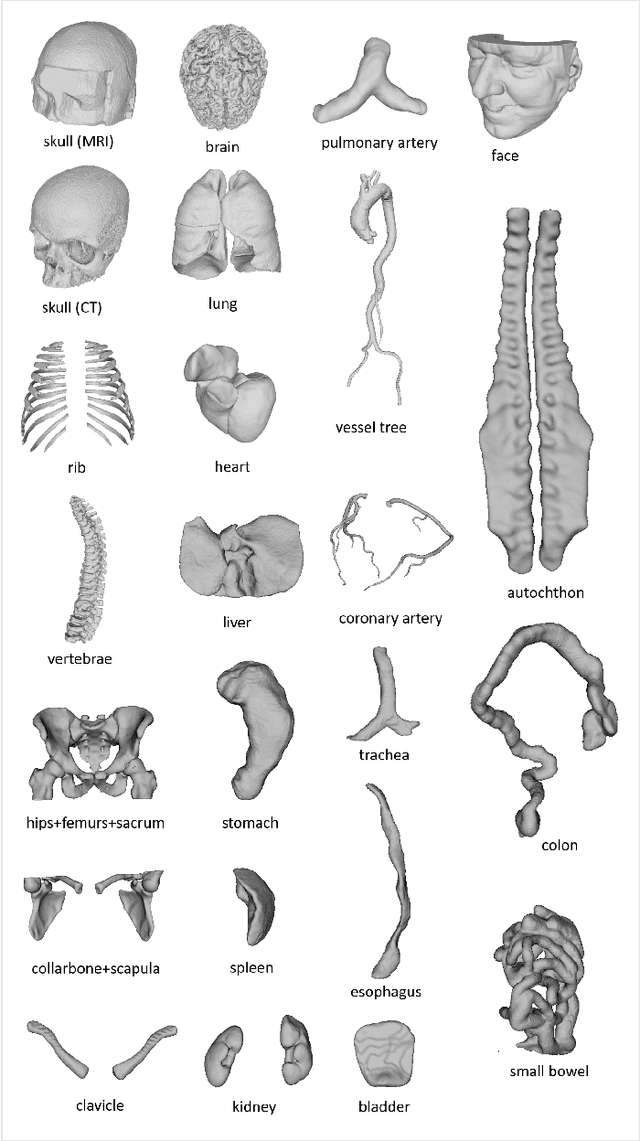

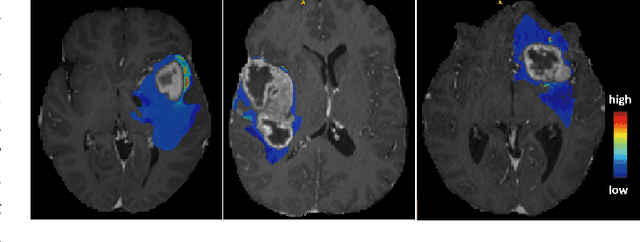
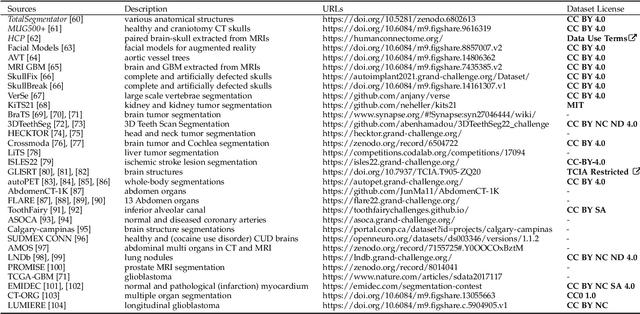
Abstract:We present MedShapeNet, a large collection of anatomical shapes (e.g., bones, organs, vessels) and 3D surgical instrument models. Prior to the deep learning era, the broad application of statistical shape models (SSMs) in medical image analysis is evidence that shapes have been commonly used to describe medical data. Nowadays, however, state-of-the-art (SOTA) deep learning algorithms in medical imaging are predominantly voxel-based. In computer vision, on the contrary, shapes (including, voxel occupancy grids, meshes, point clouds and implicit surface models) are preferred data representations in 3D, as seen from the numerous shape-related publications in premier vision conferences, such as the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), as well as the increasing popularity of ShapeNet (about 51,300 models) and Princeton ModelNet (127,915 models) in computer vision research. MedShapeNet is created as an alternative to these commonly used shape benchmarks to facilitate the translation of data-driven vision algorithms to medical applications, and it extends the opportunities to adapt SOTA vision algorithms to solve critical medical problems. Besides, the majority of the medical shapes in MedShapeNet are modeled directly on the imaging data of real patients, and therefore it complements well existing shape benchmarks comprising of computer-aided design (CAD) models. MedShapeNet currently includes more than 100,000 medical shapes, and provides annotations in the form of paired data. It is therefore also a freely available repository of 3D models for extended reality (virtual reality - VR, augmented reality - AR, mixed reality - MR) and medical 3D printing. This white paper describes in detail the motivations behind MedShapeNet, the shape acquisition procedures, the use cases, as well as the usage of the online shape search portal: https://medshapenet.ikim.nrw/
Liver Segmentation in Time-resolved C-arm CT Volumes Reconstructed from Dynamic Perfusion Scans using Time Separation Technique
Feb 09, 2023



Abstract:Perfusion imaging is a valuable tool for diagnosing and treatment planning for liver tumours. The time separation technique (TST) has been successfully used for modelling C-arm cone-beam computed tomography (CBCT) perfusion data. The reconstruction can be accompanied by the segmentation of the liver - for better visualisation and for generating comprehensive perfusion maps. Recently introduced Turbolift learning has been seen to perform well while working with TST reconstructions, but has not been explored for the time-resolved volumes (TRV) estimated out of TST reconstructions. The segmentation of the TRVs can be useful for tracking the movement of the liver over time. This research explores this possibility by training the multi-scale attention UNet of Turbolift learning at its third stage on the TRVs and shows the robustness of Turbolift learning since it can even work efficiently with the TRVs, resulting in a Dice score of 0.864$\pm$0.004.
Complex Network for Complex Problems: A comparative study of CNN and Complex-valued CNN
Feb 09, 2023



Abstract:Neural networks, especially convolutional neural networks (CNN), are one of the most common tools these days used in computer vision. Most of these networks work with real-valued data using real-valued features. Complex-valued convolutional neural networks (CV-CNN) can preserve the algebraic structure of complex-valued input data and have the potential to learn more complex relationships between the input and the ground-truth. Although some comparisons of CNNs and CV-CNNs for different tasks have been performed in the past, a large-scale investigation comparing different models operating on different tasks has not been conducted. Furthermore, because complex features contain both real and imaginary components, CV-CNNs have double the number of trainable parameters as real-valued CNNs in terms of the actual number of trainable parameters. Whether or not the improvements in performance with CV-CNN observed in the past have been because of the complex features or just because of having double the number of trainable parameters has not yet been explored. This paper presents a comparative study of CNN, CNNx2 (CNN with double the number of trainable parameters as the CNN), and CV-CNN. The experiments were performed using seven models for two different tasks - brain tumour classification and segmentation in brain MRIs. The results have revealed that the CV-CNN models outperformed the CNN and CNNx2 models.
Liver Segmentation using Turbolift Learning for CT and Cone-beam C-arm Perfusion Imaging
Jul 20, 2022
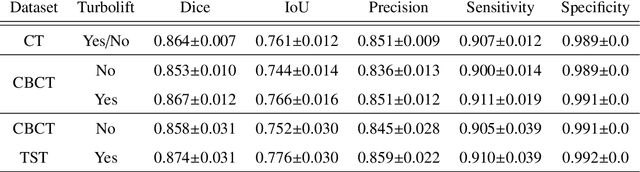
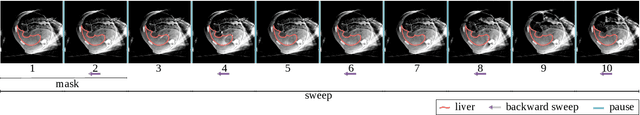
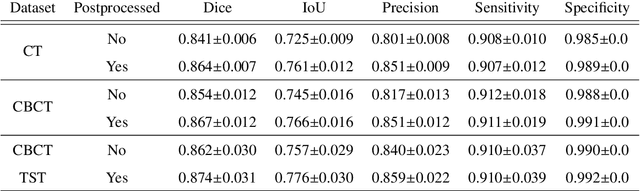
Abstract:Model-based reconstruction employing the time separation technique (TST) was found to improve dynamic perfusion imaging of the liver using C-arm cone-beam computed tomography (CBCT). To apply TST using prior knowledge extracted from CT perfusion data, the liver should be accurately segmented from the CT scans. Reconstructions of primary and model-based CBCT data need to be segmented for proper visualisation and interpretation of perfusion maps. This research proposes Turbolift learning, which trains a modified version of the multi-scale Attention UNet on different liver segmentation tasks serially, following the order of the trainings CT, CBCT, CBCT TST - making the previous trainings act as pre-training stages for the subsequent ones - addressing the problem of limited number of datasets for training. For the final task of liver segmentation from CBCT TST, the proposed method achieved an overall Dice scores of 0.874$\pm$0.031 and 0.905$\pm$0.007 in 6-fold and 4-fold cross-validation experiments, respectively - securing statistically significant improvements over the model, which was trained only for that task. Experiments revealed that Turbolift not only improves the overall performance of the model but also makes it robust against artefacts originating from the embolisation materials and truncation artefacts. Additionally, in-depth analyses confirmed the order of the segmentation tasks. This paper shows the potential of segmenting the liver from CT, CBCT, and CBCT TST, learning from the available limited training data, which can possibly be used in the future for the visualisation and evaluation of the perfusion maps for the treatment evaluation of liver diseases.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge